emi412262-sup-0001-si
advertisement

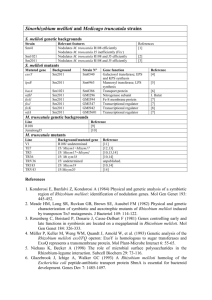
Supporting Information To monitor the invasion of mucoid cultures on agar by the expR mutants, a variety of strains and conditions were used (Fig. S1A). WT strains Sm2B3001, Rm41, and Rm8530 were repeatedly sub-cultured on TY agar using three different regimes. In experiment 1, bacteria were grown on TY agar containing 10 µg/ml of nalidixic acid and incubated at 30 oC for 3.5 day cycles. The medium and incubation temperature remained identical in experiment 2 but bacteria were incubated for 7 day cycles. Longer incubations decreased the NCC 50 (number of consecutive cultures needed until ≥50% of the colonies have a dry phenotype). In experiment 3, 20 µg/ml of gentamicin was used instead of nalidixic acid. This was because the wild type strains carried the plasmid pWBPwgeAmCh which contains mCherry fused to the promoter of the wge operon. This operon encodes multiple genes necessary for galactoglucan production (reviewed by Janczarek, 2011). Thus the wge promoter-mCherry fusion provides an additional indicator of galactoglucan production. The two phenotypes were easily distinguishable as colonies on agar: mucoid/red for the wild type and dry/white for the mutant (Fig. S1B). Furthermore, in experiment 3, bacteria were firstly incubated at 30oC for 2 days, followed by incubation at room temperature for 5 days. Additionally, strains Sm2B3001A (strain Rm2011 with a functional expR locus inserted into pSymA and Sm2B3001exoB (loss of EPS production) were included in experiment 3. Biological replicates were included in each experiment, e.g., represented as 8/10 for the strain Sm2B3001 in experiment 1. In experiment 3, NCC50 of strain Sm2B3001exoB could not be determined (n. d.) because the white mutant phenotype failed to dominate the culture by the 20th consecutive culture. Using one dry/white colony from each S. meliloti strain, restoration of the mucoid/red phenotype via the plasmid pPHUPexpRexpR was observed in each case. This is a low-copy plasmid carrying a functional expR controlled by its native promoter. The expR locus was sequenced for a collection of dry mutants. A map of these mutations is shown in Fig. S1C, and in more detail in Table S1. Also noteworthy is that the mutations which accumulated in our lab are comparable to those from other laboratories, previously reported in two dry phenotype S. meliloti strains Rm102F34 by Pellock et al in 2002 and GR4 by Martínez-Abarca et al in 2013 (see mutant 12 and 13, respectively, in Fig. S1C and Table S1). We also sequenced the expR gene in two other independently isolated S. meliloti strains with a dry phenotype, RU11 and L5.30, and found mutations (mutants 8 and 21, respectively). The ISRm1 element inserted in expR of strains Rm2011 and Rm1021 (Pellock et al., 2002) is included in Fig. S1C (mutant 17 and 18, respectively), however none of the new expR mutants contained an IS element. Figure S1 Loss-of-function mutation in expR is the cause of the non-mucoid phenotype. S1A, mucoid strains eventually turn dry upon serial sub-culturing. S1B, mutant phenotypes were complemented via a plasmid-borne copy of expR. S1C, amino acid sequence of ExpR and mapped mutations. Nterminal AHL-binding domain is in blue and C-terminal DNA binding domain in coral red. Locations of each mutation are indicated (underlined) and numbered. (See Table S1 for detailed description of the mutations.) S1D, mutations in expR correlates with loss of ExpR activity. A total of 10 variants of expR (Fig. S1D, numbers in left column correlate with those in Fig. S1C), each causing a single amino acid change, were selected for over-expression and purification via a His-tag, and tested for activity. Fig. S1D summarizes the results of these analyses (see Fig. S2A for data). S. meliloti strains with a dry phenotype were also analyzed for the presence of ExpR via Western Blot. A polyclonal rabbit-based anti-ExpR-peptide antibody (CASLO Laboratory ApS, Lyngby, Denmark) was used to probe for ExpR. The peptide sequence