Student Protocol and Teacher Key
advertisement

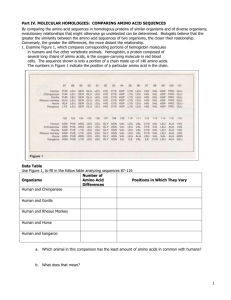
Biology Name: ___________________________ For grading: Staple Worksheets A and B to your lined paper. Submit as a packet. Procedure MOLECULAR SEQUENCES & PRIMATE EVOLUTION: AN AMINO ACID EXAMPLE In this lab, you will apply a bioinformatics tool to investigate the similarities and differences between hemoglobin molecules in various primate species. Remember that hemoglobin is the protein in red blood cells that carries oxygen. Hemoglobin is constructed from several tertiary level protein chains to make the quaternary level hemoglobin structure. The composition of the beta chain within hemoglobin is very similar across a variety of different species. In constructing cladistic trees and determining evolutionary relationships, biologists use hemoglobin structure as one measure of similarity or dissimilarity between different species. Materials: Procedure (in hand) Computer Data sheet (to be downloaded) BLAST search tool Worksheet A Worksheet B (to be obtained later) Essential Questions: How is macromolecular data used to determine potential evolutionary relationship and common ancestry? What tools might contemporary biologists use to conduct such an inquiry? Procedure: SECTION A – Building a Matrix From a Set of Sequences. This will help to process the raw sequence data into a format suitable for building a diagram showing relationships. A hemoglobin molecule consists of two protein chains: an alpha chain and a beta chain. There are 147 amino acids in the beta chain of hemoglobin. On the DATA SHEET, you will find the amino acids listed for the beta hemoglobin sequences in seven different species. You will use the BLAST search tool to compare these seven sequences to one another. The BLAST tool was developed to assist biologists in utilizing the growing amount of genetic data that genomics research has generated. With BLAST you can compare a sequence of nucleic acid bases or of amino acids to one another or to known genes and proteins. In this activity, you will use BLAST to compare the 7 sequences to one another. You will also then use BLAST to search for the beta hemoglobin of an 8th species and compare this to the other seven. At the top of Worksheet A, you will find a partially completed matrix: Biology Section A Matrix, Worksheet A: Differences Among Amino Acid Sequences. Compare each species with each of the other species and determine how many differences there are in the amino acid chains. You only need to complete the spaces in the upper portion of the matrix because the same numbers would go in the corresponding positions below the diagonal of “s’s” You will find an "x" in each redundant position below the diagonal. Each of the values in the matrix is the total number of positions where the amino acids differ between the two different species being compared. Using BLAST to compare the amino acid sequences. 1. Download the data sheet as directed. 2. Access the National Center for Biotechnology Information at http://www.ncbi.nlm.nih.gov/ 3. On the right hand side of the website, click on the link for BLAST. 4. On the lower left hand side of the BLAST site click on ‘protein blast’ Biology 5. In protein blast, click the box for ‘align 2 or more sequences’. 6. Two boxes should appear. In each box, cut and paste one of the two amino acid sequences you plan to compare. It is critical that you know which two species you are comparing at any given time, BLAST will not keep track of this for you. 7. Click the ‘BLAST’ button when you are ready to compare sequences. Be patient, as BLAST may take up to 60 seconds to process your request. Biology 8. Your results will look like this (when you scroll down). The ‘S’ value should be relatively higher and the ‘E’ value should be very low. The “query” and “subject” sequences are the two sequences you are comparing. The row in between these two shows similarities and places where the sequences differ. You will, of course, be counting the differences here. Section A 1. If you have not done so, compare each species to each of the other species. Count each spot where there is a difference in amino acid. Record this in the proper spot on the matrix. 2. Calculate and record the average value for each column. SECTION B – From Differences to Similarities Questions, Answer on another sheet of paper. Label as Section B please. 1. How many of the 147 amino acids in the beta chain of hemoglobin do the two most similar sequences share? How many do the two least similar sequences share? 2. The number of amino acid differences increases as the matrix moves to the left? What does this suggest about the biological relationships of each species to the species on its left? Biology SECTION C–Primate Similarities SPECIES ID: The first seven species in the data table and matrix are primates I: Human V: Rhesus Monkey (an Old World Monkey: OWM) II: Chimpanzee (a Great Ape) VI: Squirrel Monkey (a New World Monkey: NWM) III: Gorilla (a Great Ape) VII: Ring-tail lemur (a Prosimian) IV: Common Gibbon (a "Lesser" Ape) VIII: (to be revealed later) Section C Questions, Answer #2-4 on a separate piece of paper 1. On worksheet A, place these names (or abbreviations) to the left of their Roman numerals in the Species column in Matrix A, and above their numerals that run across the top of the matrix. 