Suffix tree - Wikipedia, the free encyclopedia
advertisement

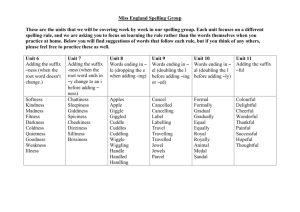
Lo g in / create acco unt Article Discussion Read Edit Search Suffix tree Fro m Wikipedia, the free encyclo pedia Main page Co ntents Featured co ntent Current events Rando m article Do nate to Wikipedia Interactio n Help Abo ut Wikipedia Co mmunity po rtal Recent changes Co ntact Wikipedia To o lbo x Print/expo rt Languages Česky Deutsch Français Bahasa Indo nesia Italiano ע ברית Lietuvių 日本語 Po lski Русский 中文 In computer science , a suf f ix t ree (also called PAT t ree or, in an earlier form, posit ion t ree) is a data structure that presents the suffixes of a given string in a way that allows for a particularly fast implementation of many important string operations. The suffix tree for a string S is a tree whose edges are labeled with strings, such that each suffix of S corresponds to exactly one path from the tree's root to a leaf. It is thus a radix tree (more specifically, a Patricia trie) for the suffixes of S . Constructing such a tree for the string S takes time and space linear in the length of S . Once constructed, several operations can be performed quickly, for instance locating a substring in S , locating a substring if a certain number of mistakes are allowed, locating matches for a regular expression pattern etc. Suffix trees also provided one of the first linear- time solutions for the longest common substring problem. These speedups come at a cost: storing a string's suffix tree typically requires significantly more space than storing the string itself. Co nt e nt s [hide] 1 Histo ry 2 Definitio n 3 Functio nality 4 Applicatio ns 5 Implementatio n 6 External co nstructio n Suffix tree for the string BANANA. Each substring is terminated with special character $. The six paths from the root to a leaf (shown as boxes) correspond to the six suffixes A$, NA$, ANA$, NANA$, ANANA$ and BANANA$. The numbers in the boxes give the start position of the corresponding suffix. Suffix links drawn dashed. 7 See also 8 References 9 External links History [edit] PDFmyURL.com 中文 The concept was first introduced as a position tree by Weiner in 1973 [1], which Donald Knuth subsequently characteriz ed as "Algorithm of the Year 1973". The construction was greatly simplified by McCreight in 1976 [2] , and also by Ukkonen in 1995[3][4]. Ukkonen provided the first linear- time online- construction of suffix trees, now known as Ukkonen's algorithm. Definition [edit] The suffix tree for the string S of length n is defined as a tree such that ( [5] page 90): the paths from the root to the leaves have a one- to- one relationship with the suffixes of S , edges spell non- empty strings, and all internal nodes (except perhaps the root) have at least two children. Since such a tree does not exist for all strings, S is padded with a terminal symbol not seen in the string (usually denoted $). This ensures that no suffix is a prefix of another, and that there will be n leaf nodes, one for each of the n suffixes of S . Since all internal nonroot nodes are branching, there can be at most n − 1 such nodes, and n + (n − 1) + 1 = 2n nodes in total (n leaves, n − 1 internal nodes, 1 root). Suf f ix links are a key feature for linear- time construction of the tree. In a complete suffix tree, all internal non- root nodes have a suffix link to another internal node. If the path from the root to a node spells the string χα , where χ is a single character and α is a string (possibly empty), it has a suffix link to the internal node representing α . See for example the suffix link from the node for ANA to the node for NA in the figure above. Suffix links are also used in some algorithms running on the tree. Functionality [edit] A suffix tree for a string S of length n can be built in Θ(n) time, if the letters come from an alphabet of integers in a polynomial range (in particular, this is true for constant- siz ed alphabets). [6] For larger alphabets, the running time is dominated