ppt
advertisement

Fine Scale Mapping and the Coalescent
•The Fundamental Problem
•The Data
•Genotype to Phenotype Functions
•Types of Mapping
•Population Set-up & Measures of Dependency
•The Calculations
•Practical Considerations
Genotype and Phenotype Covariation: Gene Mapping
Sampling Genotypes and Phenotypes
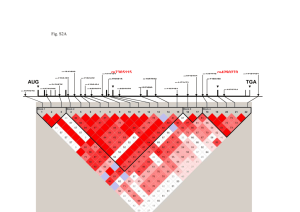
Decay of local dependency
Time
Reich et al. (2001)
Genetype -->Phenotype Function
Result:The Mapping Function
Dominant/Recessive.
Penetrance
A set of characters.
Binary decision (0,1).
Spurious Occurrence
Quantitative Character.
Heterogeneity
genotype
Genotype Phenotype
phenotype
Pedigree Analysis & Association Mapping
Association Mapping:
Pedigree Analysis:
M
r
D
Pedigree known
Few meiosis (max 100s)
D
2N generations
M
r
Resolution: cMorgans (Mbases)
Pedigree unknown
Many meiosis (>104)
Adapted from McVean and others
Resolution: 10-5 Morgans (Kbases)
Causes of linkage disequilibrium
D
M
Time t ago
D
M
Now
Creates LD
Breaks down LD
Drift
Selection
Admixture
Recombination
Gene conversion
Significance of a Single Association
Disease locus
Marker locus
Disease locus
Marker locus
Test for independence in 2 times 2 Contingency Table
XA,B Xa,B
X.,B
XA,b Xa,b
X.,b
XA,.
X.,.
Xa,.
Measuring Linkage Disequilibrium between 2 Loci with 2 Alleles
Remade from McVean
DA,B =fA,B-fAfB =-Da,B =-DA,b =Da,b
Correlation Coeffecient Measure [0,1]
Hill & Robertson (1968)
2
DAB
2
r
AB
f A fa f B fb
2
AB
Range constrained by allele frequencies [0,1]
Lewontin (1964)
'
DAB
if ( D 0)
Odds-ratio formulation
Devlin & Risch (1995)
AB
f AB
fB f A
DAB
DAB
else
min( f A , f b , f a , f B , )
min( f A , f b , f a , f B , )
Examples of Associations: Pairwise, Triple,...
Disease locus
Marker loci
Combine Single (Pairwise) to Multiple Tests
Bonferroni
Sharper bounds using linkage information.
ApoE and Alzheimers Syndrome
Causative SNP
6 markers with
low association
Martin et al 2000
The coalescent with recombination or gene conversion
Adapted from Hudson 1990
Recombination:
Gene Conversion:
Local trees for recombination and gene conversion
Gene conversion
Recombination
1 2
3 4
Tree 1
1
2
4
Tree 2
3
1 2 3
Tree 3
4
1 2
3 4
Tree 1
1
2
4
Tree 2
3
1 2 3
4
Tree 1
Target tree
Measures of tree similarity
Target
Region with no recombination
Same tree as target
Same topology as target
1
2
3
4
Same tree
1
2
3
Same MRCA as target
5
Same topology
4
5
1 2
3
4
Same MRCA
5
1
2
3 4 5
Local trees of the target and other positions
Sample size = 20
Only recombination, r=2.
Also gene conversion g/4
From Mikkel Schierup
Probability that the largest segment does not include the target
Recombination/gene conversion
rate
R=2, G=0
R=2,
G=8
#segments with same tree
1.02
1.8
P(target segment not largest)
0.2%
14%
#segments same topology
1.02
2.1
P(target segment not largest)
0.3%
20%
1.1
2.9
1.5%
25%
#segments same TMRCA
P(target segment not largest)
From Mikkel Schierup
Quantifying the mosaicism caused by Gene Conversion
A and B are the most distant markers in significant LD with target
A
Target
B
What is the proportion of markers between these also in significant LD?
Rho=4
G=0
56%
G=16
33%
From Mikkel Schierup
Development of multi-locus association methods
Single Marker Methods
•Kaplan et al. (1995), Rannala & Slatkin (1998)
Problem: Difficult to combine markers.
Haplotype methods with star-shaped genealogies
•Terwilliger (1995), Graham & Thompson (1998), McPeek &
Strahs(1999), Morris et al.(2000)
Problem: wrong genealogy, gives overconfidence in result.
