Lab Optional: Hyperspectral Image Analysis with ENVI
advertisement

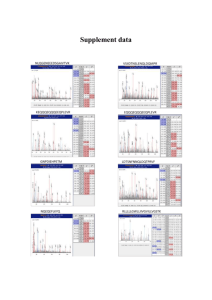
GISC 7365: Remote Sensing Digital Image Processing Instructor: Dr. Fang Qiu Optional Lab Hyperspectral Analysis Objective: This lab is designed to introduce you to the concepts and procedures for analyzing imaging spectrometer data or hyperspectral images. We will use 1995 Airborne Visible/Infrared Imaging Spectrometer (AVIRIS) data from Cuprite, Nevada, to investigate the hyperspectral data and the associated spectral mapping techniques. Data: Data to be used for this lab is in T:\Qiu\RSDIP\Data\lab10 Quickview Open the 1995 calibrated (Effort-Corrected) Data Start ENVI. Open the image file CUP95EFF.INT (Integer image file) from the data folder. This is the 1995 AVIRIS ATREM-calibrated apparent reflectance data with the EFFORT correction applied. Load band 193 as a gray scale image in the new image display by clicking on the Gray Scale radio button, selecting the band, selecting New Display, and clicking on the Load Band button. Part I: MNF Transform Use the MNF Rotation (minimum noise fraction) transforms to determine the inherent dimensionality of image data, to segregate noise in the data, and to reduce the computational requirements for subsequent processing. Both the resulting eigenvalues and the MNF images (eigenimages) are used to evaluate the dimensionality of the data. Generate MNF Image In the main menu, choose Spectral->MNF Rotation->Forward MNF>Estimate Noise Statistics From Data In the box of Select Input Files, choose CUP95EFF.INT from the data folder and then press OK In the dialogue, Click Choose to enter the Output name of the first two files if you want. Then check the Output Results to File and enter the output name as cup95eff_MNF in your workspace. Set “Select Subset from Eigenvalue” to YES, and then press OK. From the result, you can see Eigenvalue of each MNF band and cumulative percentage of variance. MNF bands whose Eigenvalue larger than 1 contain more spatial information and the bands whose Eigenvalue less than 1 contains primarily noise. We can choose the first 15 bands to output. And then click OK. Save the output Eigenvalues graph as an image. From the available bands list, display each of MNF bands to examine their spatial coherence. Compare MNF Images Load some MNF bands. Use ToolsProfilesZ Profile and ToolsLinkLink Displays to compare MNF spectra with the Apparent Reflectance spectra from the EFFORT image. Examine the relationships between them. Use Tools2D Scatter Plots in the Main Image window. Be sure to examine the high variance (low band number) MNF bands. Also be sure to examine at least one scatter plot of a low variance (high band number) MNF bands (change the bands by selecting OptionsChange Bands in the Scatter Plot window). Notice the corners (pointed edges) on some MNF scatter plots (fig.1) Fig.1 MNF pointed edge Fig.2 MNF fuzzy edge Use linked windows and Z profiles to understand the reflectance spectra of the MNF corner pixels. Look for areas where the MNF data stops being “pointy” and begins being “fuzzy” (fig.2). How do you explain these patterns? Homework: Make a map to display two of the resulted MNF bands (one with low and one with high band number). Also enclose the screen shot of the eigenvalues and cumulative percentage graph. Part II Pixel Purity Index In this part, you will examine the role of purity of pixels. The Pixel-Purity-Index (PPI) is a means of finding the most “spectrally pure,” or extreme, pixels in multispectral and hyperspectral images. The most spectrally pure pixels typically correspond to mixing endmembers. Generate PPI In the main menu, choose Spectral->Pixel Purity Index->New Output Band. You have the choice of a disk-based PPI method and a FAST PPI method. The FAST PPI places the image data into memory and performs the computations in memory, which is much faster than the disk-based PPI method, but requires adequate memory space. In the dialogue box, select CUP95EFF.INT as input file. Press OK. Set threshold factor as 2. A threshold of 2 marks all pixels greater than two digital numbers (DN) from the extreme pixels (both high and low) as being extreme. This threshold selects the pixels on the ends of the projected vector. Choose Output Results to File, and enter the output name as cup95eff_PPI in your workspace. Patiently wait for the computer to finish the job. From the available bands list, we can see the PPI image list there. Choose it to open display it as a gray scale image. Brighter pixels represent more spectrally extreme finds (hits) and indicate pixels that are more spectrally pure. Darker pixels are less spectrally pure. Open both the MNF (cup95mnf) and Effort apparent reflectance (CUP95EFF.INT) files and load bands from each of these two into two new displays. The PPI image is the result of several thousand iterations of the PPI algorithm on the MNF data. Link all windows using ToolsLinkLink Displays and create Z-profiles in the MNF and reflectance image displays. In this configuration, you can examine the spectral profiles of certain pixels as they are selected in the PPI display. Select or ToolsCursor Location/Value from the Display menu to determine the range of values present in the image. Select ToolsRegion of InterestBand Threshold to ROI to create an ROI containing only the pixels with high PPI values. Use a minimum threshold. For example, a minimum of 10 will include all of those pixels with PPI values greater than 10 in the ROI. In