Supplementary Information (doc 2963K)

Supplementary Material for:
Linkage and Association Analysis of ADHD Endophenotypes in Extended and Multigenerational
Pedigrees from a Genetic Isolate
Claudio A. Mastronardi
1*
, Eva Pillai
1*
, David A. Pineda
2*
, Ariel F. Martinez
3
, Francisco Lopera
2
, Jorge I.
Velez
1
, Juan D. Palacio
2
, Hardip Patel
1
, Simon Easteal
4
, Maria T. Acosta
3
, F. Xavier Castellanos
5
,
Maximilian Muenke
3*
, and Mauricio Arcos-Burgos
1, 2, 3*
,
Affiliations:
1.
Genomics and Predictive Medicine, Genome Biology Department, John Curtin School of Medical
Research, ANU College of Medicine, Biology and Environment, The Australian National
University, Canberra, ACT, Australia;
2.
Group of Neurosciences, University of Antioquia, Medellin, Colombia;
3.
Medical Genetics Branch, National Human Genome Research Institute, National Institutes of
Health, Bethesda, Maryland, USA;
4.
National Center for Indigenous Genomics, Genome Biology Department, John Curtin School of
Medical Research, ANU College of Medicine, Biology and Environment, The Australian National
University, Canberra, ACT, Australia;
5.
The NYU Child Study Center at NYU Langone Medical Center, New York, NY, US, and Nathan
Kline Institute for Psychiatric Research, Orangeburg, NY, US.
* Contributed equally in the study
Correspondence to be addressed to:
Dr. Mauricio Arcos-Burgos
E-mail: Mauricio.Arcos-Burgos@anu.edu.au or
Dr. Maximilian Muenke
E-mail: mamuenke@mail.nih.gov
Non-Parametric Linkage Results For Endophenotypes
1) WISC Block Design
The greatest peak LOD score was found on chromosome 2, the position of the peak was 3.333cM from marker D2S1360 (LOD=2.51, p= 0.00034; Fig. 1A). The next highest score was at chromosome 12 at marker D12S398 (LOD=1.86, p=0.0017; Fig. 1G). Chromosome 14 peaked at
3.5cM from marker D14S1434 (LOD=1.75, p=0.0023; Fig. 1I). The peak for chromosome 10 was
1.666cM away from marker D10S1221 (LOD=1.73, p=0.0024; Fig. 1E). Chromosome 15 had a peak 1.333cM away from marker D15S131 (LOD=1.42, p=0.0053; Fig. 1J). Chromosome 11 peaked 3.333cM away from marker D11S4464 (LOD=1.41, p=0.0054; Fig. 1F). The peak at chromosome 19 was found at marker D19S1034 (LOD=1.41, p=0.0054; Fig. 1M). Chromosome 9 peaked at marker D9S938 (LOD= 1.33 and p=0.0067; Fig. 1D). Chromosome 3 peaked at marker
D3S1311 (LOD=1.25, p=0.0082; Fig. 1B). Chromosome 5 peaked 1cM away from marker
D5S1471 (LOD=1.11, p=0.0118; Fig. 1C). Chromosome 18 peaked at marker D18S542
(LOD=1.05, p=0.0141; Fig. 1L). Chromosome 16 peaked 4.5cM from marker D16S3103
(LOD=1.02, p=0.015; Fig. 1K).
A B
E
C
L
O
D
Sc or e
D
G
J
H
K
F
I
L
M
Marker Position (cM)
Figure 1S. Linkage and Association analysis of WISC Block Design for chromosomes with nominal linkage. A. chromosome 2; B. chromosome 3; C. chromosome 5; D. chromosome 9; E. chromosome 10;
F. chromosome 11; G. chromosome 12; H. chromosome 13; I. chromosome 14; J. chromosome 15; K. chromosome 16; L. chromosome 18; M. chromosome 19.
2) WISC PIQ
The greatest peak LOD score was found on chromosome 15, the position of the peak was at marker D15S131 (LOD=2.06, p= 0.00103; Fig. 2H). The next highest score was at chromosome
13 at marker D13S317 (LOD=2.01, p=0.00118; Fig. 2G). Chromosome 12 peaked at 2.333cM from marker C12S916 (LOD=1.78, p=0.0021; Fig. 2F). The peak for chromosome 2 was at marker D2S405 (LOD=1.68, p=0.0027; Fig. 2A). Chromosome 5 had a peak at marker D5S1471
(LOD=1.61, p=0.0032; Fig. 2D). Chromosome 11 peaked 1.167cM away from marker D3S2418
(LOD=1.34, p=0.0064; Fig. 2B). Chromosome 4 peaked at marker D4S391 (LOD=1.28, p=0.0076;
Fig. 2C). Chromosome 9 peaked at marker D9S9381 (LOD=1.21, p=0.0091; Fig. 2E).
