molecules buffer
advertisement

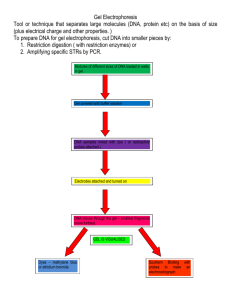
ELECTROPHORESIS Definition "Electrophoresis" refers to the electromotive force (EMF) that is used to move the molecules through the gel matrix. By placing the molecules in wells in the gel and applying an electric field, the molecules will move through the matrix at different rates determined largely by their mass when the charge to mass ratio (Z) of all species is uniform, toward the anode if negatively charged or toward the cathode if positively charged. It is one of the most popular ways to analyze macromolecules such as proteins, DNA and RNA. There are several types of electrophoresis, but the concepts are similar. The machine has an anode (positive charge) and a cathode (negative charge). Negative ions move toward the anode, and positive-charged ions move towards the cathode. The rate and distance traveled by these molecules help scientists classify and study different biomolecules. Several alternative types of gel electrophoresis exist and can be classed as one-dimensional such as agarose gels, polyacrylamide gel electrophoresis (native or SDS-PAGE) and Isoelectricfocusing (IEF) or two dimensional such as 2D-PAGE. Gel electrophoresis can also be carried out using narrow gauge capillaries. Types 1. Agarose Gels Agarose gels are an electrophoresis method to separate RNA and DNA molecules. It separates the molecules based on charge and size. DNA molecules are negatively charged, so they move through the gel quickly depending on size. Smaller DNA fragments move more quickly than larger ones due to friction resistance. 2. PAGE (Polyacrylamide Gel Electrophoresis) a) SDS-PAGE SDS-PAGE (Sodium Dodecyl Sulfate - Polyacrylamide Gel Electrophoresis) is a common form of electrophoresis for analyzing proteins. The SDS part of the name is a protein denaturing detergent that causes the molecule to unfold. The detergent binds to the polypeptide in a 1:1 ratio with each segment of the protein to give it a charge. The protein polypeptides move through the gel at different rates depending on mass, allowing researchers to study proteins based on size. b) Native-PAGE Native gels are similar to SDS-PAGE, except the detergent (SDS) is not used to denature proteins. Native gels are only able to separate proteins up to 2,000 kDa in size. Because the proteins are left folded, the dyes used are also different than SDS-PAGE. 3. Electrofocusing Electrofocusing takes advantage of charge and pH values of proteins. A container is filled with a gel solution that has an increasing pH gradient. The amino acids that form polypeptides have different acidic or basic charges. The protein travels through the gel, obtaining or losing protons depending on its charge. As the protein particle moves through the gel, it eventually becomes neutral and gets stuck in an isoelectric position. 4. 2-DIMENSIONAL GEL ELECTROPHORESIS Using this technique, proteins are separated by two different properties. Initially proteins or polypeptides are separated on the basis of their net charges by isoelectric focusing. Gels are then turned 90° and separation continues based on their molecular weight. Because it is unlikely that two molecules will be similar in both properties, molecules are more effectively separated in 2-D electrophoresis than in 1-D electrophoresis gels. In 2-D electrophoresis, proteins are initially separated by isoelectric focusing. This isoelectric focusing occurs across a pH gradient. The protein band will stop moving across the gel at its isoelectric point when the charge associated with the different amino groups is nullified by the pH. The gel is then turned 90o and the second electrophoresis occurs using SDS to separate the proteins by molecular weight. This second dimension of focusing gives a series of spots across the gel and each spot is a specific protein. Typical representation of a 2-D SDS-PAGE with proteins separated by charge and molecular weight. 5. Capillary Capillary electrophoresis is a method similar to SDS-PAGE. It separates molecules based on their charge and mass. Molecules are placed in rows called capillaries filled with conductive, electrolyte fluid. The analytes move in a speed relative to their charge and mass. This method is an older technique introduced in the 1960s that’s why SDS-PAGE is usually preferred in labs. Electroendosmosis What is Electroendosmosis (EEO)? A requirement of the gel is that it is neutral and that included the absence of charged impurities. If the gel contains any charged groups then an effect known as electroendosmosis will take place. As these charged groups are immobilised onto the gel they cause a solvent flow towards one of the electrodes, usually the cathode (negative) and thus in opposition to the sample flow. This is obviously undesirable as this will slow down and may distort the migration of the samples reducing resolution. Principle of