recognized by this antibody is RSDPHRKRMESMMVEA, located at approximately the middle of ExpR. A signal corresponding to ExpR was detected in each of the wild type parental strains Rm41, Rm8530 and Sm2B3001 (represented as WT in Fig. S1D), but not in the dry phenotype strain Rm2011. Only 3 ExpR-variants were detected via Western Blot: M107V, T215A and W218C (indicated by ‘+’ in Fig. S1D). All 10 expR variants were tagged with (his)6 and expressed in E. coli. Again, only the same 3 (M107V, T215A and W218C, indicated by ‘+’ in Fig. S1D) from 10 were sufficiently soluble for purification (Fig. S2B). The purified mutant proteins were tested for both their DNA binding activity, and transcription activity using the promoter regions of wgeA and sinI (Fig. S2C and S2D) as previously described (Bartels et al., 2007; McIntosh et al., 2009; Charoenpanich et al., 2013). Compared to the wild type, M107V bound to the wgeA and sinI promoter regions (Fig. S2C and S2D, respectively) with a low affinity (score of ‘++’, Fig. S1D), but did not activate transcription (Fig. S2E, S2F). In contrast, T215A and W218C were inactive (score of ‘-’) and weak (score of ‘+’), respectively, in both DNA binding and transcription activation. The severe defect in DNA binding activity of T215A and W218C is consistent with current knowledge of LuxR homologues, as reviewed by Nasser & Reverchon, 2007. The residue T215 is highly conserved among the LuxR homologues and interacts with DNA as part of the DNA-binding C-terminal domain. ExpRM107V is probably affected in its AHL binding, since the mutation is located only 3 residues away from an AHL interacting site of the ExpR homologue, TraR. Consistent with this hypothesis, M107V did not activate the promoter activities of wge and sinI. However, it did bind weakly to these promoters in an EMSA, as does wild-type ExpR in the absence of AHLs (see Fig. S2D). Table S1 List of expR sequence variants from dry/white phenotype mutants which invaded WT strains during the consecutive sub-culturing experiments described in Fig. S1. Each of the mutant variants is assigned a number from 1 to 36. This number is correlated with the location of the mutation in the expR sequence relative to the translation start. Each variation is described at the DNA and the amino acid level. Lastly, the parental strain is indicated. Of the 36 mutants listed, 30 spontaneously arose in our experiments. The other six (mutants 8, 12, 13, 17, 18, and 21) arose elsewhere. Figure S2 All ExpR variants are defect in activity. A, Western Blot analysis of ExpR and mutant derivatives in S. meliloti strains as indicated (3001, Sm2B3001; 2011, Rm2011). His-tagged proteins purified from E. coli (far right) were included as controls. B, His-tagged proteins from each step of purification from E. coli were collected and analyzed using SDS-PAGE. Lane are as follows: 1) uninduced whole cells, 2) induced whole cells, 3) crude extract, 4) 1st wash with 20 mM imidazole, 5) 3rd wash with 100 mM imidazole, and 6) eluate with 1 M imidazole. A239P represents one of the mutant variants which could be over-expressed but not purified. M107V and W218C represent mutant variants which could be purified. C & D, EMSA with purified (His)6-ExpR was used to test DNA binding activity. Labeled DNA included the promoter region of pstS as a negative control (upper panel, left), while the promoter region of wgeA (upper panel, right) and sinI (lower panel) were used to show binding activity. E & F, promoters of wgeA and sinI fused to egfp were used to test transcription activation by the ExpR variants. Promoter activity was induced by IPTG-induced ectopic expression of expR and variants in an expR/sinI double mutant strain (Sm2B4011) which lacks both a functional expR and the ability to produce AHLs. Only the WT ExpR and the W218C variant were able to activate promoters of wgeA and sinI in the presence of C16:1-HL. However, W218C showed a significantly lower activation. Error bars were calculated from four biological replicates. Figure S3 Growth curves of WT (Sm2B3001), expR mutant (Rm2011) and the exoB mutant (Sm2B3001∆exoB) grown in TY broth with shaking at 30°C. OD600 was measured at the time points indicated. The results are the average of three biological replicates. Error bars indicate standard deviation. Table S2 Bacterial strains and plasmids used in this work. References Bahlawane, C., Baumgarth, B., Serrania, J., Rüberg, S., and Becker, A. (2008) Fine-tuning of galactoglucan biosynthesis in Sinorhizobium meliloti by differential WggR (ExpG)-, PhoB-, and MucR-dependent regulation of two promoters. J Bacteriol 190: 3456-3466. Bartels, F.W., McIntosh, M., Fuhrmann, A., Metzendorf, C., Plattner, P., Sewald, N. et al. (2007) Effector-stimulated single molecule protein-DNA interactions of a quorum-sensing system in Sinorhizobium meliloti. Biophys J 92: 4391-4400. Brockwell, J., and Hely, F.W. (1966) Symbiotic characteristics of Rhizobium meliloti: an appraisal of the systematic treatment of nodulation and nitrogen fixation interactions between hosts and rhizobia of diverse origins. Australian Joicrnal of Agricultural Research 17: 885-899. Casadesús, J., and Olivares, J. (1979) Rough and fine linkage mapping of the Rhizobium meliloti chromosome. Mol Gen Genet 174: 203-209. Charoenpanich, P., Meyer, S., Becker, A., and McIntosh, M. (2013) Temporal expression program of quorum sensing-based transcription regulation in Sinorhizobium meliloti. J Bacteriol 195: 3224-3236. Ditta, G., Stanfield, S., Corbin, D., and Helinski, D.R. (1980) Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci U S A 77: 7347-7351. Dsouza, M., Larsen, N., and Overbeek, R. (1997) Searching for patterns in genomic data. Trends Genet 13: 497-498. García-Rodríguez, F.M., and Toro, N. (2000) Sinorhizobium meliloti nfe (nodulation formation efficiency) genes exhibit temporal and spatial expression patterns similar to those of genes involved in symbiotic nitrogen fixation. Mol Plant Microbe Interact 13: 583-591. Glazebrook, J., and Walker, G.C. (1989) A novel exopolysaccharide can function in place of the calcofluor-binding exopolysaccharide in nodulation of alfalfa by Rhizobium meliloti. Cell 56: 661-672. Green, M.R., and Sambrook, J. (2012) Molecular cloning : a laboratory manual. Cold Spring Harbor, N.Y.: Cold Spring Harbor Laboratory Press. Janczarek, M. (2011) Environmental signals and regulatory pathways that influence exopolysaccharide production in rhizobia. Int J Mol Sci 12: 7898-7933. Kowalski, M. (1970) Transducing phages of Rhizobium meliloti. Acta Microbiol Pol A 2: 109113. Martínez-Abarca, F., Martínez-Rodríguez, L., López-Contreras, J.A., Jiménez-Zurdo, J.I., and Toro, N. (2013) Complete Genome Sequence of the Alfalfa Symbiont Sinorhizobium/Ensifer meliloti Strain GR4. Genome Announc 1. McIntosh, M., Krol, E., and Becker, A. (2008) Competitive and cooperative effects in quorumsensing-regulated galactoglucan biosynthesis in Sinorhizobium meliloti. J Bacteriol 190: 53085317. McIntosh, M., Meyer, S., and Becker, A. (2009) Novel Sinorhizobium meliloti quorum sensing positive and negative regulatory feedback mechanisms respond to phosphate availability. Mol Microbiol 74: 1238-1256. Meade, H.M., Long, S.R., Ruvkun, G.B., Brown, S.E., and Ausubel, F.M. (1982) Physical and genetic characterization of symbiotic and auxotrophic mutants of Rhizobium meliloti induced by transposon Tn5 mutagenesis. J Bacteriol 149: 114-122. Nasser, W., and Reverchon, S. (2007) New insights into the regulatory mechanisms of the LuxR family of quorum sensing regulators. Anal Bioanal Chem 387: 381-390. Pellock, B.J., Teplitski, M., Boinay, R.P., Bauer, W.D., and Walker, G.C. (2002) A LuxR homolog controls production of symbiotically active extracellular polysaccharide II by Sinorhizobium meliloti. J Bacteriol 184: 5067-5076. Pleier, E., and Schmitt, R. (1991) Expression of two Rhizobium meliloti flagellin genes and their contribution to the complex filament structure. J Bacteriol 173: 2077-2085. Szende, K., and Ördögh, F. (1960) Die Lysogenie von Rhizobium meliloti. Naturwissenschaften 47: 404-405.