TA extra office hours this week: Reminders: Molecular evolution
advertisement

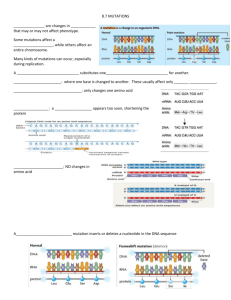
Molecular evolution Reminders: • First writing assignment due in class on Wednesday • Exam 2 next Monday, Nov 3 TA extra office hours this week: Early (1960s) analyses of amino acid variation among different species (Kimura; Zuckerkandl & Pauling) e.g., Cytochrome c Hemoglobin human horse bird fish Kyra: Wed (10/29), 11 am - Noon Nic: Fri (10/31), 9-10 am Pu: Fri (10.31), 10-11 am Cytochrome c TA review session (Katie and Pu): 2:00 - 3:00 Friday, McMillan 149 Amount of aa differences between species ~correspond to the length of time since species diverged from a common ancestor. e.g., horses and humans, good fossil data on time of divergence Suggested constant, steady rate of amino acid substitution Molecular clock: constant, steady rate of change at the molecular level (amino acids, DNA sequences). e.g., for a hypothetical protein sequenced in several species Humans and horses last shared a common ancestor ~45 MY ago. Differ at 6 out of 100 aa sites reptile-fish Molecular clock: constant, steady rate of change at the molecular level (amino acids, DNA sequences). e.g., for a hypothetical protein sequenced in several species Humans and horses last shared a common ancestor ~45 MY ago. Differ at 6 out of 100 aa sites Humans and fruitflies last shared a common ancestor ~600 MY ago. Differ at 90 out of 100 aa sites reptile-fish Linear relationship indicates molecular clock Humans and fruitflies last shared a common ancestor ~600 MY ago. Differ at 90 out of 100 aa sites 1 Implications of molecular clock observations… • Inconsistent with evolution by natural selection: — Would expect far fewer changes (since the protein’s function didn’t change — e.g., cytochrome c) — Would expect changes to be episodic, not steady: associated with periods of NS (e.g, environmental change, rapid speciation, etc.) • Suggested that most mutations that arise and go to fixation do so by genetic drift: selectively neutral Neutral Theory of Evolution Most mutations that arise and go to fixation are neutral with respect to natural selection; fixation by genetic drift. This doesn’t mean that natural selection isn’t acting on these genes: — We assume that most mutations that affect protein function are deleterious and immediately selected against. Many mutations are quickly eliminated by natural selection (not observed). — The Neutral Theory focuses on mutations that arise and go to fixation Revolutionary at the time, since it had been assumed that natural selection was the primary mechanism of evolution The proportion of a gene showing neutral evolution depends on proportion of sites under functional constraint (where mutations will likely be deleterious). — Very few mutations are driven to fixation through natural selection Molecular clock varies by protein: depends on functional constraint e.g., functional constraint is lower for a gene that is mostly introns e.g., functional constraint is lower for a gene where amino acid changes are less likely to disrupt protein function Variation in functional constraint within and among genes: — pseudogenes (no expressed protein): no functional constraints — noncoding regions (e.g., introns): very few functional constraints — 3rd codon position (often synonymous): some functional constraints —1st position (nonsynonymous): high functional constraints Cyt c is under high functional constraint: a low proportion of mutations evolve neutrally Molecular clock will vary for different genes/proteins, gene regions 2 Level of functional constraint varies within a gene Degree of functional constraint on non-synonymous varies depending on gene function: Rates of nucleotide substitition between humans and mice/rats Influenza A strains with different degrees of divergence Gene Lower proportion of nonsynonymous sites evolve neutrally: slower molecular clock Non Synonymous* Synonymous* Histone 3 0.00 6.38 Actin α 0.01 3.13 Thyrotropin 0.33 4.66 Immunoglobulin Ig VH 1.07 5.66 Interleukin I 1.42 4.60 Interferon γ 3.06 5.50 (*avg # substitutions per site per billion years) For sites that are not under functional constraint… Evolution purely by genetic drift: u = rate of mutation (per gamete per generation) For a diploid population of size Ne, the number of new mutations per generation = 2Neu Rate of fixation at neutral sites equals the mutation rate (u)… Implications: (2Neu)(1/2Ne) = u 1. Does not depend on population size! Large N: more mutants, but lower likelihood that any one is fixed Recall: probability of fixation of an allele equals its frequency For a newly arisen allele, this is 1/(2Ne) (e.g., for N=5, 2N= 10, p=1/10 = 0.1) So the number of mutations that arise per generation that eventually get fixed is (2Neu)(1/[2Ne]) = u Rate of fixation equals the mutation rate = u 3 Rate of fixation at neutral sites equals the mutation rate (u)… Implications: (2Neu)(1/2Ne) = u Rate of fixation at neutral sites equals the mutation rate (u)… This would explain the molecular clock too, right?