ANSWERS 2 (57 Marks) - Cerebralenhancementzone
advertisement

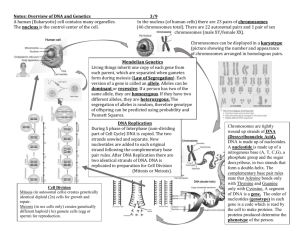
ANSWERS 2 (57 Marks) IB Standard and Higher level Biology Dulwich College Shanghai Topic 4 (SL) and 10 (HL): Genetics 4.1 4.1.1 4.1.2 4.1.3 4.1.4 Chromosomes, genes, alleles and mutations (SL) State that eukaryote chromosomes are made of DNA and proteins. Define gene, allele and genome. Define gene mutation. Explain the consequences of a base substitution mutation in relation to the processes of transcription and translation, using the example of sickle-cell anaemia. 4.2 4.2.1 4.2.2 4.2.3 Meiosis (SL) State that meiosis is a reductive division of a diploid nucleus to form a haploid nuclei. Define homologous chromosomes. Outline the process of meiosis, including pairing of homologous chromosomes and crossing over, followed by two divisions, which results in four haploid cells. Explain that non-disjunction can lead to changes in chromosomes number, illustrated by references to Down syndrome (trisomy 21). State that, in karyotyping, chromosomes are arranged in pairs according to their size and structure. State that karyotyping is performed using cells collected by chorionic villus sampling or amniocentesis, for pre-natal diagnosis of chromosome abnormalities. Analyse a human karyotype to determine gender and whether non-disjunction has occurred. 4.2.4 4.2.5 4.2.6 4.2.7 10.1 10.1.1 10.1.2 10.1.3 Meiosis (HL) Describe the behaviour of the chromosomes in the phases of meiosis. Describe the behaviour of the chromosomes in the phases of meiosis. Explain how meiosis results in an effectively infinite genetic variety in gametes through crossing over in prophase 1 and random orientation in metaphase 1. 10.1.4 State Mendel’s law of independent assortment. 10.1.5 Explain the relationship between Mendel’s law of independent assortment and meiosis 4.3 4.3.1 Theoretical Genetics (SL) Define genotype, phenotype, dominant allele, recessive allele, codominant alleles, locus, homologous, heterozygous, carrier and test cross. 4.3.2 Determine the genotypes and phenotypes of the offspring of a monohybrid cross using a Punnett grid. 4.3.3 State that some genes have more than two alleles (multiple alleles). 4.3.4 Describe ABO blood groups as an example of codominance and multiple alleles. 4.3.5 Explain how the sex chromosomes control gender by referring to the inheritance of X and Y chromosomes in humans. 4.3.6 State that some genes are present on the X-chromosome and absent from the shorter Y chromosome in humans. 4.3.7 Define sex linkage. 4.3.8 Describe the inheritance of colour blindness and haemophilia as examples of sex linkage. 4.3.9 State that a human female can be homozygous or heterozygous with respect to sexlinked genes. 4.3.10 Explain that female carriers are heterozygous for X-linked recessive alleles. 4.3.11 Predict the genotypic and phenotypic ratios of offspring of monohybrid crosses involving any of the above patterns of inheritance. 4.3.12 Deduce the geneotypes and phenotypes of individuals in pedigree charts. 10.2 Dihybrid Crosses and Gene Linkage (HL) 10.2.1 Calculate and predict the genotypic and phenotypic ratio of offspring of dihybrid crosses involving unlinked autosomal genes 10.2.2 Distinguish between autosomes and sex chromosomes 10.2.3 Explain how crossing over between non-sister chromatids of a homologous pair in prophase I can result in an exchange of alleles. 10.2.4 Define linkage group 10.2.5 Explain an example of a cross between two linked genes. Alleles are usually shown side by side in dihybrid crosses, for example TtBb. In representing crosses involving linkage, it is more common to show them as vertical pairs for example: 10.2.6 Identify which of the offspring are recombinants in a dihybrid cross involving linked genes. 10.3 Polygenic Inheritance 10.3.1 Define polygenic inheritance 10.3.2 Explain that polygenic inheritance can contribute to continuous variation using two examples, one of which must be human skin colour 4.4 4.4.1 4.4.2 Genetic Engineering and Biotechnology (SL) Outline the use of polymerase chain reaction (PCR) to copy and amplify minute quantities of DNA. State that, in gel electrophoresis, fragments of DNA move in an electric field and are 4.4.3 4.4.4 4.4.5 4.4.6 4.4.7 4.4.8 4.4.9 4.4.10 4.4.11 4.4.12 4.4.13 separated according to their size. State that gel electrophoresis of DNA is used in DNA profiling. Describe the application of DNA profiling to determine paternity and also in forensic investigations. Analyse DNA profiles to draw conclusions about paternity or forensic investigations. Outline three outcomes of the sequencing of the complete human genome. State that, when genes are transferred between species, the amino acid sequence of polypeptides translated