Development of new microsatellite loci for the genus Polistes
advertisement

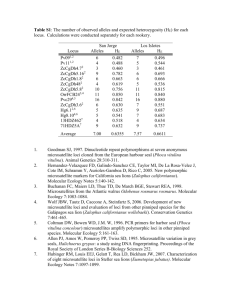
Insect. Soc. (2011) 58:581–585 DOI 10.1007/s00040-011-0164-z Insectes Sociaux TECHNICAL ARTICLE Development of new microsatellite loci for the genus Polistes from publicly available expressed sequence tag sequences M. T. Henshaw • A. L. Toth • T. J. Young Received: 21 January 2011 / Revised: 11 April 2011 / Accepted: 13 April 2011 / Published online: 24 April 2011 Ó International Union for the Study of Social Insects (IUSSI) 2011 Abstract Over the last 20 years, microsatellites have revolutionized the study of cooperation in the social insects. The Polistes paper wasps have been an important model system for investigations of cooperative behavior. Recently, an expressed sequence tag (EST) library has been developed for P. metricus, allowing researchers to investigate the genetic basis of cooperative behavior in primitive social insect societies for the first time. We searched these freely available EST sequences for microsatellite motifs. This represents a relatively new approach to the development of microsatellite loci that allows for the development of a greater number of loci at less expense. We designed 32 PCR primer pairs, of which 23 amplified PCR products and 18 were polymorphic. These loci exhibited high levels of polymorphism, comparable to anonymous loci isolated via screens of partial genomic libraries. Thus, they are appropriate for population genetic studies as well as the reconstruction of colony genetic structure. A screen of the entire EST database found a total of 708 di-, tri-, tetra- and penta-nucleotide repeats with large repeat units typical of polymorphic loci and at least 30 bp of flanking sequence for primer design. This pool of potential loci represents a new genetic tool for P. metricus, as well as Polistes more generally, as there is great promise for cross amplification in other species. Electronic supplementary material The online version of this article (doi:10.1007/s00040-011-0164-z) contains supplementary material, which is available to authorized users. M. T. Henshaw (&) T. J. Young Department of Biology, Grand Valley State University, 1 Campus Dr., Allendale, MI 49401, USA e-mail: henshawm@gvsu.edu A. L. Toth Department of Ecology, Evolution, and Organismal Biology, Iowa State University, 253 Bessey Hall, Ames, IA 50014, USA Keywords Polistes metricus Vespidae Polistinae EST-SSR Microsatellite loci Introduction Polistes paper wasps are a prominent model system for investigations of the evolution of cooperation in primitively eusocial societies (Reeve, 1991). Polistes wasps work cooperatively to build paper nests in which to rear brood, and the first females to mature each season typically become workers, caring for the offspring of the queen and defending the nest (Reeve, 1991). However, reproductive roles are more flexible in Polistes than in more advanced eusocial groups, such as ants or honeybees, and the workers are capable of reproduction should opportunities arise (Arevalo et al., 1998; Tibbetts, 2007). Thus, Polistes wasps are an ideal system in which to investigate the relationships between relatedness, conflict, and cooperation in societies where individuals who could reproduce independently choose to altruistically cooperate instead. Polistes metricus is a common paper wasp found throughout the eastern United States. It occurs as far north as Michigan, east and south to the Atlantic and gulf coasts respectively, and west to the Great Plains (Carpenter, 1996). This is an expansive range encompassing a wide variety of ecological contexts, and its wide distribution, as well as the diversity of habitats it occupies, makes P. metricus an interesting target for population genetic studies because of the potential for genetic differentiation associated with isolation by distance, or with distinct habitats or barriers. Studies characterizing patterns of genetic variation have become even more important because new genomic resources have recently been developed for P. metricus, including an EST (expressed sequence tag) library and a 123 582 microarray (Toth et al., 2007; Toth et al., 2010). As a result, P. metricus has emerged as an important model system for studies of the genetic basis of social behavior in primitively eusocial societies. However, these studies have been conducted at widely distributed sites (Hunt et al., 2007; Hunt et al., 2010; Toth et al., 