by first sorting the letters to bring them into a range of siz e O(n); in general, this takes O(nlogn) time. The costs below are given under the assumption that the alphabet is constant. Assume that a suffix tree has been built for the string S of length n , or that a generalised suffix tree has been built for the set of strings of total length . You can: Search for strings: Check if a string P of length m is a substring in O(m) time ([5] page 92). Find the first occurrence of the patterns Find all z occurrences of the patterns of total length m as substrings in O(m) time. of total length m as substrings in O(m + z) time ([5] page 123). Search for a regular expression P in time expected sublinear in n ([7]). Find for each suffix of a pattern P , the length of the longest match between a prefix of time ([5] page 132). This is termed the mat ching st at ist ics for and a substring in D in Θ(m) P. PDFmyURL.com Find properties of the strings: Find the longest common substrings of the string S i and S j in Θ(n i + n j ) time ([5] page 125). Find all maximal pairs, maximal repeats or supermaximal repeats in Θ(n + z) time ([5] page 144). Find the Lempel–Ziv decomposition in Θ(n) time ([5] page 166). Find the longest repeated substrings in Θ(n) time. Find the most frequently occurring substrings of a minimum length in Θ(n) time. Find the shortest strings from Σ that do not occur in D, in O(n + z) time, if there are z such strings. Find the shortest substrings occurring only once in Θ(n) time. Find, for each i , the shortest substrings of S i not occurring elsewhere in D in Θ(n) time. The suffix tree can be prepared for constant time lowest common ancestor retrieval between nodes in Θ(n) time ([5] chapter 8). You can then also: Find the longest common prefix between the suffixes S i [p..n i ] and S j [q..n j ] in Θ(1) ([5] page 196). Search for a pattern P of length m with at most k mismatches in O(kn + z) time, where z is the number of hits ( [5] page 200). Find all z maximal palindromes in Θ(n)([5] page 198), or Θ(gn) time if gaps of length g are allowed, or Θ(kn) if k mismatches are allowed ([5] page 201). Find all z tandem repeats in O(nlogn + z), and k - mismatch tandem repeats in O(knlog(n / k) + z) ([5] page 204). Find the longest substrings common to at least k strings in D for Applications in Θ(n) time ([5] page 205). [edit] Suffix trees can be used to solve a large number of string problems that occur in text- editing, free- text search, computational biology and other application areas.[8] Primary applications include:[8] String search, in O(m) complexity, where m is the length of the sub- string (but with initial O(n) time required to build the suffix tree for the string) Finding the longest repeated substring Finding the longest common substring Finding the longest palindrome in a string Suffix trees are often used in bioinformatics applications, searching for patterns in DNA or protein sequences (which can be viewed as long strings of characters). The ability to search efficiently with mismatches might be considered their greatest strength. Suffix trees are also used in data compression; they can be used to find repeated data, and can be used for the sorting stage of the Burrows–Wheeler transform. Variants of the LZW compression schemes use suffix trees (LZSS). A suffix tree is also used in suffix tree clustering, a data clustering algorithm used in some search engines (first introduced in [9]). Implementation [edit] PDFmyURL.com If each node and edge can be represented in Θ(1) space, the entire tree can be represented in Θ(n) space. The total length of all the strings on all of the edges in the tree is O(n 2 ), but each edge can be stored as the position and length of a substring of S, giving a total space usage of Θ(n) computer words. The worst- case space usage of a suffix tree is seen with a fibonacci word, giving the full 2n nodes. An important choice when making a suffix tree implementation is the parent- child relationships between nodes. The