Chromosome 18 peaked 3.667cM away from marker D18S851 (LOD=1.07, p=0.0132; Fig. 2I).
A B C
E F
L
O
D
Sc or e
D
G H I
J
Marker Position (cM)
Figure 2S. Linkage and Association analysis of WISC PIQ for chromosomes with nominal linkage. A. chromosome 2; B. chromosome 3; C. chromosome 4; D. chromosome 5; E. chromosome 9; F. chromosome 12; G. chromosome 13; H. chromosome 15; I. chromosome 18; J. chromosome 19.
3) WISC FSIQ
The greatest peak LOD score was found on chromosome 12, the position of the peak was at marker D12S1042 (LOD=2.05, p= 0.00106; Fig. 3C). The next highest score was at chromosome
19 at marker D19S591 (LOD=1.12, p=0.00116; Fig. 3D). Chromosome 3 peaked at marker
D3S2427 (LOD=1.06, p=0.0137; Fig. 3B). The peak for chromosome 2 was at 2.167cM away from marker D2S1394 (LOD=1.000, p=0.016; Fig. 3A).
A B
L
O
D
Sc or e
C D
Marker Position (cM)
Figure 3S. Linkage and Association analysis of WISC FSIQ for chromosomes with nominal linkage. A. chromosome 2; B. chromosome 3; C. chromosome 12; D. chromosome 19.
Quantitative Trait Loci (QTL) Linkage Analysis
Nominal linkage (LOD score ≥1) was found for the WISC Block Design, WISC Performance IQ (PIQ),
WISC Full Scale IQ (FSIQ), ACVT Correct Response test and ACVT Omissions test in chromosomes 2,
5, 7, 8, 12, 15, 16, 18, 19 and 20. A summary of the peak LOD scores, p-values, and closest marker to the peak is presented in Table 1S.
Table 1S. Summary of endophenotype peak LOD scores, p-values, peak LOD position, closest marker and marker position (cM) for linkage analysis of QTL.
Endophenotype
p-value
Peak LOD
Position
(cM)
Closest
Marker
Marker
Position
(cM)
WISC Block 2
WISC PIQ 2
5
15
16
18
20
WISC FSIQ 8
12
15
19
ACVT CR
ACVT
Omissions
7
7
1) WISC Block Design
1.03
1.17
1.21
1.27
1.08
1.31
1.13
1.04
1.04
1.02
1.14
1.12
1.12
0.015
0.01
0.009
0.008
0.013
0.007
0.011
0.014
0.014
0.02
0.011
0.012
0.012
38
0
130
60
23
75
80
125
36
36
0
19.833
19.833
D2S405 38
D2S2976
D5S1505
0
130
D15S131 60
D16S3103 23
D18S858
D20S451
75
80
D8S1179 125
D12S1042 36
D15S659
D19S591
D7S3051
36
0
18
D7S3051 18
Nominal linkage was found on chromosome 2; the position of the peak was at marker D2S405
(LOD=1.03, p= 0.015; Fig. 4S).
Chromosome 2
L
O
D
Sc or e
Marker Position (cM)
Figure 4S. Nominal linkage finding for QTL Analysis of WISC Block Design in chromosome 2.
2) WISC PIQ
The greatest peak LOD score was found on chromosome 18, the position of the peak was at marker D18S858 (LOD=1.31, p= 0.007; Fig. 5E). The next highest score was at chromosome 15 at marker D15S5131 (LOD=1.27, p=0.008; Fig. 5C). Chromosome 5 peaked at marker D5S1505
(LOD=1.21, p=0.009; Fig. 5B). The peak for chromosome 2 was at marker D2S2976 (LOD=1.17,
p=0.01; Fig. 5A). Chromosome 20 peaked at marker D20S451 (LOD=1.13, p=0.011; Fig. 5F).
Chromosome 16 peaked at marker D16S3103 (LOD=1.08, p=0.013; Fig. 5D).
A B C
L
O
D
Sc or e
D E F
Marker Position (cM)
Figure 5S. QTL Linkage Analysis of WISC PIQ with nominal findings in A: chromosome 2; B: chromosome 5; C: chromosome 15; D: chromosome 16; E: chromosome 18, F: chromosome 20.
