DNA Isolation and Genetic Transformation page 66
advertisement

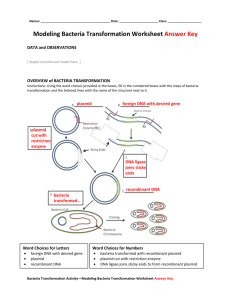
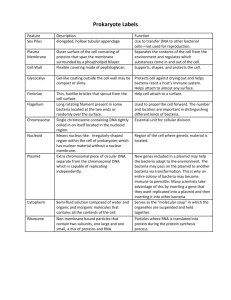
DNA Isolation and Genetic Transformation Objectives 1. Isolate DNA from cells of plant tissue (onion). 2. Use a bacterial plasmid to insert a gene into a bacterium (transformation). Introduction In one sense, people have been manipulating the genotypes of other organisms ever since they began the domestication of animals and plants. Up until very recently, however, these manipulations were purely by selection. That is, we chose plants and animals that had desirable characteristics, and we encouraged them to reproduce. By careful selection, it was possible to develop populations that differed dramatically from their wild ancestors, such as corn and cows. Selective breeding is limited, however, because it does not directly manipulate or "engineer" the genetic makeup of organisms. Over the past few decades, however, biologists have developed methods for transferring genes from one organism to another. An important step took place in 1928, when Franklin Griffith was experimenting with two strains of the bacterium Streptococcus pneumoniae. One of these strains killed mice if it was injected into them, while the other strain was harmless. If Griffith killed the harmful bacteria with heat, the dead bacteria were harmless when injected. But, if he mixed live bacteria of the harmless strain, with heat-killed bacteria of the harmful strain, the injected mice died. Somehow, the dead bacteria transformed the live ones, and whatever factor was involved was not destroyed easily by heat. Griffith called this phenomenon transformation. We know now that Griffith's transforming factor was DNA. Moderate heat denatures proteins, without destroying DNA. Grif- page 66 fith's bacteria were able to recover functional (and deadly) genes from the debris of their heat-killed brethren. As biologists learned more about the function of the DNA, and the universality of the genetic code and machinery, they realized that it would also be possible to transfer genes among unrelated organisms. In 1980, biologists first succeeded in placing a human gene into bacteria, which then manufactured a human antiviral protein known as interferon. From these early efforts, genetic engineering has grown to become a multibillion dollar growth industry, generally known as biotechnology, and come to be a critically important part of medicine, industry, and agriculture. In lab today we will perform two procedures that will give you a feeling for the sorts of manipulations involved in working with DNA and placing genes into cells. The first of these procedures is the isolation of DNA from plant tissue. The second is a bacterial transformation, which is similar in some ways to the pioneering work done by Griffith. 1- Isolation of DNA from onion We could extract DNA from any living tissue, but it is convenient to use a plant, because it won't object when we put it into a blender. Several obstacles must be dealt with, however. The DNA is within the nuclear membrane, the cell membrane, and the cell walls. The cells themselves are massed into compact tissues. We will homogenize an onion in a blender, in order to disperse the cells and mechanically break the cell walls. Knowing what you do about onions, you don't want to get too close to this operation! (Incidentally, the "tear gas" that onions release when cut is called synpropanethial-S-oxide. It is produced when enzymes called allinases are released from the cells and act on compounds called amino acid sulfoxides. This reaction proba- DNA Isolation and Genetic Transformation bly protects the onion from being eaten by some organisms). We will homogenize the onion in an "extraction solution" that contains the laundry liquid "Woolite" and NaCl. Woolite contains detergents that will dissolve the cell membranes, and also proteolytic enzymes that will break down proteins. These proteolytic enzymes will help to free the DNA from histone proteins and also help to inactivate other lytic enzymes that would break up the DNA. Heating at 60C will further aid these steps- many proteins denature at 60 degrees Celsius, while DNA is stable up to 80 C. The salt will help to stabilize and solubilize the DNA by providing cations to interact with the acid anions of the DNA. After the DNA is in solution, the trick is to separate it from all the other cellular debris in the homogenate. It turns out that this is fairly easy to do. We will create a layer of another solvent (ethanol) on top of the extraction solution. The DNA is not soluble in ethanol, and it will precipitate (come out of solution) at the interface between the ethanol and the extraction solution. The strands of DNA are long enough to wind up on a glass or plastic rod. Procedure 1. The TA will puree 1 medium onion in 100 ml of extraction solution, and distribute 30 ml of homogenate into a 50 ml centrifuge tube for each group. 2. Quickly cap the tube of homogenate and place it the at 60 C water bath for 15 minutes. 3. Remove the tube from the water bath and place it on ice for at least 5 minutes. 4. Use a coffee filter to collect 6 ml of filtrate in the plastic tube provided. page 67 5. Slowly pipette 9 ml of ice-cold 95% ethanol down the side of the tube. Be careful- you do not want the ethanol to mix with the filtrate- you want it to form a layer on top. 6. Allow the tube to sit for undisturbed for 2-3 minutes. After this time you should be able to see strands of DNA precipitating at the interface of the two layers. 7. Use the glass rods provided to "spool" the DNA strands by twisting the rod (not by stirring). 