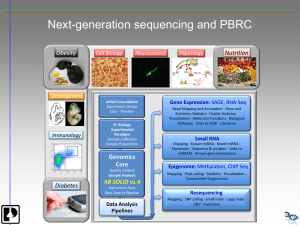
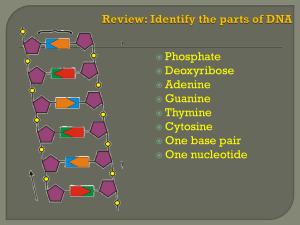
ppt presentation
advertisement

Epigenetics - RNA interference Discovery of RNA interference (1998) - silencing of gene expression with dsRNA Cenorhabditis elegans Antisense RNA (= RNA komplementární k mRNA) can silence gene expression (již počátek 80. let 20. století) - direct introduction of antisense RNA (or transcription in reverse orientation) - interaction of mRNA and antisense RNA, formation of dsRNA What is the mechanism behind? Original (!) hypotheses: - antisense RNA mechanically prevents translation - dsRNA is degraded (RNases) Cosuppression in Petunia Aim: increase expression of pigment-synthetizing enzym Result: loss of pigmentation in flower segments Napoli et al. 1990 Plant Cell 2:279–289 Cosuppression in Petunia Expression of antisense RNA was less efficient!!! Napoli et al. 1990 Plant Cell 2:279–289 Mechanism see later! What they get the Nobel prize for? - making proper controls pays off! - Introduction of even very small amount of dsRNA induce specific silencing (antisence RNA is less efficient!) dsRNA has to be a signal! - for sequence specific silencing Andrew J. Hamilton, David C. Baulcombe*(1999): A Species of Small Antisense RNA in Posttranscriptional Gene Silencing in Plants, Science 286 (5441): 950-952 RNA interference (RNAi) = silencing of gene expression mediated by small RNAs (small RNA, sRNA) in plants predominantly - 21-24 nt The precise role of 25-nt RNA in PTGS remains to be determined. However, because they are long enough to convey sequence specificity yet small enough to move through plasmodesmata, it is possible that they are components of the systemic signal and specificity determinants of PTGS (Hamilton and Baulcombe, 1999). RNA interference (RNAi) gene silencing at • transcriptional level (TGS) (transcriptional gene silencing) - induction of DNA methylation (mRNA not formed) • posttranscriptional level (PTGS) (posttranscriptional gene silencing) - transcript cleavage - block of translation Basic mechanism of RNAi dsRNA in cell is cleaved by RNase DICER into short dsRNA fragments – sRNA Argonaute with a single strand (from sRNA) mediates recognision of complementary sequences, which should be silenced (TGS, PTGS) Small RNA in plants - 3’ end of sRNA methylated (HEN1) - protection • miRNA (micro) – from transcipts of RNA Pol II (pre-miRNA) – hunderds MIR genes (in trans) pre-miRNA Wang et al. 2004 • siRNA (small interfering) – from dsRNA of various origin (both internal and external – thousands types (both in cis and in trans) ….. (+ piRNA in animals) Dicer-like - cleavage of dsRNA (21-24nt) - in Arabidopsis 4 paralogues (different functions) DCL1 – 21 nt miRNA (pre-miRNA) DCL3 – 24 nt siRNA (TGS - RdDM) DCL2, 4 – 21-22 nt siRNA (antiviral defence, secondary siRNAs) Argonaute RNA binding protein (20-26 nt RNA) - strand selection (5’ nt, participation of HSP90) - 10 genes in Arabidopsis - main component of RISC (RNA induced silencing complex) - block of translation or slicer (RNAse H-like endonuclease - PIWI doména) - role in TGS (RdDM) (RNA directed DNA methylation) Mechanism of small RNA action - overview Pol V PTGS (21-22 nt): - specific cleavage of transcript - block of translation TGS (24 nt): - methylation of promoter, heterochromatin formation - preventing interaction of transcription factors sRNA mode of action also depends on complementarity - incomplementarity in cleavage site prevents RNase activity dsRNA formation + secondary structures of viral RNAs (!) (MIR genes) ? mRNA fragment after Ago cleavage (secondarysiRNA ) • RdRP = RNA-dependent RNA Polymerase – synthesis of compl. RNA strand templates: - transcripts cleaved by RISC - impaired mRNAs (without