Intracellular Protein Degradation
advertisement

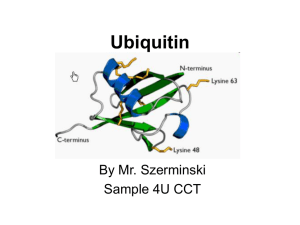
Intracellular Protein DegradationThe lysosome and Ubiquitin Proteasome System Scott Wilson Department Neurobiology 5-5573 Wilson@nrc.uab.edu Outline Sites of proteolysis Gastrointestinal tract Circulatory system Intracellular proteolysis Lysosome Biogenesis and function Degradation of extracellular material Degradation of intracelluar components by autophagy Ubiquitin proteasome pathway Components Ubiquitin and UBLs Ubiquitin conjugating enzymes Ubiquitin deconjugating enzymes The proteasome- generation and activity Gastrointestinal tract Destruction of antigenicity Controlled but no specificity- everything that enters gut is proteolyzed Production of energy Remember that destruction of proteins is an energy producing process (exergonic) Circulatory system Blood coagulation Conversion of prothrombin to thrombin which converts fibrinogen to fibrin and a blood clot is formed. Process is highly controlled (1-antitrypsin deficiency) The question: Is there turnover of cellular constituents? Or is food intact a function primarily for energyproviding (fuel for a car), that is independent from the structural and functional proteins of the body? • Studies on -galactosidase in E. coli indicated that there was no conclusive evidence that proteins within cells are in a dynamic state and that they are likely to be stable and static • Without metabolic labels (ex. 35S cysteine or 3H leucine) the problem of determining protein stability was not approachable How do you “tag” proteins to study protein dynamics? 1939 Rittenburg and Urey succeeded in generating radiolabeled Nitrogen (15N) Schoenheimer found that following administration of 15N-labeled tyrosine to rats, they found that only ~50% of the label was found in excretions. Where was the rest? The label was found incorporated in body proteins! Therefore the proteins of the body are in a dynamic state of synthesis and degradation! It is thought that we are degrading and resynthesizing ~3-5% of our cellular proteins daily. Paradigm that cellular processes are controlled mainly by only transcription and translation must be changed. Why are proteins degraded? Quality control Proteins become denature/misfolded/damaged Elevated temperatures (37°C) Proteins being synthesized are folded incorrectly Regulation of biological pathways Cell cycle Receptor mediated endocytosis Synaptic remodeling Now that we know proteins are in a “dynamic state” in cells…. How are proteins degraded within cells? Is protein degradation regulated? Selective? Compartmentalized? The discovery of the lysosome De Duve discover the lysosome in the 1950’s Vacuolar structure that contains hydrolytic enzymes that are optimal at acidic pH. Latency of of enzymatic activity- researcher found that hydrolyase fractionated from rat liver were more active after they were stored in the refrigerator for several days? The latency was due to the slow breakdown of the lysosomal membrane which protected the cells from the destructive forces of the acid hydrolyases. This compartmentalization of the peptidases by a membrane protects cellular components from inappropriate degradation. Generation of a functional lysosome Lysosomal proteases belong to the aspartic, cysteine, or serine proteinase families of hydrolytic enzymes. contain about 40 types of hydrolytic enzymes, including proteases, nucleases, glycosidases, lipases, phospholipases, phosphatases, and sulfatases. All are acid hydrolyase that have optimal activity at pH 5.0 Sorting acid hydrolyases to the lysosome is accomplished by post-translation modification Soluble lysosomal enzymes are synthesized as N-glycoslyated precursors in the ER and trafficked to the Golgi mannose 6-phosphate (M6P) groups are added to the hydrolyases The M6P groups are recognized by transmembrane M6P receptor proteins, which are present in the trans Golgi network M6P receptors release hydrolyases when pH is below 6.0 and the M6P is removed Lysosomes use an H+ ATPase pump in the membrane to generate acidic pH Overview of lysosomal trafficking Proteases in the lysosome Cysteine protease- cathepsins