2010), and it is important that we characterize genetic differences and similarities between populations so that the results of these studies can be compared. We have developed new polymorphic microsatellite loci for P. metricus, the first microsatellites developed in this species. Microsatellites can be used to estimate relatedness and to determine which individuals are reproductively active, allowing researchers to characterize queen number, mating frequency, and to determine the relative fitness consequences of cooperation and selfishness in these cooperative societies (Queller et al., 1993). Microsatellites are also commonly used to characterize genetic differences within populations. Thus, these loci represent a new genetic tool which can be used to characterize both colony-level and population genetic structure in P. metricus. We have identified new loci by searching publicly available EST sequence data for P. metricus. This relatively new approach to the development of microsatellite loci identifies more loci with fewer costs than previous approaches which require the construction of partial genomic libraries, enrichment for microsatellite repeats and sequencing of genomic fragments (reviewed in Ellis and Burke, 2007). As large-scale sequence databases become available for a broader array of species, especially species that are not classic genetic model systems, this approach will become increasingly important. Methods We identified microsatellite repeats within a previously developed P. metricus EST library (Toth et al., 2010; Toth et al., 2007), available from the NCBI Trace archive. We performed BLAST searches of assembled contigs in the ‘‘Old Polistes brain/abdomen contigs’’ database at http:// stan.cropsci.uiuc.edu/454/blast/waspblast.html for the following sequences: (AAC)5, (AAG)5, (AAT)5, (ACC)5, (CAG)5, (GAC)5, (CAT)5, (CCG)5. We retrieved 32 consensus sequences containing repeats (contigs can be downloaded from: ftp://stan.cropsci.uiuc.edu/download/Polistes/), and designed primers using the web-based program Web Primer (http://www.yeastgenome.org/cgi-bin/web-primer). We genotyped a total of eight P. metricus females collected from widely separated sites in the US states of Arkansas, Georgia, Ohio, Missouri, Tennessee, Texas and West Virginia. Genomic DNA was extracted from half a thorax from each wasp using a salt precipitation protocol 123 M. T. Henshaw et al. (Miller et al., 1988; Strassmann et al., 1996), and PCR products were amplified in 10 ll reactions (Final concentrations: 19 Colorless GoTaq reaction buffer with 1.5 mM MgCl2, 0.25 mM dNTP mix, 0.1 lM forward and reverse primers, 0.1 lM M13 primer labeled with TET, HEX or FAM, 0.35 units GoTaq DNA polymerase, unquantified diluted genomic DNA) with an annealing temperature of 50°C for all amplifications. Fragments were visualized at the University of Illinois Core Sequencing Facility on an ABI Prism 3730xl DNA analyzer and the sizes of fragments were scored using the computer program Peak Scanner (Applied Biosystems). The numbers of alleles (A), allele sizes, observed heterozygosities (Ho), expected heterozygosities (He), and tests for disequilibrium were calculated using the computer program GDA 1.1 (Lewis and Zaykin, 2001). Because these loci were isolated from an EST library, they are located within transcribed regions and are inherited as a unit with functionally important loci. As a result, they might be expected to exhibit unique characteristics due to the effects of selection. We compared the EST-derived loci isolated in this study to 19 anonymous loci which were previously isolated from the related species Polistes dominulus (Henshaw, 2000). Anonymous loci are isolated via probes of genomic libraries, and are unlikely to be closely linked to transcribed, functionally important regions. All statistical comparisons were performed using PASW 18 (SPSS, Inc.). To determine the potential for the development of additional loci from P. metricus EST sequences, we downloaded the entire EST database and searched for microsatellite repeats using the Tandem Repeats Finder algorithm (v. 4.0.4) in the Tandem Repeats Database (Yevgeniy et al., 2006). We used the default parameter settings (match, mismatch, indels = 2,7,7; minimum alignment score = 50) and eliminated redundant repeats with the Redundancy Elimination tool (90% overlap, prefer smaller unit sizes). We eliminated repeats with motif sizes \2 or [5 to retain only the motif sizes most