3) WISC FSIQ
The greatest peak LOD score was found on chromosome 19, the position of the peak was at marker D19S591 (LOD=1.14, p= 0.011; Fig. 6D). The next highest scores were at chromosomes
8 and 12 at markers D8S1179 and D12S1042 respectively (LOD=1.04, p=0.014; Fig. 6A & 6B).
Chromosome 15 peaked at marker D15S659 (LOD=1.01, p=0.02; Fig. 6C).
A B
L
O
D
Sc or e
C D
Marker Position
(cM)
Figure 6S. QTL Linkage Analysis of WISC FSIQ with nominal findings in A: chromosome 8; B: chromosome 12; C: chromosome 15; D: chromosome 19.
4) ACVT Correct Response
Nominal linkage was found on chromosome 7; the position of the peak was at marker D7S3051
(LOD=1.12, p= 0.012; Fig. 7).
Chromosome 7
L
O
D
Sc or e
Marker Position (cM)
Figure 7S. Nominal linkage finding for QTL Analysis of ACVT Correct Response test in chromosome 7.
5) ACVT Omissions
Nominal linkage was found on chromosome 7; the position of the peak was at marker D7S3051
(LOD=1.12, p= 0.012; Fig. 8).
Chromosome 7
L
O
D
Sc or e
Marker Position (cM)
Figure 8S. Nominal linkage finding for QTL Analysis of ACVT Omissions test in chromosome 7.
Endophenotype
ACVT-CR
ACVT-O
ROCFT
ROCFT
WISC_Block-Design
WISC_Block-Design
ACVT-CR
WISC_Block-Design
ROCFT
ACVT-CR
WISC_PIQ
ACVT-CR
ACVT-O
ACVT-CR
ACVT-O
WISC_Block-Design
ROCFT
WISC_FSIQ
WISC_FSIQ
ACVT-CR
ACVT-O
ACVT-CR
ACVT-O
WISC_FSIQ
ROCFT
First
Marker rs2242373 rs2242373 rs333117 rs2301582 rs9683662 rs7212228 rs8078571 rs4790689 rs2242373 rs2253820 rs248161
Chr
17
17
17
17
4
17
17
17
17
17
5 rs4791641 rs4791641 rs3826517 rs3826517 rs2159112 rs4791473 rs7669283 rs2047200 rs335271 rs335271 rs1456862 rs1456862 rs861340 4 rs10521258 17
4
4
4
4
4
4
17
17
17
17
17
17
Position
8700981
8700981
4395169
5436262
62748383
12468792
6733671
4605199
8700981
8048168
135293448
8161148
8161148
4645323
4645323
12578120
11574958
62443714
62422529
62361458
62361458
62246327
62246327
69687963
14014193
8
8
8
8
6
6
8
4
8
7
5
7
7
7
7
8
5
3
3
6
8
6
6
8
8
#
Haplotypes
7
7
7
7
5
5
7
3
7
6
4
6
6
6
6
7
5
2
3
5
7
5
5
7
7
# Haps
Regressed
Full-Model
P-Value (FDR
Adjusted)
0.0005
0.0005
0.0023
0.0029
0.0031
0.0038
0.0045
0.0047
0.0049
0.0051
0.0051
0.0052
0.0052
0.0053
0.0053
0.0058
0.0063
0.0064
0.0064
0.0075
0.0075
0.0075
0.0075
0.0077
0.0083
R
Squared
0.0650
0.0650
0.0516
0.0476
0.0846
0.0925
0.0593
0.0693
0.0426
0.0433
0.0879
0.0430
0.0430
0.0445
0.0445
0.0874
0.0403
0.0532
0.0646
0.0393
0.0393
0.0393
0.0393
0.0996
0.0376
Mean Y
14.1105
1.8895
23.8680
23.9321
9.4946
9.4946
14.1056
9.4946
23.9321
14.1111
101.9839
14.1111
1.8889
14.1098
1.8902
9.4946
23.9321
103.8404
103.8404
14.1105
1.8895
14.1105
1.8895
103.8404
23.9321
WISC_FSIQ rs11131334 4 62457453 5 4 0.0090 0.0712 103.7166
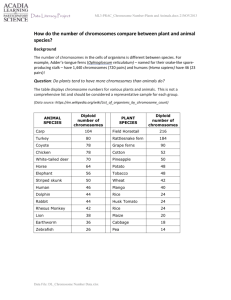
Table 2S. Haplotype Trend Regression Analyses for ADHD Endophenotypes. Haplotype analysis results for the top associated haplotypes to
ADHD endophenotypes in multigenerational pedigrees. P values for the whole LD block, after adjusting by FDR, are presented.