2. Which group of primates is least similar to the others? (Prosimians, Old World Monkeys, New-World Monkeys, Lesser Apes, Great Apes or Humans?) Are the differences between this least similar group and the other groups all about the same, OR do they gradually increase from bottom to top, suggesting a “Ladder of Progress?” 3. Are Gorillas more similar to Humans, or to Chimpanzees based on the sequence data? Which species are most similar to Gibbons on the sequence data? Chimps and Gorillas are Great Apes. 4. According to the data, should Humans be thought of as Great Apes as well? How could you explain these patterns in terms of common ancestry? Section D, Building a Cladistic Tree The working assumption for the classification scheme known as cladistics is that every group of organisms arose by branching off from a previous group. Each branch is called a clade. All the individuals in a clade share one or more carefully selected traits. Each trait must be identical or very similar within a clade, but appears to be modified (derived) from earlier (primitive, or ancestral) forms of the trait. Ideally, a cladistic tree follows the gradual accumulation of two or more traits (or their modifications) over time, showing a likely sequence of their evolution. Section D Questions. Answer on worksheet A 1. What two species have the fewest differences? Put these two species in the blanks indicating closest similarity (shortest branches, in the middle) on Cladistic Tree A (found on Worksheet A). 2. What third species is most similar to the first two species? Put it in the appropriate place on Tree A. Select the next most similar species and put this fourth species appropriately on the tree. 3. Now put the remaining species on Tree A. Biology SECTION E – Cladistic Trees and Evolutionary Relationships. Using BLAST to identify an unknown beta-hemoglobin sequence. Get WORKSHEET B with Cladistic Tree “B” and graph (at bottom) from your teacher. Cladistic Tree A has been rearranged to form Cladistic Tree B. This was done by simply rotating each branching point horizontally, starting at the lowest branching (1), then 2, 3, etc. 1. Attenpt to identify species VIII. Follow the steps in Part A above for using BLAST, except you will omit step 5. Cut and paste your sequence and clock the ‘BLAST’ button. When you scroll down to your results typically the top result with the lowest E value is the best match and thus identifies the species. Please fine a common name for this species. SECTION F – Introduction to Molecular Clocks. The dates (in Millions of Years Ago = MYA) next to the nodes (branching points) of Cladistic Tree B represent the divergence (branching) dates based on fossil evidence and radioisotope dating (not on molecular evidence). Section G Questions. Answer #1-3 on worksheet B, #4 on a separate piece of paper. 1. Fill in the average number of molecular differences (as “Changes”) for each node on Tree B on worksheet B (using your numbers from the appropriate columns in the matrix table “A” (Part “A” Matrix). 2. On the Graph of Amino Acid Sequence Differences vs. Time, plot the data for the relationship between the average number of amino acid differences (changes) and the time since divergence (age of the node). What is the general relationship between the time and average number of differences? NOTE: Species VIII in the data table and the matrix of differences diverged from the primate lineage around 90 million years ago. This explains the large number of differences. As you have likely identified species VIII, you have realized that it is not a primate, nor is it closely related to primates. 3. To Cladistic Tree B, add the branching line leading to species VIII, and add the changes and time to the left of the branching node. Add the appropriate data point to the graph of Differences vs Time. 4. On your lines paper, Summarize what you learned with this lesson. Include what it suggests (or confirms) about human evolution and references to common ancestors. Biology http://www.indiana.edu/~ensiweb/lessons/mol.wsa.pdf Biology http://www.indiana.edu/~ensiweb/lessons/mol.wsb.pdf Biology Teacher Key: Section B: 1. The two most similar sequences are identical, sharing all 147 amino acids (I and II, or the chimp and the human). The two least similar species, VII and VIII, share 114 of 127 amino acids (the lemur and the horse). 2. This would suggest that the biological relationship is more distant, as more time has passed since the two species shared a common ancestor and more mutations have accumulated to differentiate the species. Section C: 2. The prosimians are the least similar. The differences between the prosimians and all the other groups is about the same, suggesting NOT a “ladder of progress”, but rather that the prosimians diverged from the common ancestor of the other groups as a unique group farther back in the past. 3. Gorillas are equally different from both humans and chimpanzees. Humans and chimps are equally as similar to the gibbons. 4. According to the data, humans should be, biologically, categorized as Great Apes. Humans similarity to other Great Ape suggests a common ancestor in the more recent past. Biology http://www.indiana.edu/~ensiweb/lessons/mol.wsak.pdf Biology http://www.indiana.edu/~ensiweb/lessons/mol.wsbk.pdf