commonly used when developing microsatellites. Finally, we only retained repeats with at least 30 bp of flanking sequence for primer design. Results Of the 32 primer pairs designed, 23 pairs successfully amplified products and 18 were polymorphic. Polymorphic loci exhibited from 2 to 8 alleles with expected heterozygosities ranging from 0.125 to 0.875 (Table 1). All polymorphic loci were in Hardy–Weinberg equilibrium (Table 1). While the range of heterozygosities was similar to that observed in the anonymous P. dominulus loci, the mean expected heterozygosity of 0.47 (polymorphic loci Development of new microsatellite loci 583 Table 1 Summary of the characteristics of 18 polymorphic microsatellite loci developed from EST sequences in Polistes metricus Locus Sequenced repeat A (size range, bp) Ho He HWE p value Pmet40472 (TTG)9 4 (166–175) 0.429 0.396 1.000 Primer sequences (50 ?30 ) F:TACTTGGCCTCTTCCCCAGTT R:TTGGAGACTTATTTTCCACCC Pmet41635 (TTG)7 6 (209–224) 0.500 0.542 0.605 Pmet42388 (GCT)5 2 (144–147) 0.000 0.233 0.067 F:TGTATTTGCCTAGGTTGCGA R:ATAGGAGGTAACCGTCCTGCA F:AACGACCCCCTTGAATGATT R:ACCTCGACGTCAACGTTGC Pmet42482 (TTG)9 3 (230–230) 0.125 0.342 0.065 F:TTATCCCCCCTCATCACCA R:AAACACCACCAGGACATTCTT Pmet43733 (GTC)6 2 (159–162) 0.125 0.125 1.000 Pmet43777 (ACC)5 2 (125–128) 0.125 0.125 1.000 Pmet44190 (TTG)6 2 (206–209) 0.250 0.233 1.000 F:TTTCGGTGTGTGCGACTACG R:ATGCAAAATGGTACTGCGGA F:GGCGAGTGTCAACACCTTTTT R:ATTCGCGAAAGAAATTAGGG F:TGTCCTGCGATAGAGGTCTTT R:TCGGGATAATGAAATTCTCGT Pmet44592 (AAG)10 4 (135–144) 0.750 0.692 0.283 F:TTGATCGATCGAGGAGACCAT R:CGACTAACATTCGAAGGAACA Pmet45195 (TTG)10 4 (217–226) 0.750 0.750 0.780 F:TGCTGCTTTATCGTATTTGGA R:GGACAGATGATGGCTCAAAA Pmet45548 (CCG)6 5 (131–143) 0.750 0.817 0.296 Pmet45730 (AAT)8 2 (146–149) 0.000 0.233 0.067 F:TCTTTTCGGCTTCCTCTTGT R:CGAAGGGACTTAGGAAAGTTG F:CGGATGGAATTCAAGTTCTCG R:CACACGCACATACCTTTACGA Pmet46215 (TTG)7 2 (222–228) 0.143 0.143 1.000 F:TTGTTCCAATCTCCATTCTTC R:TTCGAGGTCGAGATCAAAACA Pmet46480 (GTC)7 5 (146–174) 0.375 0.450 0.380 Pmet46483 (AAG)7 3 (111–117) 0.375 0.342 1.000 Pmet46588 (GAC)11 8 (186–216) 1.000 0.875 0.923 F:TCTCGTCATCTTCGTTATGCT R:TTTCACCACCACCACTACCA F:TCCGAACAACTTGTCCCACA R:AAGAAGAATTCGGTGATGACG F:CTATATCGTCATTTGCGTTGG R:ATTTGATGAACGCACAGGAG Pmet46597 (CAT)10 5 (156–177) 0.625 0.750 0.318 F:CTCATTGATTCGTTTGTGGCA R:TTTCGCTATGTTCTCTGATGA Pmet46789 (CAT)11 4 (235–244) 0.750 0.642 0.383 F:CAGCGATTTTCGCTTATTCTT R:CGATCAACGAAATATTTGGGG Pmet46823 (AAT)9 5 (130–143) 0.714 0.725 0.356 F:ATTTTGCTTTGCCCACCCTT R:TCGGATGTGCAATTGAACGA The locus name contains the contig number in the EST database only) was significantly lower than the mean of 0.68 observed in the anonymous loci (unequal variances t test, t = 2.825, p = 0.009). We observed a positive relationship between the number of uninterrupted repeats and the expected heterozygosity at P. metricus loci (Fig. 1; He = 0.114 9 Repeats - 0.474, R2 = 0.60, p \ 0.001) and this relationship was not significantly different (t = 1.863, p = 0.07) from that observed in the anonymous loci isolated from P. dominulus (Fig. 1; He = 0.082 9 Repeats - 0.230, R2 = 0.45, p = 0.002). While similar numbers of the EST-derived loci and the anonymous loci were monomorphic (5/23 vs. 7/19; Fisher’s exact test, p = 0.3227), a greater proportion of the polymorphic EST-derived loci exhibited low expected heterozygosities between 0.0 and 0.4 (9/18 vs. 0/12; Fisher’s exact test, p = 0.004). 123 584 M. T. Henshaw et al. Fig. 2 The distributions of the number of perfect repeats for 708 di-, tri-, tetra- and penta-nucleotide microsatellite loci identified in the P. metricus EST sequences Fig. 1 The relationship between the expected heterozygosity and the number of uninterrupted repeats in the sequenced allele for each of 23 newly developed P. metricus EST-linked microsatellite loci (closed circles) as well as previously published P. dominulus anonymous loci (open circles) (Henshaw, 2000). Regression lines are shown for P. metricus (solid line) and P. dominulus (dashed line) We found a total of 5,307 repeats in the EST database and 2,507 of these repeats had motif sizes between 2 and 5 bp. These microsatellite repeats were located in 2,422 unique contigs and 708 of them had at least 30 bp of flanking sequence in which to design primers. Of these, 91, 368, 161 and 88 were di-, tri-, tetra- and penta-nucleotide repeats, respectively. The 708 repeats we isolated exhibited high numbers of repeats, characteristic of polymorphic