… 1. Does not depend on population size! Large N: more mutants, but lower likelihood that any one is fixed 2. Often expect to find variation (polymorphism) at a site: Recall: time to fixation by drift: 4Ne generations (don’t need to know derivation) e.g., Ne = 1000 4000 generations to fixation of 1 new mutation… If u = 1 X 10-9 mutations per gamete per generation, And if we’re looking at a 1000 bp gene, Then over a period of 4000 generations we expect 8 new mutations: Higher proportion of neutrally evolving sites. Lower proportion of neutrally evolving sites. (1 X 10-9)(2000 gametes)(4000 generations)(1000 bp) = 8 mutations 2Ne Variation! (=No fixation) Tomoko Ohta saves the day: she proposes Nearly Neutral Model Wait! We have a problem… Rate of fixation equals the mutation rate = u u = rate of mutation per gamete per generation NOT per absolute time But the molecular clock seems to follow absolute time… Even though species vary widely in generation time Why aren’t more mutations accumulating in lineages with short generation time?? If we allow for the possibility that most mutations are slightly deleterious instead of strictly neutral, then the probability of drifting to fixation will depend on population size: Small N = drift overrides weak selection, so most mutations are evolving as if neutral: ‘effectively neutral’ Large N = drift is weak, so most mutations not neutral and are selected against Mutations are effectively neutral (i.e., s=0) for s < 1 2Ne So why does this save the day?… Absolute time 4 Because species with short generation times tend to have larger populations: Detecting natural selection on DNA sequences Using (nearly) neutral evolution as the null hypothesis: look for deviations from neutral expectations In other words, look for patterns of evolution that don’t fit evolution by genetic drift alone. Short generation time many mutations per year but fewer mutations effectively neutral s< 1 2Ne Long generation time few mutations per year but many mutations are effectively neutral So the difference in generation time is balanced out by N Method 1. dN/dS ratios Nature Vol. 335, 8 September 1988 dN = rate of non-synonymous substitutions per site (measured as # nonsynonymous polymorphisms) dS = rate of synonymous substitutions per site (measured as # synonymous polymorphisms) MHC: major histocompatibility complex ARS receptor: antigen recognition site Antigen recognition site (57 codons) dN/dS < 1 aa replacements largely deleterious (e.g., normal functioning gene) dN/dS = 1 aa replacements are neutral (e.g., pseudogene, no functional constraint) dN/dS > 1 aa replacements are advantageous and are favored by selection dN 13.3 per 100 sites ds 3.5 per 100 sites dN/dS: 3.8 Selection favors amino acid changes in ARS (new alleles favored) MHC protein Remaining codons in Exons 2 and 3 1.6 per 100 sites 2.5 per 100 sites dN/dS: 0.64 Typical for nonsynonymous vs synonymous sites (functional constraint) 5 Nature Genetics 25, 410 - 413 (2000) Adaptive evolution of the tumour suppressor BRCA1 in humans and chimpanzees Gavin A. Huttley, Simon Easteal, Melissa C. Southey, Andrea Tesoriero, Graham G. Giles, Margaret R.E. McCredie, John L. Hopper & Deon J. Venter One problem with using dN/dS ratios to infer selection: it’s extremely conservative if you average across the entire gene e.g., 363 genes examined in mice/rats: only one with dN/dS > 1 Most useful for comparing different domains within a protein: e.g., abalone lysin protein domains Recall: sperm competition to penetrate egg rapid evolution. Suggests selection dN/dS ratios Exposed protein regions are evolving rapidly: Black: dN/dS >3.0 Gray: dN/dS <1.0 White: dN/dS ~1.0 dN/dS >3.0 MK test… Detecting natural selection on DNA sequences… Adh (alcohol dehydrogenase) gene in Drosophila species Method 2. McDonald-Kreitman (MK) test: Neutral theory: # polymorphic sites within species should be directly proportional to number of # differences fixed between species Might expect selection on Adh for alcohol tolerance in species whose larvae live in fermenting fruit Within species: look at polymorphic sites (# nonsynon., # synon.) Between species: look at fixed differences (# nonsynon., #synon.) MK test reveals selection where dN/dS alone does not: Within-species polymorphism Differences fixed between species Nonsynonymous: 2 8 Synonymous: 10 40 These ratios should be equal under neutral evolution An excess of non-synonymous fixed differences would indicate selection driving amino acid change. Within-species polymorphism Differences fixed between species Nonsynonymous: 2 7 Synonymous: 42 17 By itself, dN/dS just looks like normal functional constraint 0.048 (=dN/dS) Excess nonsynonymous fixed differences 0.412 6 Detecting natural selection on DNA sequences… Method 3: test for excess of old or new mutations… With no selection (genetic drift acting alone), we expect alleles (=haplotypes) to continuously arise and to go extinct Haplotype tree with no selection: Haplotype tree with positive selection: Most alleles will be recent descendents of favored allele Expect mixture of closely related alleles (=recently diverged from an ancestral allele), and those that are more distantly related (=older common ancestor allele) * = selectively favored mutation With directional selection for an advantageous mutation (= Positive Selection), we expect fewer older alleles than with neutrality time Method 3: test for whether there is an excess of old or new polymorphisms compared to neutral expectations. Alleles closely related, no long branches: shallow haplotype tree (blue= extinct alleles) (blue= extinct alleles) Method 3: test for excess of old or new mutations… With selection to maintain two or more allele classes (=balancing selection), find maintenance of allele lineages that would otherwise go extinct Haplotype tree with balancing selection: Some alleles will be very distantly related: long branches e.g., heterozygote advantage negative freq-dep. selection diversifying selection (blue= extinct alleles) 7