Haplotype methods based on the coalescent
•Rannala & Reeve (2001), Morris et al. (2002), Larribe et al. (2003).
Problem: computationally intensive
Based on Morris et al. 2002
Probability of Data I:
3 step approach:
I Probability of Data given topology and branch lengths
Felsenstein81 for each column
Multiply for all columns
GCAGGTT
TCAGCCT
TCAGCAT
II Integrate over branch lengths
III Sum over topologies
0
0
0
P(Data Topo,tk tk1 ..t2)e
k
tk /
2
k1
tk 1 /
2 t
dtk e
j
2
j n
3
Conclusion: Exact Calculation Computationally Intractible!!
e dtk1 ....dt2
Probability of Data II:
Griffiths & Tavavé TPB46.2.131-149
q(n’’) – determined by equilibrium distribution.
q(n)
ACCTAGGAT
n'n
TCCTAGGAT
(1,2) coalescence
3*9*3 mutations
ACCTAGGAT
n=
q(n') f (n,n')
TCCTAGGAT
TCCTAGGAT
Griffiths-Ethier-Tavare Recursions
nk (nk 1)
pa (T , n ek )
n
(
n
1
)
k :nk 2
pa (T , n)
n=(3,1,2)
n
d ,1
pa (T ' , n' )
n=(2,1,2)
n=(3,1,2)
1
1
2
3
1
2
3
1
2
3
2
Griffiths-Marjoram (1996) included recombination in the equations.
Example: Solving Linear System
q( x ) r ( x, y )q( y ) r ( x, z )q( z ), for x B
yA
zB
q( x ) known when x A and unknown when x B.
??
q( )
r(,)
r(,)
r(,)
r(,)
??
??
r(,)
q( )
r(,)
r(,)
r(,)
r(,)
q( )
??
q( x ) r ( x, y )q( y ) r ( x, y1 )r ( y1 , y )q( y )
yA
y1B yA
r( x, y )r( y , y )r( y , y )q( y ) ....
y1B y2B yA
1
1
2
2
{ ...... r ( x, y1 )...r ( yk , y )q( y )}
k 0
y1B
yk B yA
Example: Solving Linear System
Construct Markov transition function, A(x,y), with following properties:
i) A(x,y) > 0 when r(x,y) >0
ii) The chain visits A with certainty.
j
r ( X kj1 , X k )
q( x0 ) E x0 {q( X )
}
j
k 1 A( X k 1 , X k )
j
j
j
r
(
X
,
X
)
1 m
k 1
j
k
qˆ q( X j )
}
j
j
m j 1
k 1 A( X k 1 , X k )
•Introduced in coalescence theory by Griffiths & Tavare (1994)
•Griffiths & Marjoram (1996) included recombination
•Donnelly-Stephens-Fearnhead (2000-) accelerated these algorithms
The position of the marker locus is missing data
Larribe and Lessard.(2002)
Data:
haplotype
phenotype
multiplicity
15
3
6
2
1
2
1
Where is the disease causing disease?
Likelihood as function of disease locus position
Bayesian approach to LD mapping
Continuous version of Bayes formula
P(data | parameters ) f (parameter s)
f (parameters | data)
P(data)
f (parameters) = prior distribution of parameters
P(data|parameters) = L(parameters) = likelihood function
f (P|D) = posterior distribution of parameters given data
The evolutionary parameter (e.g. disease location) is considered to have
prior distribution (any prior knowledge we may have)
and we learn about parameters through data
Advantage: f (parameters|data) is the full distribution of parameters of
interest given data, e.g. confidence intervals
The basic equation
P(data | parameters ) f (parameter s)
f (parameters | data)
P(data)
Marginal posterior distribution of disease position:
P(disease position x | data)
P(paramete rs | data) dP ...dP
1
Parameters except x
n
Parameters in Shattered Coalescent Model
Morris, Whittaker and Balding (2001,,2003,2004..
P(x,h,W,T,z,N,|A,U) ~ L(A,U|x,h,W,T,z,N) p(W,T,z|) p()
p() = 2,p(W,T,z|) prior distribution of genealogies (coalescent like)
x
h
W
T
Z
N
A, U
Location of disease locus
Population marker-haplotype proportions
branch lengths of genealogical tree
topology (branching pattern)
Parental-status
effective population size Probability of Haplotypes associated Mutant
shattering parameter
cases, controls
At recombination markers
are incorporated from the
population distribution.
