Automated Cycle Sequencing - Albert Einstein College of Medicine
advertisement

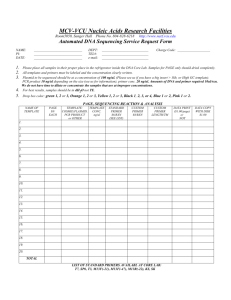
SAMPLE INFORMATION FORM DNA SEQUENCING FACILITY Albert Einstein College of Medicine 713 Ullmann, x2657 Facility e-mail:SEQLAB@AECOM.YU.EDU David Reynolds: DREYNOLD@AECOM.YU.EDU PROVIDE 2.0 l OF DNA TEMPLATE DISSOLVED IN H2O OR EB BUFFER AND 3.2 l OF PRIMER DISSOLVED IN H2O Investigator information Principal Investigator Department MIXED TOGETHER FOR A TOTAL VOLUME OF 5.2 l IN A 0.2 ML MICROAMP TUBE AT THE CONCENTRATIONS LISTED BELOW Date Submitted Contact Person Telephone Number E-Mail Address Rm/Bldg Address Grant # 9 - 526 - - 431 PURIFICATION: DNA for the automated sequencing facility should be prepared with a QIAGEN spin column and dissolved in H2O or EB buffer (do NOT dissolve in TE buffer). This will assure the most consistent and highest quality sequence data. QUANTIFICATION: You must quantify your DNA by running 1ul on a gel adjacent to a DNA concentration standard. In addition, we recommend you take an OD reading of your sample. Label on Tube 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 The purity and concentration of both the template and the primer is critical for automated sequencing. Template Type Size (kb) DNA TEMPLATE: Plasmid (double-stranded DNA) Cosmid DNA or DNA BAC DNA PCR product PRIMER: For Plasmid or PCR product For cosmid DNA or DNA For BAC DNA Concentration Volume 200 ng/l 600 ng/l 1200 ng/l 15 - 90 ng/l 2.0 l 2.0 l 2.0 l 2.0 l 1 pmol/l 5 pmol/l 15 pmol/l 3.2 l 3.2 l 3.2 l SEQUENCE RESULTS: The facility provides a printout of the sequence chromatogram and an e-mail text file. Special Request: ___________________________________ Primer Name Primer Tm Notes