the ROI tool window, select File Save ROIs … to your workspace. Homework: Make a map to display the resulting PPI image and threshold ROIs. Part III Spectral Angle Mapper Classification The Spectral Angle Mapper (SAM) is an automated method for comparing image spectra to individual spectra or a spectral library. SAM assumes that the data have been reduced to apparent reflectance (true reflectance multiplied by some unknown gain factor controlled by topography and shadows). The algorithm determines the similarity between two spectra by calculating the “spectral angle” between them, treating them as vectors in a space with dimensionality equal to the number of bands. Select Image Endmembers Open the 1995 AVIRIS ATREM-calibrated apparent reflectance data file, CUP95EFF.INT. From the ENVI main menu, SpectralMapping MethodsSpectral Angle Mapper to start the SAM endmember selection process. In the Classification Input File dialog, select the file CUP95EFF.INT as the input file and click OK. In the Endmember Collection: SAM dialog, select Importfrom ASCII file. Select the file CUP95_EM.ASC from data folder and click Open. In the Input ASCII File dialog, hold down the Ctrl key and deselect the spectra Dark/black, Bright/playa, Silica? (Dark), and Alunite (2.18) in the Select Y Axis Columns list. This will leave you with the mean spectra Zeolite, Calcite, Alunite (2.16), Kaolinite, Illite/Muscovite, Silica (Bright), and Buddingtonite. Click OK to load all of the endmember spectra into the Endmember Collection:SAM dialog. From the Endmember Collection:SAM dialog menu, click Plot button to plot the endmember spectra. From the Endmember Collection Spectra window, select OptionsStack Plots for improved comparison of spectral features, then right click in the plot window and select Plot Key from the shortcut menu to display the legend. Click Apply on the Endmember Collection:SAM dialog to start the classification, enter Output Class File Name and Output Rule Filename in the Spectral Angle Mapper Parameters dialog, and click OK. Examine SAM Open the SAM classification image by selecting FileOpen Image File and choosing resulting SAM classification image as the input file name. The classification image is one band with coded values for each class (for example, alunite is coded as “1”). When opened, the classified image will appear in the Available Bands List dialog. In the Available Bands List dialog, ensure that the Gray Scale radio button is selected. Click DisplayNew Display, select the SAM classification image, then click Load Band. In the Available Bands List dialog, click DisplayNew Display. Display each of the rule images in Gray Scale. Homework: Make a map of the resulting classification map and two of the rule images. Part IV Linear Spectral Unmixing Use Linear Spectral Unmixing to determine the relative abundances of materials that are depicted in multi- or hyper-spectral imagery based on the materials’ spectral characteristics. The reflectance at each pixel of the image is assumed to be a linear combination of the reflectance of each material (or endmember) present within the pixel. So given the resulting spectrum (the input data) and the endmember spectra, the linear unmixing is solving for the abundance values of each endmember for every pixel. The number of endmembers must be less than the number of spectral bands and all of the endmembers in the image must be used. Spectral unmixing results are highly dependent on the input endmembers and changing the endmembers changes the results. Select SpectralMapping MethodsLinear Spectral Unmixing. When the Unmixing Input File dialog appears, set input file to CUP95EFF.INT. Click OK to continue. The Unmixing Endmember Collection dialog appears. Use the dialog to select endmember spectra from a variety of sources, in this case ASCII spectra (CUP95_EM.ASC). Spectra from all sources are automatically resampled to match the wavelengths of the multiband image being unmixed. In the Input ASCII File dialog, hold down the Ctrl key and deselect the spectra Dark/black, Bright/playa, Silica? (Dark), and Alunite (2.18) in the Select Y Axis Columns list. This will leave you with the mean spectra Zeolite, Calcite, Alunite (2.16), Kaolinite, Illite/Muscovite, Silica (Bright), and Buddingtonite. Click OK to load all of the endmember spectra into the Endmember Collection Dialog. Click Select All, click Apply at the bottom of the Endmember Collection: Unmixing dialog.The Unmixing Parameters dialog appears. Select output to File and enter output filename Click OK to begin the spectral unmixing. Open and Display the Unmixing Results The results of spectral unmixing appear as a series of gray-scale images, one for each endmember, plus a root-mean-square (RMS) error image. Higher abundances (and higher errors for the RMS error image) are represented by brighter pixels (larger floating-point numbers). Open the resulting file LSU classification image. Load the Kaolinite abundance image by clicking on the Kaolinite band name and then Load Band in the Available Bands List dialog. Brighter areas correspond to higher abundances. Load the other fractional abundance images into one or more displays and compare the distribution of endmembers. Select ToolsCursor Location/Value and investigate the values at specific pixels to determine abundance percentage. View the RMS error image to help determine areas of missing or incorrect endmembers. Homework Make a map of the resulting RMS error image and two of the abundance images.