from them is unchanged because the genetic code is universal. Outline a basic technique used for gene transfer involving plasmids, a host cell (bacterium, yeast or other cell), restriction enzymes (endonucleases) and DNA ligase. State two examples of the current uses of genetically modified crops or animals. Discuss the potential benefits and possible harmful effects of one example of genetic modification. Define clone. Outline a technique for cloning using differentiated animal cells. Discuss the ethical issues of therapeutic cloning in humans. Paper 1 Multiple Choice (10 Marks) 1. Two genes A and B are linked together as shown below. A b a B If the genes are far enough apart such that crossing over between the alleles occurs occasionally, which statement is true of the gametes? A. All of the gametes will be Ab and aB. B. There will be 25% Ab, 25% aB, 25% ab and 25% AB. C. There will be approximately equal numbers of Ab and ab gametes. D. The number of Ab gametes will be greater than the number of ab gametes. 2. A polygenic character is controlled by two genes each with two alleles. How many different possible genotypes are there for this character? A. 2 B. 4 C. 9 D. 16 3. A cross is performed between two organisms with the genotypes AaBb and aabb. What genotypes in the offspring are the result of recombination? A. Aabb, AaBb B. AaBb, aabb C. aabb, Aabb D. Aabb, aaBb 4. The diagram below shows a cell during meiosis. How many chromosomes would each daughter cell have at the end of meiosis? A. 1 B. 2 C. 4 D. 8 5. The pedigree below shows which members of a family were Rhesus positive and Rhesus + negative. The allele for Rhesus positive blood (Rh ) is dominant over the allele for Rhesus negative blood (R ). Rhesus positive male I II III Rhesus negative male Rhesus positive female Rhesus negative female Which are possible genotypes of the individuals numbered I, II and III? I II III + + + + + – A. Rh Rh Rh Rh Rh Rh + + + – + + B. Rh Rh Rh Rh Rh Rh + + + – + – C. Rh Rh Rh Rh Rh Rh + – + – + + D. Rh Rh Rh Rh Rh Rh 6. The diagram below shows the results of DNA profiling using gel electrophoresis. origin band I band II What conclusion can be drawn about the DNA in bands I and II? A. The DNA in the two bands has the same base sequence. B. The DNA in the two bands consists of fragments of the same length. C. The DNA in the two bands has the same ratio of bases. D. The DNA in the two bands came from the same source. 7. The diagram below shows chromosomes during prophase I of meiosis. How many chromosomes and chiasmata are visible? A. B. C. D. Number of chromosomes 2 4 2 4 Number of chiasmata 2 2 4 4 8. In peas the allele for round seed (R) is dominant over the allele for wrinkled seed (r). The allele for yellow seed (Y) is dominant over the allele for green seed (y). If two pea plants with the genotypes YyRr and Yyrr are crossed together, what ratio of phenotypes is expected in the offspring? A. 9 round yellow : 3 round green : 3 wrinkled yellow : 1 wrinkled green B. 3 round yellow : 3 round green : 1 wrinkled yellow : 1 wrinkled green C. 3 round yellow : 1 round green : 3 wrinkled yellow : 1 wrinkled green D. 1 round yellow : 1 round green : 1 wrinkled yellow : 1 wrinkled green 9. What is a difference between autosomes and sex chromosomes? A. Autosomes are not found in gametes but sex chromosomes are. B. Sex chromosomes are found in animal cells and autosomes are found in plant cells. C. Autosomes are diploid and sex chromosomes are haploid. D. Sex chromosomes determine gender and autosomes do not. 10. The diagram below shows a cell in meiosis. What can be deduced from this diagram? [Source: J W Saunders, (1968), Animal Morphogenesis, MacMillan, page 7] Stage of meiosis shown Haploid number of chromosomes in this cell A. Metaphase I 6 B. Prophase I 3 C. Prophase I 6 D. Metaphase I 3 Paper 2 Section A Data Analysis (17 marks) 1. Rats were bred for several generations to prefer alcohol (ethanol) consumption. When tested, it was discovered that the brains of these rats possessed lower quantities of the chemical neuropeptide Y (NPY). To test the hypothesis that lower quantities of NPY leads to a preference for alcohol, rats were genetically engineered to be NPY deficient (genotype NPY –/–), or to produce an excess of NPY (NPY-EX). In separate experiments, the two groups were compared to normal rats (in terms of their alcohol preference) possessing the genotype NPY +/+. The groups were offered solutions of increasing alcohol concentration. The quantity of each solution consumed per day was measured. Figure 1 Figure 2 NPY-EX NPY +/+ Π1 12 Π1 12 15 Π1 Consumption / g kg day Π1 NPY Π/Π NPY +/+ Consumption / g kg day 15 9 6 3 0 9 6 3 0 0 5 10 15 20 25 Concentration of alcohol solution / % 0 5 10 15 