Morris et al: The Shattered Coalescent
Advantages: Allows for multiple origins of the disease mutant
+ sporadic occurrences of the disease without the mutation
Coalescent tree
Morris, Whittaker & Balding,2002
Monte-Carlo (Metropolis) sampling and integration
Metropolis et al.(1953)
P(disease position x )
P(paramete rs | data) dP ...dP
1
n
Parameters except x
•Evaluate the function in the current point p, f(p)=x
•Suggest a new point, p'
•Evaluate the function in this point f(p') = y
•If x < y, go to point p'
•If x > y, go to point p' with the probability y/x
Due to Jesper Nymann
Monte-Carlo (Metropolis)
Projection on one axis equivalent
to integration over the remaining
parameters
1
2!
1
2?
3
2?
2
1
Due to Jesper Nymann
Example 1 - Cystic fibrosis
11
19
Morris et al. (2002).
Due to Jesper Nymann
Example 2 - BRCA2
Iceland Genomics Corporation:
1132 Cases, 54 with known mutation
758 Controls
Due to Jesper Nymann
Example 2 - BRCA2 continued
True Location
1
3
5
7
9
11
13
15
Multipoint calculation for the full BRCA2 dataset
1
3
5
7
9
11
13
15
Multipoint calculation where the 54 known
mutation cases has been removed.
Due to Jesper Nymann
The Basic Setup
Simulation Parameters:
Recombination rate = 50
Number of leaf nodes = 1000
Number of markers = 10
Diseased haplotype fraction: 0.08 – 0.12
No Heterogeneity
Simulated under the asumption of constant population size
Diplotypes (phase known)
Type of simulation
Basic (red curve)
50% quantile
0.044
Due to Jesper Nymann
The effect of marker density
Type of simulation
19 markers (blue curve)
19 markers and recombination rate = 100 (yellow curve)
Basic (red curve)
50% quantile
0.0292
0.02321
0.044
Due to Jesper Nymann
The effect of knowing phase
0
1
0/1
0
0/1
1
Type of simulation
With Genotype data (blue curve)
Basic (red curve)
0/1
0
0/1
0
1
1
0
0
1
1
0
1
0
0
1
0
0
1
1
0
0
0
0
0
50% quantile
0.05857
0.044
Due to Jesper Nymann
The Effect of knowing gene genealogy
Type of simulation
With known genealogy (blue curve)
Basic (red curve)
50% quantile
0.03516
0.044
Due to Jesper Nymann
The effect of disease fraction
Type of simulation
Disease fraction 12% - 14% (blue curve)
Disease fraction 18% - 22% (yellow curve)
Basic (red curve)
50% quantile
0.0353
0.03229
0.044
Due to Jesper Nymann
The effect of Heterogeneity
Type of simulation
50% quantile
With Heterogeneity (blue curve) 0.065587
Basic (red curve)
0.044
Due to Jesper Nymann
The effect of Impurity of cases and controls
Cases
Controls
33% cases are moved to
the controls and a similar
number of controls are
moved
to the cases
Type of simulation
With mixed cases/controls (blue curve)
Basic (red curve)
50% quantile
0.1518
0.044
Due to Jesper Nymann
LD in background population
No LD in background:
P(0) P(1) P(1) P(0) P(0) P(1) P(1) P(0) P(1) P(0)
0
LD in background:
1
1
0
0
1
1
0
1
0
P(0) P(1|0) P(1|1) P(0|1) P(0|0) P(1|0) P(1|1) P(0|1) P(1|0) P(0|1)
Gene
Pool
Type of simulation
LD in background (blue curve)
Basic (red curve)
50% quantile
0.0419
0.044
Due to Jesper Nymann
Comparing the different scenarios
Simulation Type
Mean
50% Quantile
70% Quantile
95% Quantile
Basic
19 markes rho=100
19 markers
18% - 22% cases
12% - 14% cases
Fixed topology
LD in background
Genotype Data
Heterogeneity
33% impure
0,059
0,044
0,053
0,046
0,048
0,047
0,078
0,087
0,088
0,173
0,044
0,023
0,029
0,032
0,035
0,035
0,042
0,059
0,066
0,152
0,070
0,043
0,047
0,052
0,050
0,058
0,072
0,099
0,092
0,217
0,193
0,142
0,176
0,146
0,136
0,111
0,273
0,305
0,246
0,452
Random
0,303
0,273
0,407
0,696
Due to Jesper Nymann
Summary
The Fundamental Problem
The Data
Genotype to Phenotype Functions
Types of Mapping
Population Set-up & Measures of Dependency
Methods:
Pure Coalescent Based
The Shattered Coalescent
Factors influencing mapping error.
