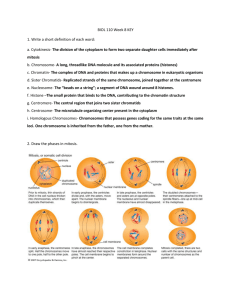
Chapter 13: Meiosis • Asexual reproduction: single individual is parent o Produces genetically identical offspring o Some multicellular organisms, but mainly single cellular • Sexual reproduction: two parents o Unites genes from both parents o Sperm cell and egg cell combine to form zygote o Greater genetic variation • Somatic cells: all body cells except gametes • Gametes: reproductive cells (sperm and egg) • Diploid (2n): two copies of each chromosome o Homologs: pairs of chromosomes o Tetrads: paired homologs o Somatic cells are diploid (46 chromosomes in humans) • Haploid (n): only one copy of each homologous chromosome o Gametes are haploid (23 chromosomes in humans) • Meiosis: diploid -> haploid, fertilization: haploid -> diploid • Sex chromosomes determine sex, autosomes are all the other chromosomes o In humans… XX is female, XY is male Meiosis • Mother cell is diploid somatic cells • 2 successive divisions: Meiosis I and Meiosis II • produces 4 haploid daughter gamete cells • DNA replicates in interphase • Cohesins: proteins that hold sister chromatids together after interphase • Meiosis I o Produces 2 haploid cells o Each cell has 1 chromosome, but 2 chromatids o Prophase I § Chromosomes condense § Homologs line up side by side § Cross over • Non-sister chromatids broken at chiasmata • Synaptonemal complex: holds homologs together • Synapsis: DNA breaks are repaired such that the DNA from one homolog is joined to the DNA from another • § Nuclear envelope breakdown § Spindle fiber formation o Metaphase I § Homologous chromosomes line up midway between poles o Anaphase I § Homologs separate, sister chromatids stay together o Telophase I § Chromosomes complete movement to poles § Nuclear envelope formation o Cytokinesis I § Cell divides Meiosis II o Produces 4 haploid cells (2 for each of the cells produced by meiosis I) o Each cell has 1 chromosome and 1 chromatid o Same as mitosis, but there is no DNA synthesis between MI and MII Mechanisms to Increase Genetic Variation • Independent assortment of chromosomes o Homologous pairs line up randomly during Metaphase I o Number of possible combinations is 2n where n = haploid number The linked image cannot be displayed. The file may have been moved, renamed, or deleted. Verify that the link points to the correct file and location. These two arrangements of chromosomes at metaphase I are equally probable. • Crossing Over o Produces recombinant chromosomes which combine DNA from each parent into a single chromosome • Random Fertilization o Any sperm can fuse with any egg to produce a zygote o Each zygote has a unique genetic identity Meiosis Worksheet Stage of Cell Cycle G1 Interphase S G2 What Happens Gap or Growth: Replace organelles etc. split during cell division Synthesis; Make an exact copy of DNA via replication Gap or Growth; Prepare materials for meiosis # of Chromosomes 46 46 46 Nuclear envelope breaks apart; Chromosomes Prophase I condense. Homologous chromosomes pair up 46 and crossing over occurs. Chromosomes line up in homologous pairs. Metaphase I 46 Independent assortment occurs. Meiosis I Homologous chromosomes separate. Homologous Anaphase I 46 Chromosomes Chromosomes move further apart; Nuclear Telophase I envelopes reform; and Chromosomes 46 decondense. Physicial cell division; Cytoplasm, and 23/cell Cytokinesis I organelles are divided into 2 cells. 46/ total Nuclear envelope breaks apart; Chromosomes 23/cell Prophase II condense 46/ total Chromosomes line up on the metaphase 23/cell Metaphase II plate, specifically here in a single file straight 46/ total line. Meiosis II Chromosomes separate, specifically here 46/cell Anaphase II Sister sister chromatids separate. 92/ total Chromataids Chromosomes move further apart; Nuclear 46/cell Telophase II envelopes reform; and Chromosomes 92/ total decondense. Physicial cell division of each cell; Cytoplasm, 23/cell Cytokinesis II and organelles are divided into 2 cells. 92/ total Meiosis begins with one cell and end with 4 cells. Each of the new cells have 1/2 the genetic information cell. Meiosis II occurs in each of the cells generated in Meiosis I. Meiosis occurs in sex cells. Replication occurs one time in meiosis during the S phase. Crossing over is the exchange of genetic material between homologous chromosomes. Independent assortment is random alignment of homologous chromosomes on the metaphase plate. Meiosis II is the "same" events as in mitosis. Mitosis vs. Meiosis Mitosis • Each daughter cell contains exact same genetic info as mother cell • DNA replicates in interphase Prophase • Chromosomes condense • Nuclear envelope breakdown • Spindle fiber formation Metaphase • Single line of chromosomes at center of cell Anaphase • Sister chromatids separate Telophase • Chromosomes complete movement to poles • Nuclear envelope formation Cytokinesis • Cell divides Meiosis I • Each daughter cell contains exactly half of the genetic information of the mother cell • DNA replicates in interphase Prophase I • Chromosomes condense • Homologs line up side by side • Sister chromatids cross over • Nuclear envelope breakdown • Spindle fiber formation • Lasts longer than in mitosis Metaphase I • Line of homologous pairs of chromosomes at center of cell Anaphase I • Homologs separate • Sister