loci (Fig. 2). Discussion Despite being linked to transcribed regions, the loci developed from the P. metricus EST database exhibited high levels of polymorphism appropriate for studies of population genetic structure, as well as colony-level structure including the estimation of relatedness, sex determination, and parentage assignment. While the mean expected heterozygosity was lower than that observed in loci that were not isolated from EST sequences, the range of expected heterozygosities did not differ, and one could easily obtain loci exhibiting similar levels of polymorphism. Selective constraints might be expected to limit the number of polymorphic loci near transcribed regions, however, the EST-linked loci did not exhibit a greater 123 number of monomorphic loci than the anonymous loci isolated from a partial genomic library. All of the loci screened for this study were trinucleotide repeats, and selection may operate differently on other motif sizes due to frameshifts (Ellis and Burke, 2007). We found four times as many trinucleotide repeats (368 loci) as dinucleotide repeats (91 loci), suggesting that selection may constrain expansion at dinucleotide repeats to a greater extent. Though the loci isolated from EST sequences are not generally monomorphic at higher rates than the anonymous loci, they do more frequently exhibit low levels of polymorphism. Some of the EST-derived loci appear to be unconstrained, with many alleles, and high levels of polymorphism. However, other EST-derived loci with low levels of polymorphism might experience more constraints on expansion. While some variation might be acceptable at these loci, large expansions might be selected against (Batra et al., 2010; Ranum and Day, 2002), maintaining them at low levels of polymorphism. In contrast, the anonymous loci are either highly polymorphic or monomorphic. It seems that once the anonymous repeats begin to expand, there is little to prevent them from expanding further and quickly becoming highly polymorphic. With a pool of more than 700 potential loci to be screened, one could easily develop hundreds of loci for P. metricus, facilitating detailed population genetic studies as well as linkage mapping (Borrone et al., 2009). The 708 contigs that possess both microsatellite repeats and sufficient flanking sequences for primer design are available in FASTA format as an associated electronic resource (Online Resource 1). The linkage of these loci to the transcribed regions makes them particularly interesting markers because of the potential to detect the effects of selection via selective sweeps (Vigouroux et al., 2002). Such effects of Development of new microsatellite loci selection would be less apparent using previously available microsatellites unlinked to ESTs. Studies examining genetic structure with these loci could not only characterize patterns of genetic differentiation, they could also elucidate the nature of that structure at fine spatial scales (because of their number) and on a gene-by-gene basis (because of their linkage), especially as P. metricus genomic resources become better annotated. Thus, these loci add to the growing list of new genetic tools in P. metricus, enhancing its status as a new genetic model system that is also amenable to ecological and evolutionary studies in a natural setting. The P. metricus EST library should also prove to be a tremendous resource for the development of new microsatellite loci in other species of Polistes. Many previous studies have demonstrated that microsatellite loci isolated in one species of Polistes frequently cross amplify in others (e.g., Henshaw, 2000; Strassmann et al., 1997), retaining high levels of polymorphism, and EST-linked microsatellites have been shown to often be more transferrable across species (Ellis and Burke, 2007). Even if the rate of cross amplification was low for these loci, with such a large number of potential loci it seems likely that one could develop microsatellites for nearly any species of Polistes from the P. metricus EST sequences. We have employed a new and powerful approach to rapidly and inexpensively generate microsatellite loci based on existing EST sequences. Due to recent innovations in sequencing technology and reductions in the cost of largescale sequencing projects (Hudson, 2008), such EST databases are becoming increasingly common for a wide range of species. This application may thus be useful for researchers interested in estimating