BIOTECH Project, University of Arizona
PCR of GFP in pGLO transformed E. coli
Name:
Date:
Period:
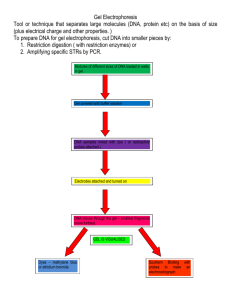
Lab Investigation: Examining a Single Gene
In this lab investigation, you will learn about a technique called polymerase chain reaction (PCR)
that allows us to examine a very small piece of DNA.
To look at a small piece of DNA, first you have to choose what DNA you want to look at, then you
have to make a bunch of copies of that piece of DNA so you can see it.
• You will make copies of DNA is just like your cells do when they replicate DNA. What ingredients
does your cell use when it replicates its DNA?
• Keeping in mind what a cell does when it replicates its DNA, make a list of steps involved in
replicating DNA:
You will use some of these same ingredients and steps to replicate DNA in a test tube instead of a cell.
The piece of DNA you will replicate is called the Green Fluorescent Protein (GFP) gene. This gene
codes for the GFP protein, a protein normally produced by jellyfish that you transformed into bacteria
in a plasmid (pGLO). The protein can be excited by UV light. In one group of plates (LB agar +
ampicillian) the bacteria transformed with the pGLO plasmid "glowed". In the other group of plates
(LB agar + ampicillian) the bacteria did not glow. Why do you think the bacteria glowed or did not
glow?
Since the same plasmid was added to both of the plates, you assume that the DNA is the same in the
bacteria on both plates. Based on the above discussion, how could you determine if this is indeed
correct?
1
BIOTECH Project, University of Arizona
PCR of GFP in pGLO transformed E. coli
Conducting PCR
Materials/Equipment Needed
• Thermocycler
• Microcentrifuge (optional)
• Micropipet and sterile pipet tips
• Bacteria colonies from both LB agar + ampicillian
For each reaction add the following to a PCR thin walled tube:
• 10 µl 2X GoTaq (contains Taq polymerase, dNTPs, MgCl2, and buffer for ideal reaction and loading
dye for electrophoresis).
• 5 µl forward primer
• 5 µl reverse primer
• a little dab of bacteria from one colony
These are small volumes; you will need to be sure to look at the pipette tip when you are pipetting to
make certain that the components are being added.
Some groups will amplify control reaction, for each control reaction add the following to a PCR thin
walled tube:
• 5 µl each primer
• 10 µl 2X GoTaq
• >0.5 µl control DNA for (+) add the GFP plasmid, and (-) add nothing to the reaction
Procedure
1. Label the PCR tube so that you can distinguish DNA from the Amp plates with glowing bacteria, or
DNA from the Amp plates with nonglowing bacteria.
2. Add 5µl primer of each primer to each tube. If necessary, gently tap you tube on the counter to get
all of the liquid to the bottom of the tube.
3. Add 10 µl GoTaq (green solution). Close the tubes and centrifuge briefly (10 seconds) to pool all
of the liquid at the bottom of the tube (if you do not have a centrifuge then tap or shake the tube
contents to the bottom of the tube).
4. Using the micropipet with a clean tip, just barely touch one of the colonies that you would like to
amplify the DNA to test for the presence of GFP. If you can see the bacteria on the tip you have
too much. Place the tip into the water in the PCR tube and pipet up and down once or twice to
dislodge the bacteria and mix. To the control PCR reaction add >0.5 µl of control DNA.
5. Place your tube into the thermocycler to run the 'GFP' program. This program is 30 cycles of:
• 94°C for 30 seconds
• 55°C for 45 seconds
• 72°C for 1.5 minutes
2
BIOTECH Project, University of Arizona
PCR of GFP in pGLO transformed E. coli
6. After the cycles are complete, PCR reactions can be refrigerated or prepared for electrophoresis.
Electrophoresis of your PCR reactions
Materials/Equipment Needed
• Electrophoresis apparatuses, electrodes, and power supplies
• Micropipet
• Micropipet tips
• Loading dye
• 0.8% agarose gel
• Molecular weight markers (1 tube per gel)
• Water bath at 55°C or hot plate
• Thermometer for water bath
• TAE buffer
• Ethidium bromide paper (1 piece per gel)
• Staining tray
• Gloves (for handling ethidium bromide)
• UV light box
• UV light Polaroid set up (including camera, film, camera connector, and light shield)
• Biohazard bag
3
BIOTECH Project, University of Arizona
PCR of GFP in pGLO transformed E. coli
Procedure
Pouring an agarose gel
1. Get your electrophoresis apparatus and seal both ends of the gel tray with tape or stoppers.
2. Make sure one comb is in place at the negative electrode (black end of the gel).
3. Pour melted agarose into the gel space until the gel is about 5 mm deep. Let the agarose
harden, which should take 5-10 minutes. Don’t touch/move your gel until it’s hard. In the
meantime, prepare your PCR reactions for electrophoresis.
Electrophoresis of your PCR reactions
1. Using the micropipet with a clean tip, pipet 5 µl gel loading dye into your PCR reaction tube.
You will load both your PCR reactions and standard DNA markers sample into the gel. A standard
DNA marker has a bunch of different sized pieces of DNA so you can compare it to the DNA from
your PCR reaction to figure out what size piece it is. Two or three groups can share a gel, but only one
molecular weight marker is needed per gel.
2. Draw a picture of your gel and label in which wells you will load which samples (PCR reaction(s),
DNA marker). Be certain to have the information of where the other groups added their samples.
3. When your gel has hardened, remove the tape or stoppers.
4. Load your samples into the wells - be sure you keep track of which samples you're loading in which
wells.
5. Pour TAE buffer carefully so it fills the electrophoresis apparatus and just covers the gel.
6. Run that gel! Plug the electrodes into your electrophoresis apparatus (red to red, black to black),
being careful not to bump your gel too much.
7. Plug the power source into an outlet and set the voltage to about 100 V (max = 120 V).
8. Let the gel run until the dye migrates about 5-6 cm from the wells (about 20-25 minutes).
9. Turn off the power supply, disconnect the electrodes, and remove the top of the electrophoresis
apparatus.
10. Carefully remove the gel. The gel can be wrapped in plastic wrap and stored in the refrigerator or
placed it in the staining tray for DNA staining.
4
BIOTECH Project, University of Arizona
PCR of GFP in pGLO transformed E. coli
Staining gels to examine PCR reactions
1. Place gel in staining tray
2. Using gloves, remove the plastic from the ethidium bromide sheet and place the ethidium bromide
paper on the gel. Gently rub the paper with your fingers to make sure it is contacting the gel all
over.
3. Stain for about 10 minutes.
4. Put the gel on the UV light box and, with the UV shield down, view your gel.
5. Take a Polaroid picture of your gel.
Analysis
• What do you see on your gel? Which bacteria samples were able to amplify the GFP gene from the
pGLO plasmid? Why do you think you saw these results? What else could you do to ascertain the
amplified DNA is the GFP gene?
5
 0
0