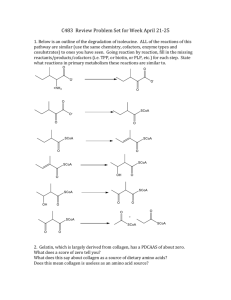
Amino Acids & Proteins
advertisement

WADE CHAPTER TWENTY-FOUR OUTLINE: AMINO ACIDS AND PEPTIDES Skip: 24-7C; 24-12:24-14 1. Structure of Twenty -amino acids (Table 24-2) 2. Acid/Base properties (Section 24-3) a. Zwitterions b. pKa’s c. Isoelectric Point: pH of Neutral zwitterion (Section 24-4) d. Gel electrophoresis o Buffered solution selects for particular charge state depending on isoelectric point o high charge density elutes first 3. Synthesis of Amino Acids (Summary p 1168) a. Reductive Amination of -ketoacid (RCOCOOH to RCHNH2COOH) b. SN2 reaction of ammonia (excess) with -haloacid (24-5B) c. Gabriel-Malonic Ester Synthesis (Section 24-5C) o N-phthalimidomalonic ester alkylation with NaOEt then RX o Hydrolysis of ester and release of N o Decarboxylation with heat d. The Strecker Synthesis: RCHO + NH3 and HCN then H3O+: (Mechanism pg 1168) o Formation of Imine o Formation of -amino propionitrile o Hydrolysis of nitrile to carboxylic acid 4. Amino Acid Reactivity (review) a. Acylation of amino group b. Esterification of Carboxylic Acid group 5. Resolution of Amino Acids (Section 24-6) a. React racemic aa with acetic anhydride b. expose to enzymatic deacylation with an “acylase” enzyme to hydrolyse selectively just the natural (L) amino acid 6. Peptides (Section 24-8) 7. The peptide bond: Amide between amino acids a. Naming convention: N terminus at left; C terminus at right b. Primary structure: Three letter designations (Arg-Pro-Gly) 8. Sequencing : Determining the Amino Acid Composition of a polypeptide (Section 24-9) a. Disulfide Linkage cleavage b. Acid-catalyzed Hydrolysis c. Trypsin Cleavage: C-terminus of basic amino acids lysine& arginine d. Chymotrypsin cleavage: C-terminus of aromatic aa Phe, Tyr, Trp 9. Edman Degradation: Grabbing the N-Terminus (Section 24-9C) a. Rxn with phenyl isothiocyanate (S=C=N-Ph) to bind to N-terminus b. Acid catalyzed Intramolecular rxn with C=O of same amino acid c. Decomposition of thiazolinone to expel N-terminus aa 10. Peptide Synthesis (aa1-aa2) a. Traditional solution synthesis (Section 24-10B) o Protect N-terminus of aa1 with benzylchloroformate (PhCH2OCOCl) to make Z-aa1 o Activate C-terminus of aa1 with ClCOOEt ethylchloroformate o Join together with aa2 protected at C terminus as ester o Remove benzyl protecting group with hydrogenation H2/Pd to make aa1-aa2(ester) b. Merrifield Solid Phase synthesis (Section 24-11B) o Protect N-terminus with BOC anhydride (Me3COOCOOEt) o Bond carboxylate of aa2 to solid support via SN2 at bonded Benzyl chloride branches o Remove N-protection w CF3COOH (release CO2 and isobutylene) o Couple to second aa with DCC (dicyclohexylcarbodiimide) o Rinse solid support to remove by-products like CH2=CMe2 o Add second N-protected amino acid via C-terminus LEARNING OUTCOMES: Identify amino acids. Identify the structure of a specific amino acid at a given pH Understand the role of protecting groups in Organic synthesis Propose a series of reactions to produce a given polypeptide. Propose a sequence of steps to sequence a polypeptide using traditional wet chemistry and using solid support Merrifield Synthesis. Given evidence from the results of a polypeptide sequencing experiment, deduce the primary structure of a polypeptide. Draw the mechanism for Edman degradation of a peptide using curved arrow formalism. Propose an appropriate laboratory method(s) for the separation and identification of a protein. SAMPLE EXAM PROBLEMS: 1. Draw the fragments that would form if the following polypeptide were treated with chymotripsin: Phe-Val-Asn-Gln-Tyr-His-Leu-Gly-Ser-His-Leu-Val-Glu-Ala 2. Draw the structure of the amino acid cysteine at pH 8. 3. A heptapeptide contains the amino acids arginine, glycine, isoleucine, phenylalanine, proline (two residues) and valine. Treatment of the heptapeptide with trypsin gives arginine and a hepeptide. Some of the fragments that were obtained by partial hydrolyses of the heptapeptide are Pro-Phe-Ile, Arg-Gly-Pro, Pro-Phe-Ile-Val and Pro-Pro. Assign a structure to the heptapeptide. 4. Draw the organic products A, B and C missing from the following reaction sequence. Include stereochemistry where appropriate: