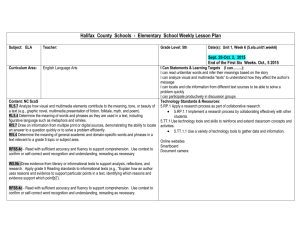
Explaining a paradox and predicting behaviour: GPCR Signalling Making Predictions
advertisement

Explaining a paradox and predicting behaviour: A dynamical systems model of the G protein GTPase cycle Benjamin Smith1, Claire Hill2, Louise Godfrey3, John Davey2 & Graham Ladds3 1MOAC DTC, Coventry House, 2Biological Sciences, 3Warwick Medical School, Gibbet Hill Road, University of Warwick, Coventry, CV4 7AL Biology Mathematics Combined Approach GPCR Signalling Simulating Experiments Making Predictions •G protein-coupled receptor (GPCR) signalling pathways regulate diverse cellular processes (Hepler, 1999). •The GTPase cycle of the G protein forms the central signalling unit of these pathways. •Regulator of G Protein Signalling (RGS) proteins regulate the response (Ross and Wilkie, 2000). •Model using a system of Ordinary Differential Equations. •A range of different initial ligand conditions → simulate dose response. •Altering parameter values and other initial conditions → simulate different strains. •With a mathematical model, it is possible to manipulate the system in any conceivable way. •It is therefore possible to make predictions about how the in vivo system will behave when the same manipulations are applied. •RGS proteins are important in both positive and negative signal regulation. Figure 1. Signalling through GPCRs. Receptor activation promotes guanine nucleotide exchange on Gα. “Active” (GTP-bound) Gα produces a downstream response through effector molecules. RGS protein “deactivates” Gα. β γ β γ Gα Gα GDP GTP GTP autohydrolysis GTP hydrolysis RGS Effector Effector Effector RGS Proteins •Accelerate GTP hydrolysis. •Increase the “switching off” rate of GTP-bound Gα. Hypothesis Development •Initial models included rapid recycling by the RGS protein (based on the model in Zhong et al., 2003). •They were unable to reproduce GTPase enhanced signalling. •We’ve seen that permanently GTP-bound Gα cannot produce maximal signalling. •Postulate an “inert” GTP-bound state. 1 Gα-GTP can only activate 1 effector molecule per round of signalling. What happens when we remove RGS protein from cells? Response Inert State Effectoractive Measuring Response •The experimental system is the pheromone response pathway in the yeast Schizosaccharomyces pombe. •Response is measured using reporter genes. •Cells are stimulated with increasing amounts of ligand to generate dose response curves (Figure 2). GTP Gα RGS Effectorinactive What happens when we increase the amount of RGS protein in cells? •A prediction was made using the mathematical model and the experiment was performed in vivo. Simulation Data 600 400 300 200 0 B RGS RGS + Pi RGSGαGTP RGS + Pi 30 ∆RGS 1xRGS 2xRGS 3xRGS 500 100 GαGTPEffector GTP RGSGα Biology Data 700 β -galactosidase activity GPCR* Response units Ligand -9 -8 -7 -6 -5 -4 25 ∆Rgs1 1xRgs1 2xRgs1 3xRgs1 20 15 10 5 0 B -9 -8 Log [Ligand] M GαGDP GαGTP Activation Figure 3. Schematic of the revised GTPase cycle hypothesis. This is the central mechanism of the mathematical model. -7 -6 -5 -4 Log [P factor] M Figure 6. Predicting the effects of increasing RGS expression in cells. Simulation predicts that doubling the amount of RGS protein in cells reduces sensitivity and further increases maximal signalling beyond wild-type levels. Trebling the amount of RGS is predicted to further reduce sensitivity, but also to reduce maximal signalling below that of wild-type. Results from in vivo assays show good agreement with these predictions. Conclusions 500 18 •RGS proteins reduce basal signalling but significantly enhance maximal signalling. RGS ∆RGS Rgs1 ∆Rgs1 400 16 Response Units β -galactosidase activity 20 14 12 10 8 6 •Current understanding of GTPase cycle insufficient. 