Supplemental material - Springer Static Content Server
advertisement

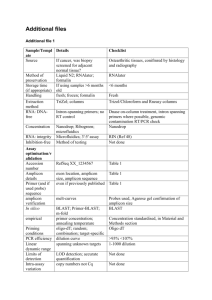
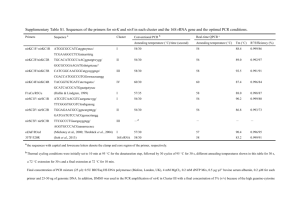
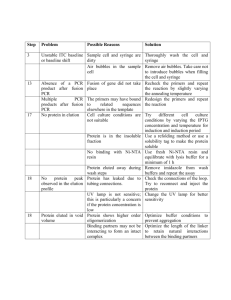
Electronic Supplementary Material Accreditation and Quality Assurance (ACQUAL), Springer, Title. Considerations for the development and application of control materials to improve metagenomic microbial community profiling Jim F. Huggett1*, Thomas Laver2, Sasithon Tamisak1, Gavin Nixon1, Denise O’Sullivan1, Ramnath Elaswarapu1, David J. Studholme2 and Carole A. Foy1 1 Molecular Biology, LGC, Queens Road, Teddington, TW11 0LY. 2 Geoffrey Pope Building, Biosciences, University of Exeter, Stocker Road, Exeter EX4 4QD. Supplementary material and methods Production of metagenomic control material ATCC purified bacterial genomic DNA (approximately 5 µg of each) was sourced from LGC Standards (detailed in ESM Table 1) and resuspended overnight at 4 °C on a tube rotator in 50 µL of TE pH 7.0 buffer (Ambion). Genomic DNA mass was estimated using the Qubit dsDNA BR Assay Kit (Life Technologies) and Qubit 2.0 Fluorometer (Life Technologies) by calculating the mean values from three separate dsDNA measurements. A 25 ng/µL mixture of metagenomic control material was prepared with defined bacterial proportion according to gram mass (ESM Table 1) and then incubated for 3 hours at 4 °C on a tube rotator. This stocks was diluted using TE pH 7.0 buffer to 1 ng/µl (25 µL aliquot) working stocks. PCR reaction Post- and pre-PCR adapter experiments were performed using 0.5 ng metagenomic control material (approximately 1.69X105 total genomic copies) as template in 250 ng of Human genomic DNA (Promega). No template DNA controls contained only 250 ng of Human genomic DNA (Promega). Post-PCR adapter experimental PCR reactions comprised 1X Fast EvaGreen® qPCR Master Mix (Biotium), 1X ROX dye, 300nM forward and reverse primers without adapters and Nuclease-Free water (Ambion) to a final volume of 25 μL (see Supplementary Table 2). PCR amplification was performed using the Applied Biosystems 7900HT Fast Real-Time PCR System (Applied Biosystems) with Absolute Quantitation run type and the following thermal cycling conditions: 96 oC for 120s, followed by 45 amplification cycles (96 oC for 30 s, 60 oC for 30 s, 72 oC for 30 s) and a dissociation stage (95 oC for 15 s, 60 oC for 15 s, 60 oC to 95 oC temperature ramp at 2 % ramp rate, 95 oC for 15 s). Data analysis was performed using Sequence Detection Software (SDS) version 2.4 with automatic baseline/threshold settings to estimate quantification cycle (Cq) and melting temperature (Tm) prior to NGS processing steps. Pre-PCR adapter experimental PCR reactions comprised 1.25 units of FastStart High Fidelity DNA polymerase (Roche), 1× FastStart buffer, 200 μM dNTP mixture, 400 nM forward and reverse fusion primers (ESM Table 2) and Nuclease-Free water (Ambion) to a final volume of 25 μL. PCR amplification was performed using a Gene Amp PCR system 9700 (Applied Biosystems) under the following the thermal cycling conditions recommended by the supplier: 94°C for 3 min, 35 amplification cycles (94°C for 15 s, 60°C for 45 s, 72°C for 60 s) and a final extension at 72°C for 8 min. Post- and pre-PCR adapter amplicons were characterised using the Agilent 2100 Bioanlyzer (Agilent) with the Agilent DNA 1000 Kit (Agilent) to determine amplicon size and purity prior to subsequent NGS processing steps (data not shown). Amplicon