Electroendosmosis The static support, the stabilizing medium (e.g. the gel) and/or the surface of the separation equipment such as glass plates, tubes or capillaries can carry charged groups: e.g. carboxylic groups in starch and agarose, sulfonic groups in agarose, silicium oxide on glass surfaces. These groups become ionized in basic and neutral buffers: in the electric field they will be attracted by the anode. As they are fixed in the matrix, they can not migrate. This results in a compensation by the counterflow of H3O+ ions towards the cathode: electroendosmosis. In gels, this effect is observed as a water flow towards the cathode, which carries the solubilized substances along. The electrophoretic and electroosmotic migrations are then additive. The results are: blurred zones, and drying of the gel in the anodal area of flatbed gels. When fixed groups are positively charged, the electroosmotic flow is directed towards the anode. Applications Gel electrophoresis is used in forensics, molecular biology, genetics, microbiology and biochemistry. The results can be analyzed quantitatively by visualizing the gel with I-IV light and a gel imaging device. The image is recorded with a computer operated camera, and the intensity of the band or spot of interest is measured and compared against standard or markers loaded on the same gel. The measurement and analysis are mostly done with specialized software. Depending on the type of analysis being performed, other techniques are often implemented in conjunction with the results of gel electrophoresis, providing a wide range of field-specific applications. DNA CHECK BY AGAROSE GEL ELECTROPHORESIS PRINCIPLE: Agarose gel electrophoresis separates DNA fragments according to their size. An electric current is used to move the DNA molecules across an agarose gel, which is a polysaccharide matrix that functions as a sieve to help "catch" the molecules as they are transported by the electric current. The phosphate molecules that make up the backbone of DNA molecules have a high negative charge. When DNA is placed on a field with an electric current, these negatively charged DNA molecules migrate toward the positive end of the field, which in this case is an agarose gel immersed in a buffer bath. The agarose gel is a cross-linked matrix i.e., a three-dimensional mesh or screen. The DNA molecules are pulled to the positive end by the current, but they encounter resistance from this agarose mesh. The smaller molecules are able to navigate the mesh faster than the larger ones. This is how agarose electrophoresis separates different DNA molecules according to their size. The gel is stained with ethidium bromide so as to visualize these DNA molecules resolved into bands along the gel. Ethidium bromide is an intercalating dye, which intercalate between the bases that are stacked in the center of the DNA helix. One ethidium bromide molecule binds to one base. As each dye molecule binds to the bases the helix is unwound to accommodate the stain from the dye. Closed circular DNA is constrained and cannot withstand as much twisting strain as can linear DNA, so circular DNA cannot bind as much dye as can linear DNA. Unknown DNA samples are typically run on the same gel with a "ladder." A ladder is a sample of DNA where the sizes of the bands are known. Unknown fragments are compared with the ladder fragments (size known) to determine the approximate size of the unknown DNA bands. Approximately 10ng is visible in a single band on a horizontal agarose gel. MATERIALS: •Agarose •TAE buffer •Gel casting tray, comb, power pack •Sample DNA •Loading dye •Sterile micro tips •EtBr staining solution •UV transilluminator or Gel Documentation System INSTRUCTIONS: For casting gel, agarose powder is mixed with electrophoresis buffer (TAE) to the desired concentration, and then heated in a microwave oven until completely melted. After cooling the solution to about 60°C, it is poured into a casting tray containing a comb and allowed to solidify at room temperature for nearly 45 min. After the gel has solidified, the comb is removed, using care not to rip the bottom of the wells. The gel, still in its plastic tray, is inserted horizontally into the electrophoresis chamber and just immersed with buffer (TAE). DNA samples mixed with loading buffer are then pipeted into the sample wells, the lid and power leads are placed on the apparatus, and a current is applied. The current flow is confirmed by observing bubbles coming off the electrodes. DNA will migrate towards the positive electrode, which is usually colored red. The distance DNA has migrated in the gel can be judged by visually monitoring migration of the tracking dyes. Bromophenol blue and xylene cyanol dyes migrate through agarose gels at roughly the same rate as double-stranded DNA fragments of 300 and 4000 bp, respectively. When adequate migration (2/3 of the gel) has