most common is using linked lists called sibling list s. Each node has pointer to its first child, and to the next node in the child list it is a part of. Hash maps, sorted/unsorted arrays (with array doubling), and balanced search trees may also be used, giving different running time properties. We are interested in: The cost of finding the child on a given character. The cost of inserting a child. The cost of enlisting all children of a node (divided by the number of children in the table below). Let σ be the siz e of the alphabet. Then you have the following costs: Lookup Insert ion Traversal Sibling list s / unsort ed arrays O(σ) Θ(1) Θ(1) Θ(1) Θ(1) O(σ) Hash maps Balanced search t ree Sort ed arrays Hash maps + sibling list s O(logσ) O(logσ) O(1) O(logσ) O(σ) O(1) O(1) O(1) O(1) Note that the insertion cost is amortised, and that the costs for hashing are given perfect hashing. The large amount of information in each edge and node makes the suffix tree very expensive, consuming about ten to twenty times the memory siz e of the source text in good implementations. The suffix array reduces this requirement to a factor of four, and researchers have continued to find smaller indexing structures. External construction [edit] Suffix trees quickly outgrow the main memory on standard machines for sequence collections in the order of gigabytes. As such, their construction calls for external memory approaches. There are theoretical results for constructing suffix trees in external memory. The algorithm by Farach et al. [10] is theoretically optimal, with an I/Os complexity equal to that of sorting. However, as discussed for example in [11], the overall intricacy of this algorithm has prevented, so far, its practical implementation. On the other hand, there have been practical works for constructing disk- based suffix trees which scale to (few) GB/hours. The state of the art methods are TDD [12], TRELLIS [13] , DiGeST [14], and B 2 ST [15]. TDD and TRELLIS scale up to the entire human genome – approximately 3GB – resulting in a disk- based suffix tree of a siz e in the tens PDFmyURL.com of gigabytes [12], [13]. However, these methods cannot handle efficiently collections of sequences exceeding 3GB [14]. DiGeST performs significantly better and is able to handle collections of sequences in the order of 6GB in about 6 hours [14]. The source code and documentation for the latter is available from [16] . All these methods can efficiently build suffix trees for the case when the tree does not fit in main memory, but the input does. The most recent method, B 2 ST [15], scales to handle inputs that do not fit in main memory. See also [edit] Suffix array References [edit] 1. ^ P. Weiner (19 73). "Linear pattern matching algo rithm". 14th Annual IEEE Symposium on Switching and Automata Theory . pp. 1–11. do i:10 .110 9 /SWAT.19 73.13 . 2. ^ Edward M. McCreight (19 76 ). "A Space-Eco no mical Suffix Tree Co nstructio n Algo rithm" do i:10 .1145/3219 41.3219 46 . . Journal of the ACM 23 (2): 26 2–272. 3. ^ E. Ukko nen (19 9 5). "On-line co nstructio n o f suffix trees" . Algorithmica 14 (3): 249 –26 0 . do i:10 .10 0 7/BF0 120 6 331 . 4. ^ R. Giegerich and S. Kurtz (19 9 7). "Fro m Ukko nen to McCreight and Weiner: A Unifying View o f Linear-Time Suffix Tree Co nstructio n" Algorithmica 19 (3): 331–353. do i:10 .10 0 7/PL0 0 0 0 9 177 . . 5. ^ a b c d e f g h i j k l m n Gusfield, Dan (19 9 9 ) [19 9 7]. Algorithms on Strings, Trees and Sequences: Computer Science and Computational Biology. USA: Cambridge University Press. ISBN 0 -521-58 519 -8 . 6 . ^ Martin Farach (19 9 7). "Optimal suffix tree co nstructio n with large alphabets" on. pp. 137–143. . Foundations of Computer Science, 38th Annual Symposium 7. ^ Ricardo A. Baeza-Yates and Gasto n H. Go nnet (19 9 6 ). "Fast text searching fo r regular expressio ns o r auto mato n searching o n tries". Journal of the ACM (ACM Press) 4 3 (6 ): 9 15–9 36 . do i:10 .1145/2358 0 9 .2358 10 . 