A, B Aspartate protease- cathepsin D Zinc protease-? Activation of protease by removal of inhibitory segment- conversion of proprotein to protein Pathways into the Lysosomal/vacuolar System 1 3 2 4 4 Model of the mechanism for multivesicular endosome formation How do proteins get into the lysosome for degradation? Microautophagy- cytoplasm is segregated into membrane -bound compartments and are then fused to lysosome Maroautophagy- entire organelles such as mitochondria, ER and other large cytoplamic entities are engulfed and then fused with the lysosome Autophagy pathway Problems that still remain Proteins vary greatly in their stability - from minutes to days! Rates of protein degradation of specific proteins changes with physiological conditions (nutrients and hormones) How could this happen by microautophagy Lysosomal inhibitors have differential affects on different populations of protein If lysosomal proteases degrade proteins in an exergonic manner, how could you explain evidence that the proteolytic machinery required energy? Still more data suggesting another pathway for degradation of intracellular proteins Poole et al were studying the mode of action anti-malaria drugs Chloroquine and other lysosomotropic (weak bases) block the activity of lysosomal proteases by neutralizing the low pH of the lysosome. Treat macrophages labeled with 3H-leucine with chloroquine and then feed them protein extracts that were labeled with 14C-leucine This allowed them to monitor the stability of phagocytosed extracellular and intracellular proteins when the lysosome is blocked What did they find? Lysosomotropic drugs only affected the stability of the engulfed extracellular proteins and not the intracellular proteins. This indicated that there must be a second pathway for the degradation of intracellular proteins and that the lysosome was the primary site of degradation of internalized extracellular proteins The search for a new proteolytic pathway The new pathway must explain several things Requirement for metabolic energy ATP depletion inhibits proteolysis Why do you need ATP? Need phosphorylation of substrates or enzymes? Remember proteolysis is exergonic Differential stability of intracellular proteins Example- RNA polymerase I t1/2= 1.5 hrs RNA polymerase II t1/2= 12 hrs How stability of proteins can change under different environmental conditions Cell-free proteolytic system Rabbit reticulocyte lysates Made from red blood cells (terminally differentiated and do not have lysosomes) New that for different hemoglobinopathies, the blood cells attempt to rid themselves of abnormal hemoglobins and therefore must have a proteolytic system that was not lysosomal based. Found that reticulate lysates were capable of degrading proteins in an ATP dependent manner A new paradigm for proteolysis Biochemical characterization of reticulate lysates Divided the lysates into two fractions (DEAE cellulose, anion exchange resin) Flow thru and high salt eluate Each fraction did not have proteolytic activity on its own. Combination of fraction I and II reconstituted proteolysis Previous work indicated that only a substrate and protease were need for degradation. This was very important in that it suggested that there was not a single protease that mediated degradation. This new system need a substrate, protease and something else Activator? Characterization of fractions I and II Analysis of Fraction I Found that fraction I contained only a single factor that was heat sensitive and required ATP This factor was termed APF-1 for ATPdependent proteolysis factor Critical finding was that APF-1 can be covalently attached to a target substrate APF-1 is shifted to high molecular mass compounds following addition of ATP to the fraction I. 125I labeled fractions following gel-filtration chromatography SDS PAGE analysis of samples run on gel-filtration Lane 1- Fraction II + 125I- APF (no ATP) Lane 2- Fraction II + 125I-AFP + ATP Lane 3- Fraction II + 125I-AFP + ATP + unlabeled lysosome as substrate Lane 4 & 5 - Increasing conc of lysosome Lane 6- Fraction II + 125I-lysosome (no ATP) + unlabeled APF Lane 7- Same as lane 6 + ATP These