Fragmentation of Protonated Peptide Ions via Interaction
advertisement

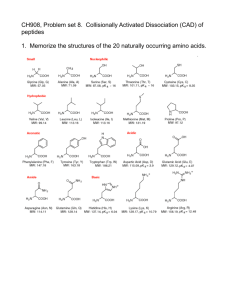
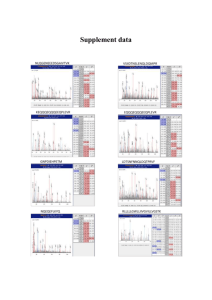
Fragmentation of Protonated Peptide Ions Group meeting Thursday, August 31, 2006 Outline Introduction on proteomics Use of mass spectrometry and different MS-MS techniques: •CID •ECD •Fragmentation patterns Details of an ETD MS-MS published recently Introduction Proteomics: The large-scale study of proteins, particularly their structures and functions. The entirety of proteins in existence in an organism throughout its life cycle are referred to as the proteome of the organism. Protein identification. Is one of the important branches of proteomics. Well-known methods are: 1-Edman degradation: low-throughput sequencing, limited to peptides of 50-60 amino acids 2-Mass spectrometry: Higher-throughput, can identify much larger peptides and even proteins Proteomics vs. genomics In the human genome there are ~22,000 genes vs. ~400,000 proteins. This makes proteomics more complex than genomics. The large increase in protein diversity is thought to be due to alternative splicing (mRNA change) and post-translational modification of proteins. Protein diversity can not be fully characterized by gene expression analysis alone, making proteomics a useful tool for characterizing cells and tissues of interest. http://www.scientificpsychic.com Mass spectrometry in proteomics Protein or peptide is identified from its molecular weight and the mass to charge value of its fragments Tandem mass spectrometry (MS-MS) is one of the most important methods for protein and peptide identification High sensitivity (femtomole level, 10-15) and specificity Collision induced dissociation (CID) of protonated peptide cations is the most common method for fragmentation Electron capture dissociation (ECD) is also used as a complement of CID Collision Induced Dissocation (CID) Multiple collisions of gas molecules with peptide. The kinetic energy of neutral gas molecule transfers to the peptide, it will rapidly distribute all over the covalent bonds in the molecule and this will lead to amide bond breakage. Cleavage of the peptide amide bond to produce b and y-type fragment ions. Incomplete backbone fragmentation and loss of labile groups that are very important in posttranslational modifications Q1 http://www.city.sapporo.jp Q2 Q3 Peptide structure Ca http://courses.cm.utexas.edu/emarcotte/ch339k/fall2005/Lecture-Ch3-1/Slide18.JPG Fragmentation nomenclature http://www.matrixscience.com/help/fragmentation_help.html Electron Capture Dissociation (ECD ) Capture of a thermal electron by a protonated peptide causes the peptide fragmentation from N-Ca bond. This will produce N-terminal c-fragments and C-terminal z-type fragment ions. Unlike CID this is nonergodic, meaning that the fragmentation happens before the energy transfer within the molecule. As a result labile modification groups will remain intact. The maximum cross section is for thermal electrons: Only successful in FT-ICR mass spectrometry. (strong RF electric field of quadruple and ion trap makes the introduction of such electrons difficult) Fragmentation of Protonated Peptide Ions via Interaction with Metastable Atoms Vadym D. Berkout MassTech, Inc., Columbia, Maryland Anal. Chem.2006, 78,3055-3061 Electron Transfer Dissociation (ETD) A negative or neutral atom will transfer the electron to peptide for fragmentation M n+ + Ar* M (n-1)+ + Ar + Fragmentation Glow discharge a source for metastable argon 25 Torr 5 mTorr GD source Asymmetric electric field: better separation of metastable atoms from ions. Cathode: 1.5 mm diameter, oxygen free copper rod Anode: off-axis planer Voltage = 300V, I = 5mA Two lenses plus a nickel grid are charged with few hundred volts negative potential to prevent any electron entrance into the quadrupole region. Octopole Ion guide Quadrupole Ion guide Only metastable argon? M + Ar .+ M .+ + Ar charge exchange ionization (15.7 eV =1520kJ/mol) M + Ar* M .+ + Ar + e Penning ionization (11.72 eV and 11.55 eV) Ionization energy for different gases (eV): Nitrogen: 15.58 Methane: 12.61 Ammonia: 10.07 Isobutene: 10.68 15.58 10.07 12.61 10.68 Argon is mainly neutral and the small amount of argon ions will be ejected from quadrupole by setting the low mass cut off at m / z =250 Fragmentation of peptides in trapping mode Peptide ions produced by electrospray were trapped in quadrupole for 100-400 ms Substance P: 11-amino acid polypeptide with the sequence: Arg Pro Lys Pro Gln Gln Phe Phe Gly Leu Met NH2. In central nervous system The regulation of mood disorders, anxiety, stress, reinforcement, neurogenesis, respiratory rhythm, neurotoxicity, nausea / emesis and pain Substance P Lots of c fragments Similar to ECD Bradykinin: Small peptide with 9 amino acids. arg - pro - pro - gly - phe - ser - pro - phe - arg Bradykinin is a potent endothelium-dependent vasodilator, causes contraction of non-vascular smooth muscle, increases vascular permeability and also is involved in the mechanism of pain. In some aspects, it has similar actions to that of histamine, and like histamine is released from venules rather than arterioles. Bradykinin complete c and z fragments Except c1 (same as ECD) Fibrinopeptide A A peptide released as part of the clotting process. Mw = 1536.6 Thrombin clips two peptides (fibrinopeptide A and B )from fibrinogen. This produces the fibrin monomer with exposed polymerization sites that can bind to other fibrin monomers. The monomers polymerize to form a loose clot http://www.hematology.org Fibrinopeptide A Lots of b and y fragments in addition to c and z fragments Potential energy diagram for interaction of peptide cation with metastable electronically excited atom. Dependence of cross section of the electron transfer on the collision energy. It shows that metastable argon atoms can be a source of electron for peptide dissociation. This allows ETD in Ion trap and quadrupole as a substitute of ECD in FTMS. Other refrences: http://www.answers.com http://en.wikipedia.org http://www.mcb.harvard.edu/DonWiley