8 . ^ a b Alliso n, L.. "Suffix Trees" . Retrieved 20 0 8 -10 -14. 9 . ^ Oren Zamir and Oren Etzio ni (19 9 8 ). "Web do cument clustering: a feasibility demo nstratio n". SIGIR '98: Proceedings of the 21st annual international ACM SIGIR conference on Research and development in information retrieval . ACM. pp. 46 –54. 10 . ^ Martin Farach-Co lto n, Pao lo Ferragina, S. Muthukrishnan (20 0 0 ). "On the so rting-co mplexity o f suffix tree co nstructio n.". J. Acm 47(6) 4 7 (6 ): 9 8 7–10 11. do i:10 .1145/355541.355547 . 11. ^ Smyth, William (20 0 3). Computing Patterns in Strings . Addiso n-Wesley. 12. ^ a b Sandeep Tata, Richard A. Hankins, and Jignesh M. Patel (20 0 3). "Practical Suffix Tree Co nstructio n". VLDB '03: Proceedings of the 30th International Conference on Very Large Data Bases. Morgan Kaufmann. pp. 36–47. 13. ^ a b Benjarath Pho o phakdee and Mo hammed J. Zaki (20 0 7). "Geno me-scale disk-based suffix tree indexing". SIGMOD '07: Proceedings of the ACM SIGMOD International Co nference o n Management o f Data . ACM. pp. 833–844. 14. ^ a b c Marina Barsky, Ulrike Stege, Alex Tho mo , and Chris Upto n (20 0 8 ). "A new metho d fo r indexing geno mes using o n-disk suffix trees". CIKM '08: Proceedings of the 17th ACM Conference on Info rmatio n and Kno wledge Management. ACM. pp. 649–658. 15. ^ a b Marina Barsky, Ulrike Stege, Alex Tho mo , and Chris Upto n (20 0 9 ). "Suffix trees fo r very large geno mic sequences". CIKM '09: Proceedings of the 18th ACM Conference on Info rmatio n and Kno wledge Management. ACM. 16 . ^ "The disk-based suffix tree fo r pattern search in sequenced geno mes" . Retrieved 20 0 9 -10 -15. PDFmyURL.com External links [edit] This article's use of external links may not f ollow Wikipedia's policies or guidelines. Please improve this article by removing excessive and inappropriate external links. (August 2010) Suffix Trees by Dr. Sartaj Sahni (CISE Department Chair at University of Florida) Suffix Trees by Lloyd Allison NIST's Dictionary of Algorithms and Data Structures: Suffix Tree suffix_tree libstree , a generic suffix tree library written in C Tree::Suffix Strmat ANSI C implementation of a Suffix Tree , a Perl binding to libstree a faster generic suffix tree library written in C (uses arrays instead of linked lists) SuffixTree a Python binding to Strmat Universal Data Compression Based on the Burrows- Wheeler Transformation: Theory and Practice BWT Theory and Practice of Succinct Data Structures Practical Algorithm Template Library , application of suffix trees in the , C++ implementation of a compressed suffix tree] , a C++ library with suffix tree implementation on PATRICIA trie, by Roman S. Klyujkov v·d·e Tre e s in co m put e r scie nce B inary t re e s Se lf - b alancing b inary se arch t re e s B - t re e s Binary search tree (BST) · Cartesian tree · Top Tree · T- tree AA tree · AVL tree · Red- black tree · Scapegoat tree · Splay tree · Treap B+ tree · B*- tree · B x- tree · UB- tree · 2- 3 tree · 2- 3- 4 tree · (a,b)- tree · Dancing tree · Htree Trie s Suf f ix t re e · Radix tree · Ternary search tree B inary sp ace p art it io ning ( B SP) t re e s Quadtree · Octree · kd- tree (implicit) · VP- tree N o n- b inary t re e s Sp at ial d at a p art it io ning t re e s O t he r t re e s [hide] Exponential tree · Fusion tree · Interval tree · PQ tree · Range tree · SPQR tree · Van Emde Boas tree R- tree · R+ tree · R* tree · X- tree · M- tree · Segment tree · Fenwick tree · Hilbert R- tree Heap · Hash tree · Finger tree · Metric tree · Cover tree · BK- tree · Doubly- chained tree · iDistance · Link- cut tree Categories: Trees (structure) | Algorithms on strings | String data structures This page was last modified on 31 January 2011 at 13:55. Text is available under the Creative Commons Attribution- ShareAlike License ; additional terms may apply. See Terms of Use for details. Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc., a non- profit organiz ation. PDFmyURL.com Contact us Privacy policy About Wikipedia Disclaimers PDFmyURL.com