lec14_2013 - Andrew.cmu.edu
advertisement

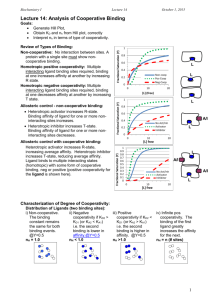
Biochemistry I Lecture 14 February 15, 2013 Lecture 14: Analysis of Cooperative Binding Key Terms: Hill coefficient, nh 1. Positive cooperativity: nh >1 2. Negative cooperativity: nh < 1 3. Non-cooperative: nh = 1 A homotropic positively cooperative system will show a _____________ in affinity as ligand binds, due to the stabilization of the _______ state. A homotropic negatively cooperative system will show a ____________ in affinity as ligand binds, due to the stabilization of the _______ state. Hill Plot: log(Y/(1 - Y)) versus log[L] x-intercept = logKD Review of Types of Binding: 1 protein with a single site must show noncooperative binding. Homotropic positive cooperativity: Multiple interacting ligand binding sites required, binding at one increases affinity at another by increasing R state. Fractional Saturation (Y) Non-cooperative: No interaction between sites. A 0.8 0.6 0.4 Homotropic negative cooperativity: Multiple 0 interacting ligand binding sites required, binding 0 Heterotropic activator increases R-state. Binding affinity of ligand for one or more noninteracting sites increases. Heterotropic inhibitor increases T-state. Binding affinity of ligand for one or more noninteracting sites decreases. Fractional Saturation (Y) at one decreases affinity at another by increasing T state. Allosteric control - non-cooperative binding: 5 10 15 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 20 [L] (free) No Act/Inh Activator Inhibitor 0 5 10 15 20 [L] free Fractional Saturation (Y) Allosteric control with cooperative binding: Heterotropic activator increases R-state, increasing average affinity. Heterotropic inhibitor increases T-state, reducing average affinity. Ligand binds to multiple interacting states (homotropic) with some form of cooperative binding, neg or positive (positive cooperativity for the ligand is shown here). Non-coop Pos-Coop Neg-Coop 0.2 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 It is possible to quantify the degree of cooperativity by 0 analysis of the binding data using a Hill Hill Coefficient plot. The outcome of this analysis is <1 the Hill coefficient, which has the =1 characteristics summarized to the right: >1 Behavior of Cooperative Systems: Single Site Binding Non- = n, number of binding sites. cooperative: 1 1 [ M ][ L] [ L] KD KD [ ML] [ L] Y 1 1 [ M ] [ ML] K D [ L] [M ] [ M ][ L] 1 [ L] KD KD No Act/Inh Activator Inhibitor 5 10 15 20 [L] free Interpretation Negative cooperativity. Non-cooperative Positive cooperativity. Infinitely strong positive cooperativity. When Y = 0.5 what is [L]? 1 Biochemistry I Lecture 14 February 15, 2013 Consider a two step binding: A1 [ M ] [ L] K [ ML ] K D1 i) Non-cooperative. The binding constant remains the same for both binding events. A2 [ ML ] [ L] K [ ML2 ] [ M ][ L] [ ML][ L] K D2 [ ML] [ ML2 ] ii) Positive cooperativity if KD2 < iii) Negative cooperativity if KD2 > KD1 (or KA2 > KA1) i.e. the KD1 (or KA2 < KA1) i.e. the second second binding is higher in binding is lower in affinity. affinity. Two Binding Sites - Infinitely Positive Cooperativity: Consider an infinitely positive cooperative system such that the second dissociation constant (KD2) is much lower than the dissociation constant for binding the first ligand (KD1). Then the only species present in solution are [M] and [ML2]. The fractional saturation under these conditions is: 1 1 [ M ][ L][ L] [ ML2 ] K D2 K D1 K D 2 [ L]2 Y 2 1 1 [ M ] [ ML2 ] [M] [ ML][ L] [ M ] [ M ][ L][ L] K D1 K D 2 [ L] K D2 K D1 K D 2 [ ML][ L] When Y = 0.5 what is [L]? Generalize to N-Ligands - Infinitely Positive Cooperativity: Y [ L]n K D1K D1 K Dn [ L]n When Y = 0.5 what is [L]? N-Ligands – General Cooperative Binding: For less cooperative systems, the fractional saturation can be approximated by: Y [ L]nh n K Dh ave [ L]nh Where nh is the Hill coefficient. This is a measure of the degree of cooperativity. KD-AVE is the "average" KD; when [L]=KDAVE, Y=1/2. The KD-AVE is the root of a polynomial of the individual dissociation constants (Note: for two binding sites KD-AVE= 2√𝐾𝐷1 𝐾𝐷2 , regardless of the degree of cooperativity.) 