Articles I
M. A. Beaumont and B. Rannala (2004) The Bayesian Revolution in genetics, Nature Reviews, Genetics vol. 5. 251
Botstein D, Risch N. (2003) Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for
complex disease. Nat Genet. 33 Suppl:228-237. Cardon, L. and J. Bell (2001) “Association Study Designs for Complex Diseases “ Nature
Review Genetics
Daly, M. J., Rioux, J. D., Schaner, S. F., Hudson, T. J. & Lander, E. S. (2001), High-resolution haplotype structure in the human genome, Nat
Genet 29(2), 229-232.
Devlin, B. & Roeder, K. (1999), Genomic control for association studies, Biometrics 55(4), 997-1004.
Frisse, L et al.(2001) Gene Conversion and Different Population Histories May Explain the Contrast between Polymorphisms and LD Levels.
AJHG 69..?-?
Gabriel, S. B. et al. (2002), The structure of haplotype blocks in the human genome, Science 296(5576), 2225-2229.
Griffiths,R & S. Tavare (1994) “ Simiulating probability distributions in the coalescent ” Theor.Pop.Biol. 46.2.131-159
Griifiths, R. and P. Marjoram (1996) “Ancestral inference from samples of DNA sequences with recombination ”J.Compu.Biol.
Hudson, R. R. (1990).Gene genealogies and the coalescent process, “Oxford Surveys in Evolutionary Biology” (D. futuyma and J. Antonovics,
Eds.) Vol 7, pp. 1-44, Oxford Univ. Press, Oxford, UK
B. Kerem, J. M. Rommens, J. A. Buchanan D. Markiewicz, T. K. Cox, A. Chakravarti, M. Buchwald and L. C. Tsui Identification of the Cystic
Fibrosis Gene: Genetic Analysis Science 245: 1073-1080, 1989
Kong A, et al. (2002) A high-resolution recombination map of the human genome. Nat Genet. 31,241-7.
Laitinen et al. (2004) Characterization of a common susceptibility locus for Asthma-related traits. Nature 304, 300-304.
Martin, E. R., et al. (2000), SNPing away at complex diseases: analysis of single-nucleotide polymorphisms around APOE in Alzheimer
disease, Am J Hum Genet 67, 383-394.
Larribe, M, S. Lessard and Schork (2002) “Gene Mapping via the Ancestral Recombination Graph”. Theor. Pop.Biol. 62.215-229.
Liu,J. et al.(2000) “Bayesian Analysis of Haplotypes for Linkage Disequilibrium Mapping” Genome Research 11.1716-24.
Martin, E. et al.(2001) “SNPing Away at Complex Diseases: Analysis of Single-Nucleotide Polymorphisms around APOE Alzheimer Disease”
AJHG 67.838-394.
N Metropolis N AW Rosenbluth, MN Rosenbluth, AH Teller, E Teller (1953) Equation of state calculation by fast computer machines, J. Chem.
Phys. 21:1087-1092
McVean,G.(2002) “A Genealogical Interpretation of Linkage Disequilibrium” Genetics 162.987-991
Morris, A., JC Whittaker and D. Balding “Fine-Scale Mapping of Disease Loci via Shattered Coalescent Modeling of Genealogies” AJHG
70.686-707.
Morris, J. C. Whittaker, and D. J. Balding (2004) Little loss of information due to unknown phase for fine-scale LD mapping with SNP
genotype data, AJHG . 74: 945-953, 2004
Andrew P. Morris, John C. Whittaker, Chun-Fang Xu, Louise K. Hosking, and David J. Balding Multipoint linkage-disequilibrium mapping
narrows location interval and identifies mutation heterogeneity, PNAS November 11, 2003, Vol. 100, 13442-13446
Articles II
McVean GA, Myers SR, Hunt S, Deloukas P, Bentley DR, Donnelly P. (2004) The fine-scale structure of recombination rate variation
in the human genome. Science 304:581-584.