20 25 Concentration of alcohol solution / % [Source: adapted from Thiele et al, Nature, (1998), 396 pages 366–369] (a) (b) Calculate the difference in consumption of the 6% alcohol solution between the (i) NPY –/– and NPY +/+ rats (figure 1); –1 –1 2.8 (±0.5) g kg day (1) (ii) (1) NPY-EX and NPY+/+ rats (figure 2). –1 –1 2.0 (± 0.5) g kg day Compare the alcohol consumption of the NPY –/– rats with the NPY-EX rats. NPY – / – consumes more alcohol (1) than NPY-EX (at all concentrations); consumption increases at a (relatively) constant rate (above 6%) with concentration for NPY-EX, but levels off at higher concentrations for NPY – / – (1); as alcohol concentration increases both NPY – / – and NPY-EX rats consume more; (1) NPY – / – consumes more alcohol than NPY + / + and NPY-EXconsumes less (3) than NPY + / +; (Accept use of word control.) biggest difference from NPY + / + for NPY-EX is at 20%, but for NPY – / – it is at 10%; (c) Identify the relationship between NPY levels and alcohol consumption. NPY levels are inversely related to alcohol consumption the lower the NPY levels,the more alcohol consumption (Accept the converse.) (1) An experiment was carried out to test the hypothesis that an increase in preference for alcohol might be related to a decrease in sensitivity to its effects. Rats were injected with a sample of alcohol and then assessed for the length of time it took for them to regain the righting reflex. (The righting reflex refers to the ability of the rat to return to its feet after being placed on its back.) Figure 3 75 Time to regain 50 righting reflex / mins 25 0 NPY Π/Π NPY +/+ NPY-EX [Source: adapted from Thiele et al, Nature, (1998), 396 pages 366–369] (d) Deduce the relationship between NPY levels and the time required to regain the righting reflex. (3) NPY – / – takes least time to regain the reflex; NPY-EX takes most time to regain the reflex; NPY-EX is the most sensitive to effects of alcohol / NPY – / – is the least sensitive to effects of alcohol; the higher the NPY levels the more time taken to regain the reflex; NPY– / – / under-expression has more of an effect than NPY-EX / overexpression; An additional experiment was carried out to determine whether differences in sensitivity to the effects of alcohol might be related to differences in the rats’ ability to remove alcohol from their blood. Rats were injected with alcohol and blood samples were taken one hour and three hours later to determine alcohol levels. The results are shown below. Figure 4 Key: NPY Π/Π 300 NPY +/+ Concentration of alcohol Π1 200 in the plasma / mg dL 100 0 1 3 Time after injection / hours (e) Evaluate the hypothesis that differences in sensitivity to the effects of alcohol might be related to differences in the ability of the rats to remove alcohol from their blood. (2) there is no difference / small difference in blood levels between groups at 1 hour and 3 hours / decrease from 1 to 3 hours is the same for both; therefore, the hypothesis does not appear to be justified; (f) (g) Using all the data, outline the relationship between preference for alcohol and sensitivity to the effects of alcohol. NPY-EX does not prefer alcohol and is sensitive to effects of alcohol NPY – / – prefers alcohol and is not sensitive to effects of alcohol; therefore, alcohol preference is inversely related to sensitivity to effects of alcohol the less sensitive the rats are to alcohol, the more they consume it; (2) (i) Define the term homozygous. having two identical alleles (of a gene) (1) (ii) State the phenotype of a rat with the genotype NPY +/+. normal preference for alcohol / moderate preference for alcohol / standard phenotype for alcohol preference / normal levels of NPY Do not accept "normal rats" only. (1) (iii) Using a Punnett grid, predict the fraction of offspring that would have the genotype NPY +/– if two rats were crossed, one homozygous for the NPY+ allele and one homozygous for the NPY– allele. (2) Punnett square correctly drawn; Can be two by two or one by one, accept + / – without NPY, square or diamond shape acceptable but clear cells expected, male or female symbols not necessary. 