2 Biochemistry I Lecture 14 Single Site - Noncooperative Y Two sites - infinitely positive cooperative Y N-sites - infinitely positive cooperative: N-sites - just plain pos. cooperative: Y Y Y=0.5 when [ L] K D [ L]1 K D [ L]1 [ L]2 K D1 K D 2 [ L]2 [ L] N K D1 K D 2 K Dn[ L] N [ L]nh n K Dh ave [ L]nh Y=0.5 when [ L] 2 K D1 K D2 [ML] = 0 [ML2] > 0 Y=0.5 when [ L] N K D1 K D2 ...K DN [ML]…[MLN-1] = 0 [MLN] > 0 Y=0.5 when [ L] K D ave Hill Equation and Plot: The Hill coefficient, and the "average" KD can be obtained from a Hill Plot. The Hill plot is based on the following transformation of the above binding equation: Plot of log [Y/(1 - Y)] versus log[L] The Hill coefficient, nh, is the slope as the line crosses the x-axis. The logKD-ave is the intersection of the Hill curve with the x-axis. Non-Cooperative Systems (n =1): Y [ L] K D [ L] log Y log K D log[ L] (1 Y ) This is a straight line with a unit slope. Intersection with x-axis (Y = 0.5) gives the true KD. 0 log February 15, 2013 1 log[L] KD Y [ML]…[MLN-1] > 0 [MLN] > 0 [ L ]n h n K Dh ave [ L]nh Y [ L]nh 1 Y K nh D Ave Y nh log K D ave nh log[ L] log (1 Y ) Log Y 1-Y 0 1 log[L] KD log K D log[L] K D [ L] log Log[L] Cooperative Systems. Low ligand: At very low ligand concentration, the binding appears non-cooperative because most of the macromolecule is in the [M] form. Therefore the Hill plot is initially linear, with a slope =1, intersecting x-axis at logKD1. High ligand: At very high ligand concentration, the binding also appears non-cooperative because most of the macromolecule is in the [MLn] form. Therefore the Hill plot is again linear, with a slope = 1, intersecting the xaxis at log KDn. Intermediate Ligand Concentration (@Y=0.5): Slope: Hill coefficient (0 ↔ 1 ↔ n) Intercept: Ligand concentration to give Y=0.5 (True KD for non-cooperative binding.) 3 Biochemistry I Lecture 14 February 15, 2013 Microscopic and Macroscopic Binding constants: Cooperativity requires a change in the affinity of the binding site. In the case of positive cooperativity the affinity increases (KA increases, KD decreases). In the case of negative cooperativity the affinity decreases (KA decreases, KD increases). For proteins that bind multiple ligands to identical sites the experimentally determined affinity constants will change even if there is no cooperativity. For a two binding sites the affinity for the second site will be lower by a factor of two. This difference is due entirely to statistical factors – there are no differences between the molecular interactions in the two sites. Microscopic KA: This is the association constant for a single site, and is just the ratio of the on- and off-rates: KA=kON/kOFF. It reflects the intrinsic affinity between the protein and the ligand: Go = -RT ln KA kon koff KA Macroscopic KA: This is the observed KA based on the concentrations of the various species, i.e. KA1=[ML]/[M][L], i.e. what you would typically consider the equilibrium constant. One-binding site: The macroscopic and microscopic are the same. M Microscopic kon kon Two binding sites: For the first binding event this is 2KA since there are two ways to form the [ML] species, i.e. KA1=2kON/kOff. For the second binding event there are two ways for the ligand to leave, so KA2=kON/2kOff. Three binding sites ML koff kon koff kon koff koff Macroscopic 2kon kon koff 2koff KA1 M ML KA1 M KA2 ML KA2 ML2 ML2 KA3 ML3 4