Patil, N. et al. (2001) Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science
294: 1719-1723.
Reich, D. E. et al. (2001), Linkage disequilibrium in the human genome, Nature 411(6834), 199-204.
Reich D. E. and Lander, E. On the allelic spectrum of human diseases. Trends in Genetics 19, 502-510.
Reich, D. E. et al. (2002), Human genome sequence variation and the influence of gene history, mutation and recombination, Nat
Genet 32(1), 135-142.
Risch, N. and Merikangas, K. (1996) The future of genetic studies of complex human diseases. Science 273, 15161-1517.
Pritchard, J. K., Stephens, M., Rosenberg, N. A. & Donnelly, P. (2000), Association mapping in structured populations, Am J Hum
Genet 67(1), 170-181.
Stefansson, H. et al. (2003), Association of neuregulin 1 with schizophrenia confirmed in a Scottish population, Am J Hum Genet
72(1), 83-87.
Stephens JC et al. (2001) Haplotype variation and linkage disequilibrium in 313 human genes. Science.;293(5529):489-93.
Strachan, T. & Read, A. P. (2003) Human Molecular Genetics 3, BIOS Scientific Publishers Ltd, Wiley, New York.
Spielman R S and W J Ewens (1996) The TDT and other family-basedtests for linkage disquilibrium and association. Am. J. Hum.
Gen. 59:983-989
The International HapMap Consortium (2003) The International HapMap Project. Nature 426, 789-795.
Weiss, KM and Clark, AG (2002) Linkage disequilibrium and the mapping of complex human traits. Trends in Genetics 18:19-24.
Pritchard, J and M. Przeworski (2000) Linkage Disequilibrium in Humans: Models and Data AJHG 69.1-14.
Pritchard, JK et al.(2000) “Association Mapping in Structured Populations” Am.J.Hum.Genet. 67.170-181 .
Pritchard and Cox (2002) “The allelic architecture of human disease genes: common disease-common variant … or not” Human
Molecular Genetics 11.20.2417-2Rannala, B and JP Reeve (2001) High-Resolution Multipoint Linkage-Disequilibrium Mapping in
the Context of a Human Genome Sequence AMJHG 69.159-178.
R S Spielman and W J Ewens (1996) The TDT and other family-basedtests for linkage disquilibrium and association. Am. J. Hum.
Gen. 59:983-989
Tabor, Risch and Myers (2002) Candidate-gene approaches for studying complex genetic traits: practical considerations Nature
Reviews Genetics 3.May.1-7
Terwilliger,JD et al(2002) A bias-ed assessement of the use of SNPs in human complex traits. Curr.Opin. Genetics & Development
12.726-34
Weiss,K and Terwilliger, J (2000) “How many diseases does it take to map a disease with SNPs” Nature Genetics vol. 26 Oct.
Books & Www-sites
Books
Encyclopedia of the Human Genome (2003) Nature Publishing Group
Liu, . J(2001) “Monte Carlo Strategies in Scientific Computation” Springer Verlag
Ott, J.(1999) Analysis of Human Genetic Linkage 3rd edition Publisher: John Hopkins
Strachan & Read (2004)
Human Molecular Genetics III
Publisher: Biosciences
Weiss,K.(1993) “Genetic Variation and Human Disease” Cambridge University Press.
Web-sites
www.stats.ox.ac.uk/mcvean
Jeff Reeve and Bruce Rannala A multipoint linkage disequilibrium disease mapping program (DMLE+) that allows genotype data to be used
directly and allows estimation of allele ages.
http://dmle.org/
Liu, J.S., Sabatti, C., Teng, J., Keats, B.J.B. and N. Risch (Version upgraded by Xin Lu, June/9/2002) This is the software for the Bayesian
haplotype analysis method developed by Liu, J.S., Sabatti, C., Teng, J., Keats, B.J.B. and N. Risch in article Bayesian Analysis of Haplogypes
for Linkage Disequilibrium Mapping. Genome Research 11:1716, 2001
http://www.people.fas.harvard.edu/~junliu/TechRept/03folder/bladev2.tar
J. N. Madsen, M.H. Schierup, C. Storm, and L. Schauser, T. Mailund CoaSim is a tool for simulating the coalescent process with recombination
and geneconversion under the assumption of exponential population growth
http://www.birc.dk/Software/CoaSim/