generation with fusion primers (Llib-A) Amplicons were produced for Lib-A protocol employing fusion primers (Primers A and B) designed to amplify distinct variable regions of 16s rRNA sequences. Two separate amplicons were produced for v2-3 (~520 bp) and v4-6 (~650 bp) regions respectively. PCR products, without purification, were analysed on 2100 Bioanalyzer (Agilent) for verifying the sizes. Amplicons were cleaned up by following a modified method consisting of three cycles of purification. The first round of clean up was performed following QIAquick PCR purification protocol, followed by 2x purification with AMPure beads (bead to DNA ratio of 1.6:1). After final purification, DNA was quantified using a Qubit 2.0 fluorometer and the concentration of each amplicon calculated in molecules/µl. emPCR reactions were set up as per Roche protocol and amplification was performed by employing recommended long-amplicon cycle protocol as described in 454 sequencing guidelines for amplicon protocol (March 2012). Ligated adapters amplicon sequencing (Lib-L) Amplicons were produced with gene-specific primers (without adapters) for v2-3 (~470 bp) and v4-6 regions (~600 bp). After fragment end repair, adapters were attached to the amplicon, using the kit ligation reaction, and purification was performed following the same protocol as described above for Lib-A products. emPCR was performed with Lib-L kit as per manufacturer's protocol with long amplicon emPCR cycle. 454 sequencing Both amplicons were mixed together in equimolar amounts prior to sequencing. Sequencing of mixed pre PCR or post PCR libraries was performed as per Roche protocol by employing 200 nucleotide cycles. ESM Table 1: Details of bacterial gDNA used to make the metagenomic control material Material ATCC Number* Genome Mol. Wt. size (MB) (Da) gDNA copy Percentage ng per Percentage 16S 16S copies Percentage Coefficient of number genomic µl Mass copies µl 16S variation per µl copy number Staphylococcus aureus BAA-1556Dsubsp. aureus 2.872769 1.8960E+09 4.00E-03 5 Rosenbach (MRSA) 0.80 % 1.27E+03 0.75 % 5 6.35E+03 0.78 % 4.00 % Staphylococcus aureus BAA-1718Dsubsp. aureus 2.872915 1.8549E+09 6.00E-02 12.00 % 5 Rosenbach (MSSA) 1.95E+04 11.52 % 5 9.74E+04 11.96 % 5.09 % Streptococcus pneumoniae 700669D-5 2.221315 1.4661E+09 1.28E-01 25.60 % 5.26E+04 31.09 % 4 2.10E+05 25.83 % 2.19 % Streptococcus pyogenes 700294D-5 1.852441 1.2226E+09 2.00E-02 4.00 % 9.85E+03 5.83 % 6 5.91E+04 7.26 % 3.49 % Streptococcus agalactiae BAA-611D-5 2.160267 1.4258E+09 7.50E-03 1.50 % 3.17E+03 1.87 % 7 2.22E+04 2.72 % 23.84 % Enterococcus faecalis 700802D-5 3.34194 2.2057E+09 5.00E-03 1.00 % 1.37E+03 0.81 % 4 5.46E+03 0.67 % 3.66 % Pseudomonas aeruginosa 47085D-5 6.264404 4.1345E+09 5.00E-03 1.00 % 7.28E+02 0.43 % 4 2.91E+03 0.36 % 9.17 % Klebsiella pneumoniae 700721D-5 subsp. Pneumoniae 5.31512 3.5080E+09 1.20E-01 24.00 % 2.06E+04 12.18 % 8 1.65E+05 20.24 % 4.34 % Acinetobacter baumannii 17978D-5 4.00075 2.6405E+09 5.00E-04 0.10 % 1.14E+02 0.07 % 5 5.70E+02 0.07 % 20.76 % Escherichia coli 700928D-5 5.231428 3.4527E+09 1.00E-02 2.00 % 1.74E+03 1.03 % 7 1.22E+04 1.50 % 20.43 % 2.194961 1.4487E+09 1.40E-01 28.00 % 5.82E+04 34.42 % 4 2.33E+05 28.60 % 13.65 % Neisseria meningitides 700532D-5 *ATCC purified gDNA materials sourced from LGC Standards, Teddington, UK ESM Table 2: List of 16S PCR primers, reagents and primer concentrations. Assay Primer type Primer sequence (5’-3’) 23_F no adapter