experiments demonstrate that APF is covalently attached to substrate (explains the requirement of ATP) Multiple APF-1’s can be added to a substrate What is APF-1 ? Amino acid analysis and its known molecular mass indicated that APF-1 is ubiquitin. Ubiquitin is a 76 aa protein found only in eukaryotes The covalent attachment of ubiquitin to a substrate stimulates its proteolysis (but by what?) Ubiquitin is covalently attached to a substrate by is C-terminal glycine to the -NH2 group of an internal lysine of the substrate Studies of fraction II defined the ubiquitin conjugation machinery Substrate recognition N-end rule: On average, a protein's half-life correlates with its N-terminal residue. Proteins with N-terminal Met, Ser, Ala, Thr, or Gly have half lives greater than 20 hours. ・Proteins with N-terminal Phe, Leu, Asp, Lys, or Arg have half lives of 3 min or less. What about the protease? Previous studies demonstrated that the activity of the protease was ATP dependent (not just ubiquitination requires ATP) What is it composed of? Where is it located? How is it selective toward ubiquitinated proteins? Why does it need ATP? Structure of the 26S proteasome Tanaka et al discovered a highmolecular mass protease that degraded ubiquitinated lysozyme but not untagged lysozyme Required ATP for activity Protease was later called the 26S proteasome Similar multi-subunit proteases found in prokaryotes Subunits of the 26S proteasome 19S regulatory particlecomposed of approximately 20 different proteins 20S core particlecomposed of 14 different subunits (1-7 and 1-7) 19S Regulatory particle (RP) Recognition and binding of ubiquitinated proteins Unfolding of ubiquitinated substrate to enter 20S mediated by AAA ATPases (ATP dependent) Removal of ubiquitin side chains to allow entry into 20S ( lumen ~1.3 nm) by deubiquitinating enzymes Activation/opening of 20S lumen 20S Core Particle (CP) Contains the endopeptidase activity The alpha subunits function is to control the opening and closing of the 20S gate (interacts with 19S) The beta subunits 1, 2 and 5 contain the endopeptidase activity of the proteasome. Proteins are not degraded into amino acids but into short peptides ( very important for immune surveillance). The UPS is enormous! The genes of the UPS constitutes ~5% of the genome E1’s- 1-2 activating enzymes E2’s- 10-20 conjugating enzymes E3’s- 500-800 ubiquitin ligase- drives specificity DUBs- 100 ubiquitin specific proteases- regulators of pathway Pathways controlled by regulated proteolysis Diseases of the lysosome and UPS pathways Lysosomal Neimann Pick Disease- ataxia, brain degeneration and spasticity. Krabbe Disease- hypertonia, seizures, deafness and paralysis Tay-Sachs Disease- cognitive disorder, deafness, paralysis Ubiquitin-dependent regulation of Ubp6 Hanna, J et al Cell 127:99-111 2006 Ubiquitin-dependent regulation of Ubp6 levels Hanna, J et al Cell 127:99-111 2006 Altered proteasome content in yeast expressing Ubp6C118A Hanna, J et al Cell 127:99-111 2006 Cellular responses to ubiquitin deficiency and proteasomal stress Hanna, J et al Cell 127:99-111 2006 Proteasome inhibition increases Usp14 ubiquitin-hydrolase activity Usp14 Uch37 Borodovsky, A et al EMBO J. 20:5187-96 2001 The proteasomal DUB Usp14 impairs protein degradation Lee, BH et al Nature 467:179-84 2010 Decrease steady-state levels of aggregate prone proteins in the absence of Usp14 Lee, BH et al Nature 467:179-84 2010 Proteasome activity can be modulated by Uch37, Rpn11 and Usp14 Proteasomal DUB functions in yeast 1) Rpn11- cleaves near base of chain to remove ubiquitin chains “en bloc” 2) Usp14 - recycling of residual ubiquitin conjugates from proteins entering the proteasome, ubiquitin chain editing regulation of proteasome activity 3) Uch37- ubiquitin chain editing Mouse models 1- Rpn11- unknown but likely lethal 2- Usp14- KO embryonic lethal (E14) hypomorphic allele viable 3- Uch37 unknown Ubiquitin is not the only small peptide to be covalently attached to proteins and or lipids SUMO 1/2 Nedd8 ISG15 ATG8 FAT10 Not thought to target proteins for destruction Each is thought to have its own conjugation and deconjugation system