chromatids stay together Telophase II • Chromosomes complete movement to poles • Nuclear envelope formation Cytokinesis I • Cell divides Chapter 14: Mendel and the Gene Idea • Gregor Mendel: “father” of genetics o Studied in late 1850s o Largely unnoticed until rediscovered in early 20th c after his death • Character: feature that differs between individuals • Trait: differences in character • Gene: a heritable factor that causes a trait • True-breeding: all individuals are identical and self fertilization gives rise to offspring like parents, generation after generation (homozygous) • Mendel’s experiments o Studied inheritance of 7 characters that existed in 2 forms o Tracked 3 generations o Pea plants • Hybridize: produce offspring between genetically different strands • Monohybrid cross: mating following one character • Parental generation (P) • First filial generation (F1): offspring of P mating • Second filial generation (F2): offspring of F1 mating • Results of monohybrid crosses: o F1: all offspring display one trait (dominant trait) o F2: both traits are displayed in 3:1 ratio (dominant:recessive) • • • • • • Mendel’s explanation: o Each individual has two copies of each gene o Alleles: different variants of the gene o In an individual with both alleles, only the dominant allele is expressed o The recessive allele will only be expressed if both alleles are recessive Phenotype: physical attributes Genotype: genetic makeup Homozygous: both alleles are the same Heterozygous: two alleles are different Punnett square: diagram of outcomes of fertilization • • • • • • • • • • • • Probability = (# of outcomes with a particular result) / (# of possible outcomes) Mendel’s Law of Segregation (actually a theory not a law) o There are different variants of some heritable factor that causes a trait o Only one allele contributes to the character o Each gamete only has one allele o Each individual receives one allele from each parent Scientific theory: explanation of a phenomenon that cannot be observed directly Dihybrid cross: follows two characters Mendel’s Law of Independent Assortment o Genes for a pair of traits are distributed independently of one another when gametes are formed o This only applies if the genes are on separate chromosomes o Segregation of alleles for one gene does not affect segregation of alleles of a second gene Incomplete dominance: heterozygote has a phenotype intermediate between the homozygous phenotypes Codominance: both alleles are expressed Multiple alleles: gene has more than 2 alleles o An individual only has two alleles o There are more than two alleles in the population Epistasis: genotype at one gene affects phenotypic expression of a second gene Polygenic inheritance: multiple genes affect the same character have additive effects on phenotype Pedigree analysis: diagrams to follow traits through families using pedigrees Prenatal diagnosis: sample of fetal tissue from amniotic fluid or fetal portion of placenta analyzed for chromosomal defects or DNA mutations Chapter 15: Genes and Chromosomes • • • • • • • • • • • • • • Chromosome Theory of Inheritance (early 20th c): genes reside on chromosomes Law of Independent Assortment: alleles separate independently of each other during metaphase I Law of Segregation: 2 alleles separate during gamete formation such that each gamete has only one allele Linked genes: genes that are close together on the same chromosome o Don’t exhibit independent assortment Thomas Hunt Morgan o Studied fruit flies o Wildtype: individual with normal phenotype o Discovered sex linked traits: involve genes on sex chromosomes Sex chromosomes: X and Y determine sex of zygote o Females have XX § Barr body: inactivated X (all females have one) o Males have XY Autosomes: all other chromosomes You can measure the relative distances between genes on the same chromosome based on the percent recombination o 1% recombination = 1 map unit = 1 centimorgan genes that are very far apart on the same chromosome obey the law of independent assortment aneuploidy: too few or too many copies of a chromosome caused by nondisjunction o nondisjunction: error in chromosome distribution during meiosis deletion: portion of chromosome is missing duplication: portion of chromosome is repeated inversion: segment within chromosome is reversed translocation: segment of chromosome is moved to another non homologous chromosome DON’T HAVE TO KNOW HOW TO USE CHI SQUARE TEST!!! Chapter 16: the molecular basis of inheritance • Genes are made of DNA DNA Structure • Polynucleotides: chains of nucleotides • Sugar phosphate backbone with nitrogenous bases as appendages • Nucleotides bound by covalent bonds • Double helix: double stranded molecule that spirals around o Watson and Crick discover, win Nobel prize 1962 • Antiparallel strands: 5’ end of one strand bonds to 3’ end of other strand • Nitrogenous bases bound to each other by h bonds o A—T o G—C o Purine bonds to pyrimidine • Base pair: a pair of complementary nucleotides DNA Synthesis • Two strands separate and each strand acts as the template for synthesis of the complementary strand • • Results in 2 identical double-stranded molecules DNA polymerase: enzyme that adds nucleotides in DNA synthesis and proofreads for incorrect bases o Nucleotides only added at free 3’ end • Nucleotide that is added is a triphosphate, it loses two Pi molec. when it’s added making it a monophosphate • Replication fork: bubble where new nucleotides are added, expands until entire molecule is replicated • Helicase: binds at the replication fork and breaks h bonds between base pairs to make the DNA into two template strands Topoisomerase: protein that binds ahead of the replication fork and breaks covalent bonds in the sugar phosphate backbone to relieve over winding in the double helix Single strand binding proteins bind after the replication fork to prevent new h bonds from forming between the base pairs, this keeps the two parental strands separate so DNA can be synthesized on them Primase adds primer o DNA can’t act on a single strand alone; it needs a few nucleotides to already be present (these nucleotides are called the RNA primer) o The primer is later removed by DNA polymerase DNA ligase fills the gap where the RNA primer used to be and connects Okazaki fragments • • • • • • Leading strand: synthesized continuously Lagging strand: synthesized discontinuously o Okazaki fragments: short fragments of DNA on lagging strand • • • • • • • Excision: a nuclease enzyme recognizes and removes a mispaired nucleotide Repair: DNA polymerase fills in the gap using an undamaged strand as a template Circular chromosomes(bacteria) have one origin of replication Linear chromosomes (eukaryotes) have many origins of replication Each chromatid has one molecule of DNA o When DNA replication occurs during s phase, there are two chromatids and therefore two molecules of DNA o After mitosis each cell has only one chromatid and one molecule of DNA Chromatin: combination of DNA and nucleosomes o DNA is wound around nucleosomes like beads on a string o Histone: small basic proteins o Nucleosome: complex of 8 histones Telomere: end of a chromosome o Every time a DNA molecule is synthesized the end will get shorter by the amount equal to the length of the RNA primer Overview of DNA Synthesis Chapter 17: Gene Expression DNA---(transcription)---> RNA ---(translation)---> protein RNA Structure • Usually single stranded • Contains ribose sugar instead of deoxyribose sugar • Contains uracil nitrogenous base instead of thymine nitrogenous base Transcription • Transcription: synthesis of RNA from a DNA template o DNA strands are separated and RNA is synthesized complementary to one of the strands o RNA processing: RNA molecules are modified in the nucleus to produce the three types of RNA (mRNA, tRNA, and rRNA) • Transcription units: shorter segments that are transcribed (genes) • Spacer dna: dna between transcription units that is never transcribed • RNA polymerase: enzyme that carries out transcription o Reused over and over again to create multiple mRNA’s • Each gene has a start and stop signal o Promoter: dna sequence that is start signal (TATA box) o Terminator: dna sequence that is stop signal • Transcription is a lot like dna synthesis except the rna detaches from the template strand and the two dna strands rejoin • RNA processing o Capping: modified guanine nucleotide is added to 5’ end of RNA § Necessary for ribosome attachment and increases stability o Polyadenylation: 50-200 adenosine nucleotides added to 3’ end § Increases stability o RNA splicing: internal segments of RNA molec are removed § Intron: removed segments § Exon: retained segments (remember the exons are expressed) § Spliceosome: large complex of proteins and small RNAs where splicing occurs Translation • Translation: synthesis of polypeptide from an mRNA template in cytoplasm • Messenger (mRNA): produced by transcription o Read by rRNA to make protein o Template to synthesize polypeptide • • • • • • o Codon: 3 nucleotides that codes for 1 amino acid Ribosome: molecule that carries out translation o Composed of several proteins and 3 rRNA molecules o 2 subunits join together during translation Transfer (tRNA): adapter molecule in translation o Folded into lowercase t shape o Aminocyl tRNA synthase: enzyme that attaches amino acid to tRNA o Anticodon: 3 nucleotide region that base paires with the codon on mRNA during translation Codon: 3 nucleotide sequence in mRNA molecule o Indicates which amino acid should be added to a polypeptide o Multiple codons can code for the same amino acid o Often the third base in a codon doesn’t matter Initiation: ribosome begins translation at start codon (methionine) Elongation: ribosome moves along mRNA strand reading each codon, tRNA attaches amino acid for each codon onto chain Termination: ribosome reads a stop codon and detaches from the mRNA and polypeptide • Both transcription and translation require nrg Mutations • Substitution: one nucleotide is substituted for another o Missense mutation: change in codon causes change in amino acid o Nonsense mutation: codon changed to stop codon causing premature termination o Silent mutation: nucleotide change has no effect on amino acid sequence because the codon was changed to another codon that codes for the same amino acid • Frameshift: insertion of 1 or 2 bases causes all following codons to be read wrong • • • Mutagen: chemical or environmental condition that causes mutations Germline mutations: occur in gametes, passed on to next gen Somatic mutations: occur in body cells, maybe cancerous