No adapter AGHGGCGRACGGGTGA 23_R no adapter No adapter CGTATTACCGCGGCTGCT 456_F no adapter No adapter AGCAGCCGCGGTAATACG 456_R no adapter No adapter CATCTCACGACACGAGCTGAC 23_F PrimerA Fusion CGTATCGCCTCCCTCGCGCCATCAGAGHGGCGRACGGGTGA 23_R PrimerB Fusion CTATGCGCCTTGCCAGCCCGCTCAGCGTATTACCGCGGCTGCT 456_F PrimerA Fusion CGTATCGCCTCCCTCGCGCCATCAGAGCAGCCGCGGTAATACG 456_R PrimerB Fusion CTATGCGCCTTGCCAGCCCGCTCAGCATCTCACGACACGAGCTGAC Reagents Fast EvaGreen Master Mix (Biotium) Fast EvaGreen Master Mix (Biotium) FastStart High Fidelity PCR System (Roche) FastStart High Fidelity PCR System (Roche) Primer concentration 300 nM 300 nM 300 nM 300 nM 400 nM 400 nM 400 nM 400 nM Data analysis The sequence reads data were split into several subsets according to target amplicon; this was achieved based on matching to the respective PCR primer. A maximum of two mismatches to the PCR primer was allowed. Sequences generated from PCR chimeras were identified using ChimeraSlayer [1]. Reads were quality trimmed based on the average quality within a 50bp sliding window; sequences were trimmed when the average quality fell below 35. Sequences that were >10% longer than the expected length of the amplicon were removed, as were those shorter than 200 bases. Reads containing ambiguous base calls or homopolymers greater than 8 nt were also removed. These filtering steps were based on the high stringency pipeline used by the Human Microbiome Project Consortium [2]. For each amplicon dataset the reads were given a taxonomic assignment by performing a megaBLAST search [3] against a custom database made up of the 16S sequences for the species contained in the LGC MCM. The BLAST results were then processed using MEGAN [4], with default settings, to place the reads on the NCBI taxonomy based on the lowest common ancestor of the top BLAST hits to that read. The relative abundances of each species are based on MEGAN’s species level assignments, normalised by the species’ 16S copy number. In order to ascertain some measure of the robustness of our bioinformatics analysis with respect to error we employed a bootstrapping approach. We performed the same bioinformatics pipeline on each of 1000 randomly sampled subsets (with replacement), each comprising 10% of the total number of reads. We performed this bootstrapping approach for each of the filtered amplicon sequencing datasets (relative standard deviations shown in ESM table 3). ESM Table 3: Coefficient of Variation of the respective bioinformatic measurements based on bootstrapping (with replacement) of 10 % of the total reads. Material Pre PCR Post PCR Pre PCR Post PCR 23 23 456 456 N. meningitidis 4.17% 3.51% 3.03% 3.85% S. pneumoniae 62.39% 5.00% 2.88% 4.98% MSSA 2.81% 4.75% 5.22% 8.80% K. pneumoniae 5.81% 5.89% 96.85% 21.95% S. pyogenes 154.75% 20.73% 12.03% 20.49% S. agalactiae 300.18% 28.57% 18.70% 26.48% E. faecalis 32.69% 41.17% 44.08% 92.13% E. coli 30.06% 29.16% 281.73% 97.62% P. aeruginosa 54.40% 65.90% 99.78% 126.39% A. baumannii 212.35% 296.87% 312.46% 278.95% References 1. 2. 3. 4. Haas, B.J., et al., Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res, 2011. 21(3): p. 494-504. A framework for human microbiome research. Nature, 2012. 486(7402): p. 215-21. Morgulis, A., et al., Database indexing for production MegaBLAST searches. Bioinformatics, 2008. 24(16): p. 1757-64. Huson, D.H., et al., Integrative analysis of environmental sequences using MEGAN4. Genome Res, 2011. 21(9): p. 1552-60.