Supplementary Information
advertisement

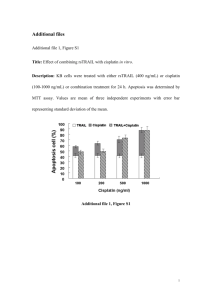
SUPPLEMENTARY MATERIALS AND METHODS DNase protection assay. Assessment of DNase protection was done as previously described (31), with modifications. Specifically,10 µL of polyplex solution (N/P = 30/1) harboring 1.0 µg of DNA was incubated with 1 µL of 40 mU/µL RQ1 RNase-free DNase (Promega, Madison, WI) solution at 37 °C for time points of 0.5, 1, 1.5, and 2 h. Gel electrophoresis was then performed on an agarose gel (3% w/v) containing 60 µg of ethidium bromide/100 mL Tris-Borate-EDTA (TBE) buffer to observe the presence and integrity of the ODN decoys. Nuclear protein extraction. Cells were released from culture plates by trypsinization and pelleted by centrifugation at 200 g for 5 min. Cell pellets were resuspended in ice-cold buffer and nuclei isolated and lysed as previously described (22). Protein concentration was determined by Bio-Rad (Hercules, CA) protein assay with bovine serum albumin as a standard. Electrophoretic mobility shift assay (EMSA). A double-stranded oligonucleotide probe for NF-κB (5’-AGTTGAGGGGACTTTCCCAGGC3’, Promega, Madison, WI) containing an NF-κB-binding site (underlined) was end-labeled using [-32P] ATP with T4 polynucleotide kinase (Promega, Madison, WI). The labeled probe was purified and EMSA performed as previously desribed (22). After densitometry measurements of EMSA bands, IC50 values were calculated using GraphPad Prism 4. MTT assay. H9c2(2-1) cells were plated into 96-well plates at 5,000 cells/well in supplemented DMEM at 5% CO2 and 37 ºC for 24 h before treatment. Each glycopolymer was dissolved in DNase-free water and diluted to the appropriate concentration in serum-free medium (Opti-MEM; 250 µL total Vol). JetPEI solution was also diluted via the same method into Opti-MEM. Supplemented DMEM (250 μL) was added to each well 4 h after transfection. Twenty four hours later, the amount of cell death was assayed using an MTT kit (Vybrant MTT Cell Proliferation kit, Invitrogen, Carlsbad, CA.) according to the manufacturer’s protocol. Absorbance of each well was read at 570 nm using a GENios Pro plate reader (TECAN US, Research Triangle Park, NC). Statistical Analysis. Results are reported as means ± standard error of the mean (SEM). Unpaired student t-test were performed and P-values reported with differences between groups considered significant at P ≤ 0.05. Power analysis was used to ensure proper sample size needed to determine significance.