occurred, DNA fragments are visualized by staining with ethidium bromide. This fluorescent dye intercalates between bases of DNA and RNA. It is often incorporated into the gel so that staining occurs during electrophoresis, but the gel can also be stained after electrophoresis by soaking in a dilute solution of ethidium bromide. To visualize DNA or RNA, the gel is placed on a ultraviolet transilluminator. Be aware that DNA will diffuse within the gel over time, and examination or photography should take place shortly after cessation of electrophoresis. Preparation of 1% Agarose gel: Weigh 0.50 g agarose; add in 50 ml 1X TAE and melt agarose in a microwave oven for 2-3 min. Cool down to about 45 to 50°C (bearable warmth) and pour into the gel platform with the comb in position. Running gel: After solidification of the gel (approx. 45 min), place the gel in a gel tank with 1 X TAE buffer. Buffer should be filled to the surface of the gel. Load the samples in the well and run the gel at 70 V till the blue dye runs to the end. Staining the gel: Prepare staining solution by adding 10 µl of 10 mg/ml stock of Ethidium bromide in 100 ml of Distilled water. Place the gel in staining solution for 30 min and view the gel in UV transilluminator. Fig 1: Agarose Gel Electrophoresis Method 10X TAE (pH 8.5): 1000 ml Tris-Base– 48.4 g EDTA – 7.44 g Glacial Acetic Acid – 11.42 mL Dissolve in 600 ml of dd water. First allow the Tris to dissolve in water, and then add EDTA. Make up the volume to one liter and autoclave. (Check and confirm the pH is about 8.5). Ethidium Bromide Stock: Stock 10 mg/ml. working concentration 1 µg/ml. a. Agarose Concentration: Agarose is a linear polymer that is derived from red algae. By using gels with different concentrations of agarose, one can resolve different sizes of DNA fragments. Higher concentrations of agarose facilitate separation of small DNAs, while low agarose concentrations allow resolution of larger DNAs. The image in the right shows migration of a set of DNA fragments in three concentrations of agarose, all of which were in the same gel tray and electrophoresed at the same voltage and for identical times. Notice how the larger fragments are much better resolved in the 0.7% gel, while the small fragments separated best in 1.5% agarose. The 1000 bp fragment is indicated in each lane. Concentration of agarose and its relation with DNA size can easily be understood from the following table. b. Voltage: As the voltage applied to a gel is increased, larger fragments migrate proportionally faster that small fragment. For that reason, the best resolution of fragments larger than about 2 kb is attained by applying no more than 5 volts per cm to the gel (the cm value is the distance between the two electrodes, not the length of the gel). b. Electrophoresis Buffer: Several different buffers have been recommended for electrophoresis of DNA. The most commonly used for duplex DNA are TrisborateEDTA (TBE = 90 mM Tris HCI, 20 mM boric acid, 2 mM EDTA, pH 8.3) and Trisacetate (TAE = 40 mM Tris HCI, 20 mM Na acetate, 1.8 mM EDTA, pH 7.8). DNA fragments will migrate at somewhat different rates in these two buffers due to differences in ionic strength. TAE is used for preparative separations when the gel is to be dissolved with NaCl solutions prior to binding of the DNA with glass powder. TBE is used for most analytical work. However for separation of large DNA fragments, electrophoresis at a low voltage gradient in TAE buffer is optimal, due to the low extent of diffusion of large molecules .Buffers not only establish a pH, but provide ions to support conductivity. If you mistakenly use water instead of buffer, there will be essentially no migration of DNA in the gel! Conversely, if you use concentrated buffer (e.g. a 10X stock solution), enough heat may be generated in the gel to melt it. c. Difference b/w TAE & TBE buffer i. Recovery of nucleic acid TAE has been prefavorably used so far because of the poor recovery of nucleic acids from TBE-gel. However, recent commercial recovery kits utilizing silica works well for either TAE-gel or TBE-gel. ii.Buffering Capacity: TBE has more buffering capacity than TAE iii. Moving Rate: Since ion strength of TBE is higher than TAE, solutes in TBE move faster than in TAE d. Effects of Ethidium Bromide: Ethidium bromide is a fluorescent dye that intercalates between bases of nucleic acids and allows very convenient detection of DNA fragments in gels, as shown by all the images on this page. As described above, it can be incorporated into agarose gels, or added to samples of DNA before loading to enable visualization of the fragments within the gel. As might be expected, binding of ethidium bromide to DNA alters its mass and rigidity, and therefore its mobility.



![Student Objectives [PA Standards]](http://s3.studylib.net/store/data/006630549_1-750e3ff6182968404793bd7a6bb8de86-300x300.png)