2– Transformation Remember that a gene is the DNA that provides the instructions for making a particular protein. The expression of genes (transcription and translation) results in the phenotypic characteristics of an organism. "Phenotype" can mean any physical or physiological characteristic. Genetic transformation means change in the phenotype that is caused by acquiring a new gene or genes. In this procedure, you will genetically transform a bacterium called Escherichia coli (E. coli for short). This bacterium is the "white rat" of microbiology, and it has been studied intensively by biologists. It is an ideal subject for biotechnology applications, because it grows very rapidly in culture (dividing every 20 minutes in ideal conditions). The phenotypic characteristic that we will alter is the ability of the bacteria to grow in the presence of an antibiotic called ampicillin. Antibiotics, by definition, are drugs that kill bacteria or prevent their growth. Normally, E. coli cannot grow if ampicillin is present. However, we will change that by giving the cells a gene for ampicillin resistance- it codes for an enzyme that breaks DNA Isolation and Genetic Transformation down ampicillin. Bacterial DNA is usually a single continuous loop, unlike the linear DNA of eukaryotic chromosomes. In addition to the big chromosome, bacteria may contain, replicate, and express other smaller loops of DNA, called plasmids. These plasmids can be thought of as little extra chromosomes, carrying a few genes that are not necessarily essential for the cell. In some ways, plasmids are similar to viruses, in that they can sometimes be passed from one bacterial cell to another. This transfer is fairly rare on a "per bacterium" basis, and only a small percentage of the cells will take up plasmids in our experiment. However, with millions of bacteria in a drop of culture, the odds of some of them taking up plasmids become pretty good. We can also facilitate the entry of plasmids into the cells by various treatments. The ability of a cell to take up a plasmid is called "competence". Plasmid pUC18 The DNA that we will use to transform E. coli is a plasmid called pUC18. Plasmid pUC18 is a small loop of DNA, containing 2686 base-pairs, and it includes only a couple of genes. Other genes can be spliced into it, so that it has been used extensively as a way to get genes into E. coli and "clone" them. After a cell takes up a plasmid, it replicates it, and each time the transformed cell divides, it passes copies of the plasmid on to its daughter cells. Remember that bacteria reproduce rapidly, so that any cell can have thousands of progeny in a matter of hours. Any cell that takes up the plasmid can rapidly produce a colony of cells that all have the plasmid. One of the pUC18 genes is for an enzyme called β-lactamase. Possessing this gene causes bacteria to be resistant to the antibi- page 68 otic ampicillin. Bacteria lacking this plasmid will not grow in the presence of this antibiotic. Acquiring this plasmid makes the bacteria able to produce ß-lactamase and able to grow in media that contains ampicillin. Procedure: 1. We will be transferring bacteria into sterile culture media, in tubes and on agar plates. You want to avoid contaminating these media with other bacteria. The instructor will wipe down the bench tops with disinfectant. Wash your hands well before beginning the procedure. The instructor will demonstrate how to transfer the bacteria without contaminating the culture ("sterile technique"). 2. Work in groups of 4. Obtain a sterile microcentrifuge tube (hereafter called a “tube”) to hold the bacterial culture while we "soften it up" a bit. Your instructor may allow you to use the micropipettes, or may designate certain students to use them. Using sterile tips, transfer 50µl of E. coli culture into each tube, and then add 500µl of CaCl2 to each tube. 3. Cap the tube and tap the end with the tip of your index finger to mix the solutions. Then put the tube on ice for about 20 minutes. This treatment improves the competence of the cells (remember what that means?). 4. While you are waiting, obtain 2 more sterile tubes and mark them "+" and "-", for "plus plasmid" and “minus plasmid”. The instructor will add 10µl of solution containing the pUC18 plasmid to the "plus" tube. 5. Transfer 100µl of the competent cells to each of the +/- tubes. DNA Isolation and Genetic Transformation 6. Incubate the +/- tubes on ice for 30 minutes. 7. Transfer the +/- tubes to a 37C water bath for 5 minutes. This heat shock facilitates the uptake of plasmid DNA. 8. Add 750µl of nutrient broth to each of the +/- tubes and incubate at 37C for 30 minutes. This incubation period allows the bacteria time to recover from the CaCl2 treatment and to begin to express the ampicillin-resistance gene on the plasmid. 9. Follow your instructor's directions to inoculate 4 agar plates. You should inoculate 2 plates from each of the 2 tubes- one plate with ampicillin and one without. You will use a micropipette to transfer 250 µl of the mixed bacterial suspension from the tube to the plate, and then use an inoculating loop to spread the bacteria evenly onto the agar surface. It is important to have areas where the culture drop is is spread very thinly, so that you can observe colonies that grow from individual cells. 10. Leave the plates at room temperature until the liquid has been absorbed into the agar (about 10 minutes). 11. Tape the plates shut, then invert them and write your name, lab section the date, and the treatment on each. Incubate the inverted plates at 37C for about 24 hours. 12. After about 24 hours, you will have to check your plates and count the bacterial colonies that have developed. Your instructor will describe how to carry out this procedure. Count the number of visible colonies on each of your plates and record these values in Table 1. page 69 Table 1. Transformation Results Treatment Number of colonies + amp, - plasmid + amp, +plasmid - amp, - plasmid - amp, + plasmid Assignment: Lab Report Your report should focus on the transformation procedure. Follow the lab report format- it wouldn’t hurt to reread the description that appeared earlier. In the introduction of your report, briefly describe the bacterial genome and plasmids, the mechanism of transformation, and the overall rationale of the experiment. End the Introduction with a statement of the predicted outcome. State the Methods in your own words- don’t transcribe the lab manual- and include any important steps added by your instructor. The Results section should have a written description of the outcome as well as a table giving the results. In the Discussion, interpret the results, and also extend your discussion to include information about either antibiotic resistance plasmids or recombinant plasmids. You can find more information in your text and the sources that are linked to the Bio 121 lab website. You will need to do some searching to find information about antibiotic resistance plasmids or recombinant plasmids. Finally, don’t forget the Literature Cited section.