SignalP-NN result: for Humant WNT-1
advertisement

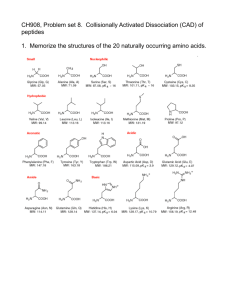
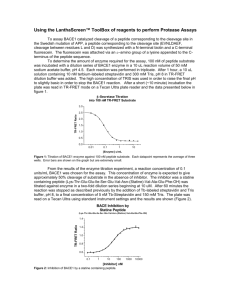
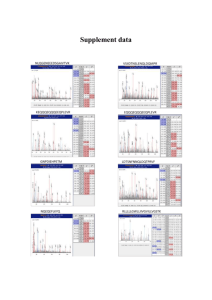
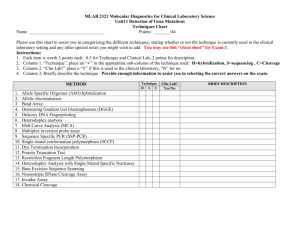
SignalP-NN result: for Humant WNT-1 # data >Sequence length = 70 # Measure Position Value Cutoff max. C 30 0.565 0.32 max. Y 30 0.690 0.33 max. S 12 0.989 0.87 mean S 1-29 0.852 0.48 D 1-29 0.771 0.43 # Most likely cleavage site between signal peptide? YES YES YES YES YES pos. 29 and 30: VTT-EI SignalP-HMM result: # data >TXN4_HUMAN Prediction: Signal peptide Signal peptide probability: 0.984 Signal anchor probability: 0.015 Max cleavage site probability: 0.962 between pos. 29 and 30 # gnuplot script for making Explain the output. Go back. the plot(s) DESCRIPTION OF THE SCORES The graphical output from SignalP (neural network) comprises three different scores, C, S and Y. Two additional scores are reported in the SignalP3-NN output, namely the S-mean and the D-score, but these are only reported as numerical values. For each organism class in SignalP; Eukaryote, Gram-negative and Gram-positive, two different neural networks are used, one for predicting the actual signal peptide and one for predicting the position of the signal peptidase I (SPase I) cleavage site. The S-score for the signal peptide prediction is reported for every single amino acid position in the submitted sequence, with high scores indicating that the corresponding amino acid is part of a signal peptide, and low scores indicating that the amino acid is part of a mature protein. The C-score is the ``cleavage site'' score. For each position in the submitted sequence, a C-score is reported, which should only be significantly high at the cleavage site. Confusion is often seen with the position numbering of the cleavage site. When a cleavage site position is referred to by a single number, the number indicates the first residue in the mature protein, meaning that a reported cleavage site between amino acid 26-27 corresponds to that the mature protein starts at (and include) position 27. Y-max is a derivative of the C-score combined with the S-score resulting in a better cleavage site prediction than the raw C-score alone. This is due to the fact that multiple high-peaking C-scores can be found in one sequence, where only one is the true cleavage site. The cleavage site is assigned from the Y-score where the slope of the Sscore is steep and a significant C-score is found. The S-mean is the average of the S-score, ranging from the N-terminal amino acid to the amino acid assigned with the highest Y-max score, thus the S-mean score is calculated for the length of the predicted signal peptide. The S-mean score was in SignalP version 2.0 used as the criteria for discrimination of secretory and nonsecretory proteins. The D-score is introduced in SignalP version 3.0 and is a simple average of the S-mean and Y-max score. The score shows superior discrimination performance of secretory and non-secretory proteins to that of the S-mean score which was used in SignalP version 1 and 2. For non-secretory proteins all the scores represented in the SignalP3-NN output should ideally be very low. The hidden Markov model calculates the probability of whether the submitted sequence contains a signal peptide or not. The eukaryotic HMM model also reports the probability of a signal anchor, previously named uncleaved signal peptides. Furthermore, the cleavage site is assigned by a probability score together with scores for the nregion, h-region, and c-region of the signal peptide, if such one is found. EXAMPLES OF STANDARD OUTPUT By default the server produces the following output for each input sequence: Example 1: secretory protein The example below shows the output for thioredoxin domain containing protein 4 precursor (endoplasmic reticulum protein ERp44), taken from the Swiss-Prot entry TXN4_HUMAN. The signal peptide prediction is consistent with the database annotation. >TXN4_HUMAN Example 2: non-secretory protein The example below shows the output for BMP-2 inducible protein kinase (EC 2.7.1.37), a nuclear protein taken from the Swiss-Prot entry BM2K_HUMAN. No signal peptide is predicted. >BM2K_HUMAN SignalP-NN result: # data >BM2K_HUMAN # Measure Position max. C 20 max. Y 20 length = 70 Value Cutoff 0.035 0.32 0.034 0.33 signal peptide? NO NO max. S mean S D 12 1-19 1-19 0.263 0.063 0.049 0.87 0.48 0.43 NO NO NO SignalP-HMM result: # data >BM2K_HUMAN Prediction: Non-secretory protein Signal peptide probability: 0.157 Signal anchor probability: 0.023 Max cleavage site probability: 0.027 between pos. 28 and 29 # gnuplot script for making the plot(s) Explain the output. Go back. EXAMPLE OF SHORT OUTPUT When selecting the short output format, the prediction for each submitted sequence (in a multisequence FASTA file) are reported on single line, one for each fasta entry. A two line header is included, showing the information of the different columns. # SignalP-NN euk predictions # SignalP-HMM euk predictions # name Cmax pos ? Ymax pos ? name ! Cmax pos ? Sprob ? TXN4_HUMAN 0.565 30 Y 0.690 30 Y TXN4_HUMAN S 0.962 30 Y 0.984 Y BM2K_HUMAN 0.035 20 N 0.034 20 N BM2K_HUMAN Q 0.027 29 N 0.157 N Smax pos ? Smean ? D ? 0.989 12 Y 0.852 Y 0.771 Y 0.263 12 N 0.063 N 0.049 N #