polyA or cap) - transcripts of RNA polymerase IV RNA polymerases in RNAi (in plants) RNA-dependent RNA polymerases (RdRP, RDR) - form the majority siRNAs in plants RDR6: - dsRNA from impaired (not by Ago) RNAs (lacking polyA or cap) = primary siRNAs - dsRNA from products of Ago = secondary siRNAs - RDR1 – close paralogue involved in antiviral defense RDR2: - dsRNA from products of RNA polymerase IV (DNA-dependent) RNA polymerases in RNAi (in plants) DNA dependent RNA polymerase (Pol IV and Pol V) - related to RNA polymerase II (share common subunits) - specific for plants RNA polymerase IV - transcibes chromatin with H3K9me2 and non-met H3K4 (SHH1) - short transcripts (hundreds nt), with cap, without polyA - automatically used by RDR2 ( dsRNA DCL3 siRNA) RNA polymerase V - necessary for RNA-directed DNA methylation (RdDM) - direct interaction with Ago4 – identification of target seq. RNA polymeráza II – unclear, but important role RNA-directed DNA methylation (in detail) Pol IV a V – RNA polymerases RDR2 - RNA dep.RNA polymerase DCL3 – dicer-like protein AGO4 – ARGONAUTE DRM2 – de novo metyltransferase DRD1 – chromatin remodelling protein SHH1 – dual histon-code reading (H3K9me2, H3K4) SHH1 SHH1 Saze et al. 2012 Zhang et al. 2013 - methylation (by DRM2) occurs only under interaction of AGO4(6,9)-siRNA with RNA Pol V (large subunit C-term domain) and RNA-RNA complementarity RNA-directed DNA methylation – why so complicated and energy consuming? - de novo methylation of TE in new insertion sites - transmission of info from histons (CMT2/3) less reliable (less dense nucleosomes) - PTGS – TGS transition SHH1 SHH1 Zhang et al. 2013 Zemach et al. 2013 Secondary siRNA formation - target RNA (mRNA, TAS transcripts) - cleaved by Ago + primary sRNA (miRNA or siRNA) - RDR6 – complementary strand synthesis: dsRNA DCL2(4) secondary siRNA Function of secondary siRNA - signal amplification - formation of siRNA from neighbor seq. (transitivity – new targets) ta-siRNAs (miRNA na TAS) (trans-acting siRNA – widening of miRNA targets) Cosupression in Petunia - overexpression of pigment gene (enzyme for pigment synthesis) caused loss of pigmentation in flower sectors dsRNA - occurrence of aberrant transcripts due to overexpression - formation of dsRNA from aberrant transcripts by RdRP (RDR6) - formation of siRNAs that silence both transgene and internal gene Use in functional genomics – overexpression and knock-out possible with a single gene construct Paramutation interaction (in trans) between homologous alleles (epialleles), resulting in heritable change in gene expression of one allele (= epimutation) paramutagenic and paramutated allele Mechanism? mediator of paramutation Paramutation interaction (in trans) between homologous alleles (epialleles), resulting in heritable change in gene expression of one allele (= epimutation) 1.Paramutagenic allele 2. Paramutable allele - 1. methylated, inactive allele transcribed by Pol IV - siRNA formation - 2. induction of RdDM of complementary sequences mediator of paramutation MOP1 = ortholog RDR2 MOP2 = subunit of Pol IV, V (mediator of paramutation) RNAi pathways in plants - overview a + Inverted repeat b Natural antisense c d (DNA methylation) Molnár et al. 2011 + secondary siRNAs (theoretically from any RNA fragment primarily cleaved by Ago) Antiviral systemic resistance - newly developed leaves resistant against infection (1928) Mechanism? siRNA are transported - via plasmodesmata - through phloem - passive – diffusion or in direction of stream Spreading of silencing - GFP silencing with siRNAs (from vasculature) green = GFP fluorescence, red = chlorophyll autofluorescence Function of RNAi and systemic spreading of sRNA - antiviral defense – preventing of