100% / all; Paper 2 Section A Short Structured (13 Marks) 1. The diagram below shows the pedigree of a family with red green colour-blindness, a sex-linked condition. Key 1st generation 1 normal male 2 normal female male with condition 2nd generation 1 2 1 2 3 4 5 3 4 female with condition 3rd generation (a) Define the term sex-linkage. a gene / trait / allele carried on a sex chromosome / X and Y / X / Y; (b) Deduce, with a reason, whether the allele producing the condition is dominant or recessive. (2) recessive; (1) evidence from the pedigree; eg 2nd generation–2 and 3 do not have the condition but have one child who does; (1) (c) (i) (ii) 2. Determine all the possible genotypes of the individual (2nd generation–1) using appropriate symbols. a X Y (where a = condition); (1) (1) Determine all the possible genotypes of the individual (3rd generation–4) using appropriate symbols. (1) A a A A X X or X X where A = normal, a = condition (must have both); (If upper case letter and lower case letter are reversed then the ECF rule applies.) In Zea mays, the allele for coloured seed (C) is dominant over the allele for colourless seed (c). The allele for starchy endosperm (W) is dominant over the allele for waxy endosperm (w). Pure breeding plants with coloured seeds and starchy endosperm were crossed with pure breeding plants with colourless seeds and waxy endosperm. (a) State the genotype and the phenotype of the F1 individuals produced as a result of this cross. C c W w; (1) all are coloured starchy; (1) (2) (b) The F1 plants were crossed with plants that had the genotype c c w w. Calculate the expected ratio of phenotypes in the F2 generation, assuming that there is independent assortment. Use the space below to show your working. gametes are C W, C w, c W, c w and c w; (1) F2 genotypes are CcWw, Ccww, ccWw and ccww; (1) (3) 1 coloured starchy: 1 coloured waxy: 1 colourless starchy: 1 colourless waxy; (1) Phenotypes must be unambiguously indicated, but not necessarily on the line. The observed percentages of phenotypes in the F2 generation are shown below. coloured starchy 37% colourless starchy 14% coloured waxy 16% colourless waxy 33% The observed results differ significantly from the results expected on the basis of independent assortment. (c) State the name of a statistical test that could be used to show that the observed and the expected results are significantly different. (1) chi-squared test (d) Explain the reasons for the observed results of the cross differing significantly from the expected results. (2) (autosomal) linkage (reject sex linkage) / genes are on the same chromosome / genes do not assort independently; coloured starchy and colourless waxy are parentals / coloured waxy and colourless starchy are the recombinants; recombinants produced by crossing over; Section B Extended Response (17 Marks) 1. Describe how sexual reproduction promotes genetic variation within a species. random orientation of bivalents / pairs of chromosomes; maternal and paternal chromosome could go to either pole; n 2 combinations; eg over 8 million in humans; crossing over; exchange of material between homologous chromosomes / non-sister chromatids; segregation of alleles in meiosis; combinations of alleles are broken up; fertilization brings together genes / alleles from two different parents; fertilization generates new combinations of genes / alleles; random fertilization / many possible combinations of male and female gamete; eg over 64 million million in humans (ignoring crossing over); (4) 2. 3. Outline the differences between the behaviour of the chromosomes in mitosis and meiosis. two divisions in meiosis, only one in mitosis; meiosis results in haploid cells, mitosis in diploid cells; crossing over only occurs in meiosis; no S phase precedes meiosis II; chromosome behaviour in meiosis II and mitosis is similar / chromosome behaviour in meiosis I and mitosis is different; chiasmata only form during meiosis; homologous chromosomes move to the equator in pairs only in meiosis; Do not accept number of cells produced - it is a result not a behaviour. Explain the use of two named enzymes in biotechnology. Examples and application: pectinase; obtained from citrus fruits / tomatoes / apples; used in fruit juice production; breaks down pectin allowing cells to separate; assisting in juice formation; juice formed is clear; high juice yield using enzyme; 2nd enzyme: name; (1) e.g restriction enzymes/ lactase/ amylase, proteases, lipases source; (1) use; (1) genetic engineering/ for lactose intolerance/ biological washing powders mode of action; (1) cutting DNA fragments/ converts lactose into galactose and glucose/ digests food stains e.g. starch to maltose, proteins to amino acids, lipids to fatty acids and glycerol advantage of using enzyme; (1) allows recombination of DNA/ allows lactose intolerant people to use milk products without consequences/ low temperatures for washing details of enzyme use; 8 max Possible second example could be meat tenderising – papain from papaya fruit / bromelain from the pineapple plant, biological washing powders – amylases / proteases / lipases, glucose biosensors – glucose oxidase / peroxidase, cheese making – rennin, high fructose syrups – glucose isomerase, breadmaking – fungal amylases / fungal proteases, DNA profiling – DNA ligase, etc. (Plus up to [2] for quality) (5) (8)