300 •Biological investigation of the “model” organism (Sz. pombe) combined with computational investigation of a mathematical model. 200 100 4 •A new state for the GTP-bound Gα is postulated. 2 -6 -5 -4 0 Basal -9 -8 Explaining the paradox • RGS proteins regulate signalling negatively and positively. •Their role in G protein recycling is already known to be important (Doupnik et al., 1997). •Is recycling alone sufficient to explain GTPase enhanced signalling? Biology Data •The model was used to predict the effects of increasing RGS expression. •The GTPase accelerating effect of RGS proteins is responsible for negative and positive regulation of signalling through the Gα. Future Work •Perform parameter and principle component analysis. Simulation Data β -galactosidase activity time •Optimise parameters based upon a wide range of data. 500 40 35 30 25 0h 20 15 10 400 •Simulate a range of other strains and mutants. 300 200 100 5 0 0 B -9 -8 -7 -6 -5 B -4 -9 -8 -7 -6 -5 -4 Log[Ligand] Log [P factor] M 35 30 25 4h 20 15 10 400 300 •Apply theory to other guanine nucleotide binding proteins e.g. Ras, Rac, Rho. 200 100 5 0 0 B -9 -8 -7 -6 -5 B -4 -9 -8 -7 -6 -5 -4 Log[Ligand] Log [P factor] M References 500 40 35 30 25 8h 20 15 10 400 Doupnik,C.A., Davidson,N., Lester,H.A., and Kofuji,P. (1997). PNAS 94, 1046110466. 300 200 100 5 0 0 B -9 -8 -7 -6 -5 B -4 -9 -8 -7 -6 -5 -4 Log[Ligand] Log [P factor] M 500 40 35 30 25 16h 20 15 10 Hepler,J.R. (1999). Trends Pharmacol. Sci. 20, 376-382. Acknowledgements 300 200 100 0 -9 -8 -7 -6 Log [P factor] M -5 -4 Zhong,H., Wade,S.M., Woolf,P.J., Linderman,J.J., Traynor,J.R., and Neubig,R.R. (2003). J. Biol. Chem. 278, 7278-7284. Ross,E.M. and Wilkie,T.M. (2000). Annu. Rev. Biochem. 69, 795-827. 400 5 0 •Extend the model to consider the effects of translocation and post-translational modifications. •Extend to include multiple receptor, G protein and RGS species. 500 40 B Can mathematical modelling help? •Simulation of time-course and dose response data shows good agreement with in vivo data. -4 Time-course comparison β -galactosidase activity Maximum signalling is dependent upon rapid GTP hydrolysis. -5 •Comparing time-course data allows validation of model. β -galactosidase activity G protein Recycling • Removing the RGS protein from cells would be expected to increase signalling. •Paradoxically, basal signalling is higher but maximal signalling is lower (Figure 2). •RGS insensitive mutant of Gα behaves like ∆Rgs1 →effects are due to the change in hydrolysis rate. •Constitutively active mutant of Gα behaves like ∆Rgs1 →all Gα is GTP-bound but high levels of signalling are never achieved. •Gα turnover is necessary for high maximal response. -6 Figure 4. Simulated dose response curves. The two strains from Figure 2 were simulated: wild-type cells (RGS) and cells lacking an RGS protein (∆RGS). β -galactosidase activity Figure 2. Biological dose response curves. Wild-type cells (Rgs1), compared to that of cells lacking Rgs1 (∆Rgs1). -7 Log[Ligand] Response Units -7 Log [P factor] M Response Units -8 Response Units -9 Response Units 0 Basal B -9 -8 -7 -6 -5 -4 Log[Ligand] Figure 5. Biological time-course vs. simulated time-course. A comparison between the two shows good qualitative agreement. •Dr Matthew Hodgkin (Biological Sciences, University of Warwick). •Professor David Rand (Systems Biology DTC, University of Warwick) . •Molecular Organisation and Assembly in Cells (MOAC) DTC, University of Warwick.