spreading of infection and new infection - TE defense – keeping genome integrity, heterochromatin structure - regulation of development – practically all phases - stress response (even heritable changes – see bellow) - regulation of nutrient uptake – e.g. phosphorus - epigenetic modulation of genetic information in meristems (heritable changes) - environmental adaptations Epigenetic regulation of gene expression in plant development - regulation mainly with miRNA (21 nt) (hundreds of MIR genes) - sometimes mediated with ta-siRNA (non-coding TAS transcript cleaved by RISC with miRNA, RDR6, …) - frequently transported between neighbour cells (non-cell autonomous) - targets – genes for regulatory proteins e.g. transcription factors, components of ubiquitination pathway, … (limited reliability of simple promoter fusion experiments with such genes!) Example: roles of miRNAs and ta-siRNAs in Arabidopsis leaf development Pulido A , Laufs P J. Exp. Bot. 2010;61:1277-1291 Modulation of RNAi regulation of gene expression in plant development Phenotypic changes caused by knock-out of a MIR gene Phenotypic changes caused by expression of miRNA-resistant variants of target genes (same protein encoded by different nucleotide sequence nonhomologous to miRNA) Parental imprinting - varying epigenetic modification of alleles inherited from father and mather (parental imprint) – it results in differencial expression of these alleles in zygote = alleles of some genes are methylated in one or the other gamet - evolved in mammals and angiospermous plants - hemizygote“ state can serve to ensure reasonable feeding of embryo (in mather body) – prevents evolution of paternal alleles causing excessive embryo feeding (in angiosperms – determination of endosperm size) - in plants – main function – repression of TE activity! Parental imprinting Mammals: imprint establishment connected with meiosis - extensive demethylation, specific methylation of some genes Plants: imprint establishes during development of gametophyte (haploid phase in plants: (polen) polen tube + embryo sac) demethylation in a part of gametophyte – genetically terminal lines - central cell of embryo sac (DME - DEMETER) - vegetative nucleus of polen tube (block of DDM1, DME) Epigenetic changes in microgametophyte Inhibition of methylation (repression DDM1) + actively (DME) In vegetative nucleus Primary function: induction of proper TE methylation in generative cells (sperm cells): Demethylation – reactivation of TE – siRNA formation – transport to spermatic cells – induction of proper TE methylation Epigenetic changes in megagametophyte Primary function: - block of TE and endosperm specific genes in egg cell - regulation of endosperm size - block of gametophytic developmental program? Repression of gene expression independent on methylation and siRNA: PcG (POLYCOMB GROUP) proteins - repression state kept primarily by di-(tri-)methylation of histon H3K27 - in vegetative Arabidopsis tissue – thousands genes! - Maintenance of H3K27me2 mediated by gene specific Polycomb repressive complex (PRC2) - keep histon methylation after replication Example: vernalization after vernalization VRN2 (VERNALIZATION) PcG protein binds with other proteins to promoter of FLC gene (repression by H3K27me2), expression of FLC blocks flowering - Repression of FLC allows formation of generative organs VIN3 – deacethylation of histons, VRN2 – induction of H3K27 methylation Vernalization – primary signal? VIN3 chromatin: - permanently H3K27me3 repressive mark (PRC2) - transcription induces activation mark H3K36me3 and acethylation (? Hypothetically low temperature induce disassembly of nucleosome in TSS?) - bivalent chromatin labeling ( animal stem cells) - quantitative response