population structure and relatedness in natural populations of a wide diversity of organisms. Acknowledgments The authors thank The Hudson lab at the University of Illinois, Urbana-Champaign for early access to assembled contig sequences and the Statistical Consulting Center at Grand Valley State University for statistical advice. This work was supported by grant #IOS-0803317 from the National Science Foundation as well as grants from the Center for Scholarly and Creative Activity at Grand Valley State University. References Arevalo E., Strassmann J.E. and Queller D.C. 1998. Conflicts of interest in social insects: Male production in two species of Polistes. Evolution 52: 797-805 Batra R., Charizanis K. and Swanson M.S. 2010. Partners in crime: bidirectional transcription in unstable microsatellite disease. Hum. Mol. Gen. 19: R77-R82 Borrone J.W., Brown J.S., Tondo C.L., Mauro-Herrera M., Kuhn D.N., Violi H.A., Sautter R.T. and Schnell R.J. 2009. An EST-SSR- 585 based linkage map for Persea americana Mill. (avocado). Tree Genetics & Genomes 5: 553-560 Carpenter J.M. 1996. Distributional Checklist of Species of the Genus Polistes (Hymenoptera, Vespidae, Polistinae, Polistini). American Museum of Natural History, New York Ellis J.R. and Burke J.M. 2007. EST-SSRs as a resource for population genetic analyses. Heredity 99: 125-132 Henshaw M.T. 2000. Microsatellite loci for the social wasp Polistes dominulus and their application in other polistine wasps. Mol. Ecol. 9: 2155-2157 Hudson M.E. 2008. Sequencing breakthroughs for genomic ecology and evolutionary biology. Mol. Ecol. Res. 8: 3-17 Hunt J.H., Kensinger B.J., Kossuth J.A., Henshaw M.T., Norberg K., Wolschin F. and Amdam G.V. 2007. A diapause pathway underlies the gyne phenotype in Polistes wasps, revealing an evolutionary route to caste-containing insect societies. Proc. Natl Acad. Sci. USA 104: 14020-14025 Hunt J.H., Wolschin F., Henshaw M.T., Newman T.C., Toth A.L. and Amdam G.V. 2010. Differential gene expression and protein abundance evince ontogenetic bias toward castes in a primitively eusocial wasp. Plos One 5: e10674 Lewis P.O. and Zaykin D. 2001. Genetic Data Analysis: Version 1.0 (d16c). Free program distributed by the authors over the internet from http://lewis.eeb.uconn.edu/lewishome/software.html. Miller S.A., Dykes D.D. and Polesky H.F. 1988. A simple salting out proceedure for extracting DNA from human nucleated cells. Nucl. Acids Res. 16: 1215-1215 Queller D.C., Strassmann J.E. and Hughes C.R. 1993. Microsatellites and kinship. Trends Ecol. Evol. 8: 285-288 Ranum L.P.W. and Day J.W. 2002. Dominantly inherited, non-coding microsatellite expansion disorders. Curr. Opin. Genetics Dev. 12: 266-271 Reeve H.K. 1991. Polistes. In: The Social Biology of Wasps (Ross K.G. and Matthews R.W., Eds). Cornell University Press, Ithaca, pp 99-148 Strassmann J.E., Solı́s C.R., Peters J.M. and Queller D.C. 1996. Strategies for finding and using highly polymorphic DNA microsatellite loci for studies of genetic relatedness and pedigrees. In: Molecular Zoology: Advances, Strategies and Protocols (Ferraris J.D. and Palumbi S.R., Eds). Wiley-Liss, Inc., New York, pp 163180, 528-549 Strassmann J.E., Peters J.M., Barefield K., Solis C.R., Hughes C.R. and Queller D.C. 1997. Trinucleotide microsatellite loci and increased heterozygosity in cross-species applications in the social wasp, Polistes. Biochem. Gen. 35: 273-279 Tibbetts E.A. 2007. Dispersal decisions and predispersal behavior in Polistes paper wasp ‘workers’. Behav. Ecol. Sociobiol. 61: 1877-1883 Toth A.L., Varala K., Newman T.C., Miguez F.E., Hutchison S.K., Willoughby D.A., Simons J.F., Egholm M., Hunt J.H., Hudson M.E. and Robinson G.E. 2007. Wasp gene expression supports an evolutionary link between maternal behavior and eusociality. Science 318: 441-444 Toth A.L., Varala K., Henshaw M.T., Rodriguez-Zas S.L., Hudson M.E. and Robinson G.E. 2010. Brain transcriptomic analysis in paper wasps identifies genes associated with behaviour across social insect lineages. Proc. R. Soc. B - Biol. Sci. 277: 2139-2148 Vigouroux Y., McMullen M., Hittinger C.T., Houchins K., Schulz L., Kresovich S., Matsuoka Y. and Doebley J. 2002. Identifying genes of agronomic importance in maize by screening microsatellites for evidence of selection during domestication. Proc. Natl Acad. Sci. USA 99: 9650-9655 Yevgeniy G., Rodriguez A. and Benson G. 2006. TRDB - The Tandem Repeats Database. Nucl. Acids Res. 35 (suppl 1): D80-D87 123