tpj12199-sup-0015-MethodS2
advertisement

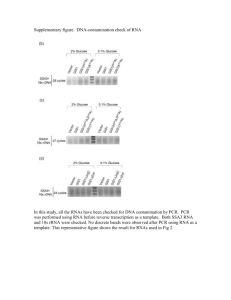
RNA isolation and transcript analysis for CAB and RBCS transcripts. After 3 days of stratification at 4°C, Col-0, mid-2 and cop1-4 seedlings of each biological replicate were grown for 4 days at 21°C in the dark in three sectors of a shared MS plates lacking sucrose. RNA was prepared using an RNeasy Plant Mini Kit (Qiagen, http://www.qiagen.com/). RNA quality was tested by photometric analysis (A260/A280 ratio between 1.9 and 2.1) and gave two distinct bands in formaldehyd agarose gel electrophoresis. Test-PCR with 35 cycles using EF1aA4 primers (Kirik et al., 2007) resulted in a band of 709 bp (cDNA) that indicated that the samples were devoid of DNA, while a sample of genomic DNA gave a clear band at 809 bp (genomic DNA). RNA was subjected to digestion with DNase I (Thermo Scientific, http://www.thermoscientific.com/fermentas) and subsequent reverse transcription was done using the Superscript III First strand System for RT-PCR including RNaseH treatment (Invitrogen, www.invitrogen.com). Transcript levels were determined by quantitative PCR using an Applied Biosystems 7300 real-time PCR system (http://www.appliedbiosystems.com/) analyzing triplicates of 1 µl cDNA in a 25 µl reaction containing POWER SYBR Green PCR Master Mix (Applied Biosystems, www.appliedbiosystems.com) and gene-specific primers for UBQ10, CAB and RBCS (Table S5). Dissociation curves were analyzed. Baseline and threshold were set manually according to the manufacturer`s instructions. Two biological replicates were used and each was analyzed in duplicate. The results were analyzed by the Ct method using UBQ10 for normalization (manufacturer`s instructions and Bookout and Mangelsdorf, 2003). Statistical analysis was performed by applying a one tailed Welch-test (Ruxton, 2006). Supplemental literature Bookout, A. L., Mangelsdorf, D. J. (2003) Quantitative real-time PCR protocol for analysis of nuclear receptor signaling pathways. Nucl Recept Signal, 1, e012.