Supplementary Fig. 1. Illustrations of 9 tested miR
advertisement

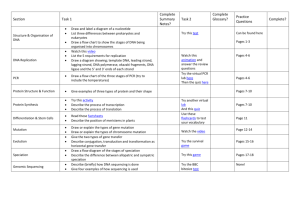
Supplementary Fig. 1. Illustrations of 9 tested miR-hosting CpG islands. Open frame, location of pre-miRNA transcript. Open arrow, pre-miRNA transcript direction; Blue-underline, locations of the forward and reversed primer-matching regions used to detect methylation of CpG island with DHPLC and bisulfite sequencing; Purple vertical bars, CpG sites 1 m 8V U 7 H2O B 6 GES-1 M CALU3 5 HCT116 H1299 4 MGC803 SGC7901 3 DU145 HEPG2 2 AGS MKN74 1 SIHA MB 0 3 4 5 min Supplementary Fig. 2. DHPLC chromatogram of methylated and unmethylated miR-9-1 in various cell lines. UV-detector; the partial denaturing temperature, 55.4C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 2 mV 10 U U H2O B H1299 SIHA M A549 DU145 SGC7901 BGC823 MKN45 RKO HELA MGC803 HEPG2 AGS MKN74 MB 0 3 4 5 min Supplementary Fig. 3. DHPLC chromatogram of methylated and unmethylated miR-9-3 in various cell lines. UV-detector; the partial denaturing temperature, 58.5C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 3 mV U 8 7 H2O B 6 PG M (partial) GES-1 A549 5 CALU3 4 MKN45 M (full) HCT116 3 RKO MGC803 2 BGC823 AGS 293T 1 MB 2 3 4 5 min Supplementary Fig. 4. DHPLC chromatogram of methylated and unmethylated miR-34b in various cell lines. UV-detector; the partial denaturing temperature, 56.8C; U, peak for the unmethylated PCR products; M (partial or full), peak for the partially or fully methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 4 200 mV U H2O B MKN45 AGS 100 HELA SIHA DU145 RKO A549 M MGC803 PC-3 KG1A HL60 MB 0 2 3 4 5 min Supplementary Fig. 5. DHPLC chromatogram of methylated and unmethylated miR-210 in various cell lines. Fluorescence-detector; the partial denaturing temperature, 58.7C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 5 U mV 900 H2O B 800 H1299 A549 700 SGC7901 M SIHA 600 MGC803 HEPG2 500 293T MKN45 400 BGC823 HELA 300 SW480 HCT116 200 AGS MKN74 100 RKO MB 0 2 3 4 5 min Supplementary Fig. 6. DHPLC chromatogram of methylated and unmethylated miR-137 in various cell lines. Fluorescence-detector; the partial denaturing temperature, 58.3C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 6 mV U H2O B CALU3 SW480 M GES-1 PC-3 H1299 MGC803 A549 RKO HELA BGC823 SGC7901 AGS MKN45 293T MKN74 MB 0 3 4 5 min Supplementary Fig. 7. DHPLC chromatogram of methylated and unmethylated miR-375 in various cell lines. UV-detector; the partial denaturing temperature, 55.7C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 7 mV U H2O 10 M DU145 HCT116 SW480 MKN74 SIHA A549 HELA BGC823 GES-1 0 SGC7901 MGC803 H1299 HL60 KG1A MKN45 MB B -10 3 4 5 min Supplementary Fig. 8. DHPLC chromatogram of methylated and unmethylated miR-200b in various cell lines. UV-detector; the partial denaturing temperature, 55.6C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 8 mV U H2O 70 B 60 M F0856 50 F1297 40 F1047 30 MKN74 20 AGS 10 MB 0 3 4 5 min Supplementary Fig. 9. DHPLC chromatogram of methylated and unmethylated miR-193b in various cell lines. Fluorescence-detector; the partial denaturing temperature, 56.5C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 9 U mV M 6 5 4 AGS 3 MKN74 MGC803 2 SGC7901 BGC823 1 MKN45 H2O 2 3 4 5 min Supplementary Fig. 10. DHPLC chromatogram of methylated and unmethylated miR-203 in various cell lines. UV-detector; the partial denaturing temperature, 57.7C; U, peak for the unmethylated PCR products; M, peak for the methylated PCR products; B, peripheral blood DNA; MB, M.sssI-methylated blood DNA 10 Supplementary Fig. 11. Distribution of miR methylation in various gastric mucosa samples with and without H. pylori infection. Each line represents methylation status of one miR gene and each column represents one sample. The proportion of methylated allele is displayed step-wisely: negative (white), positive and <30% (light gray), or 30%-60% (deep gray) or >60% (black). Not informative sample is marked as “ND”. The case with or without H. pylori-specific 23S rDNA was labeled with letter “p” (light green) and “n” (blue). 11 Supplementary Fig. 12. Comparison of levels of 7 mature miRNAs by quantitative RT-PCR in 20 pairs of fresh gastric carcinoma (GC) and the corresponding surgical margin (SM) tissue samples. The relative miRNA levels of the paired GC and SM samples from the same patient were linked by a line. 12