The Biology of Small Populations: Genetics
advertisement

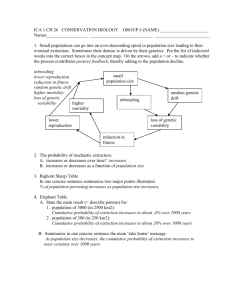
The Biology of Small Populations: Genetics Introduction In the last lecture we identified four categories of stochastic threats to small populations.1 So far we have focused on those stochastic threats that directly affect the numbers of individuals in a population. 1. Demographic stochasticity — not much of a problem in populations with more than 50–100 reproductive individuals 2. Environmental stochasticity — requires large populations, on the order of 1000-10,000 individuals to give a reasonable chance of long-term survival. 3. Demographic heterogeneity — not clear how large populations need to be to buffer its effects, probably at least as large as for environmental stochasticity. 4. Natural catastrophes — if catastrophes occur at an appreciable frequency and if they eliminate a large fraction of the population, no single population can survive over the long term. Before starting our discussion of how to use what we’ve learned in the management of threatened species, we have one more stochastic threat to discuss: genetic stochasticity or genetic drift. As I mentioned in my introductory lecture, the early literature in conservation biology — at least the early literature defined as the conservation biology literature of the early 1980’s — was dominated by studies of the threats to small populations, and genetic threats were among those most prominently identified. What exactly are those threats? Well, they can be divided into two basic categories: 1 Don’t forget that stochastic threats are primarily important only after a population is already small and endangered. Deterministic, systematic forces associated with habitat destruction, environmental degradation, invasive exotics, etc. are the most likely causes of population declines. c 2001–2015 Kent E. Holsinger 1. Short-term effects on individual viability and fecundity — These are the result of changes in the genetic structure of populations that potentially have a direct effect on demographics of the population. They could make it even more susceptible to extinction than in would be if these factors could be ignored 2. Long-term effects on the ability of populations to respond adaptively to environmental change — Even if genetic stochasticity has no immediate effect on the ability of populations to persist, it might have some effect on their ability to respond when environmental change occurs, whether we are talking about global climate change or the introduction of a new pathogen. After all, populations can respond adaptively to environmental change only if they have the necessary genetic variability, and small populations tned to harbor less variability than large ones. Before we talk about the possible effects of genetic changes on short-term and long-term survival, it is necessary first to talk about the types of genetic changes that happen in small populations. Only then can we explore their effects.2 Genetic changes in small populations As suggested by the fact that we refer to these changes as threats due to genetic stochasticity, the genetic changes specific to small populations are random. To talk about this in more detail, we need to develop a little background in population genetics, specifically the theory of genetic drift. It’s been known for more than 80 years that the randomness associated with reproduction leads to changes in the genetic composition of a population through time. Consider this: if gametes are chosen at random to be included in zygotes, the genetic composition of the zygote pool will, on average, be the same as the gamete pool, but there is a distribution of possible outcomes. We won’t go into the mathematical details here, but there are four basic properties of drift important for our purposes:3 1. Allele frequencies tend to change from one generation to the next simply as a result of sampling error. We can specify a probability distribution for the allele frequency in the next generation, but can’t say with certainty what the exact value will be. 2 This will be a review for those of you who’ve had my course in population genetics. In fact, I should probably make you give this part of the lecture, but I won’t. 3 If you want a more complete discussion, refer to chapter 7 of [8] or take my course in population genetics next fall. 2 Figure 1: Frequency distribution of predicted heterozygosity loss for 80 mammal species (from [1], Figure 11.4 [7]) 2. There is no systematic bias associated with the change in allele frequency, i.e., we can’t predict which alleles will become more common and which will become rarer. 3. Populations eventually become fixed for one of the alleles originally present in the population, unless mutation or migration introduces new genetic variation, i.e., they tend to lose genetic diversity (Figure 1). 4. Time to fixation (and other properties of drift) are inversely proportional to population size — the larger the population, the smaller the effect of drift (Figure 2). The time to common ancestry of two randomly chosen alleles in a population is 2Ne . The time to common ancestry of all alleles in a population is 4Ne [10]. This last point is perhaps the most important, and the most problematic, for it turns out that simply counting the number of reproductive plants or animals in a population is not sufficient to determine whether the population is large or small. It is not the census number4 of individuals in the population that matters, but the effective number.5 The formulas that follow are from [2, p. 362]. 4 census number — the number of individuals in a population, simply counting all that are alive effective population size — the size of a population for purposes of genetic drift, adjusted for unequal sex-ratio, age structure, variable population size, and the like. 5 3 Figure 2: Fraction of genetic diversity lost in each generation for populations of different effective size (Figure 11.3 [7]). Unequal sex ratio Ne = 4Nf Nm Nf + Nm Note: If there is only one reproductive male, the greatest the effective size can ever be is 4. Management implication — Unequal sex ratios can dramatically increase the susceptibility of a population to random genetic changes. Example: one reproductive male mates with 10 reproductive females: Ne = 4(10)(1) 40 = ≈ 3.6 (10 + 1) 11 Another example: ten reproductive males and 100 reproductive females: Ne = 4(100)(10) 4000 = ≈ 36.3 (100 + 10) 110 Variation in reproductive success Ne = 2N 2 1 + σk̄ 4 , where σ 2 is the variance in family size among individuals and k̄ is the average number of offspring per individual. Note: If σ 2 = 0, Ne = 2N . If σ 2 = 3k̄, Ne = N/2. Suppose, for example that the distribution of offspring number is negative binomial with mean 2 and variance 12.6 Then 2N Ne = 1+(12/2) ≈ 0.28N . Management implication — Reducing the differences among individuals in the number of offspring they produce can almost halve the susceptibility of a population to random genetic changes. There is a complication worth noting here, however. What I just said is true with respect to loss of genetic diversity as a result of genetic drift, but it’s true only if the population size is constant with respect to the increase in relatedness among individuals in the population.7 If the population size is increasing the effective size is close to size of progeny population (not parental population) and equal reproduction increases the effective population size by less than a factor of two. In fact, if the population size is increasing rapidly enough, equalizing the reproductive success of parents would cause relatedness to accumulate more rapidly if it limits the rate of population growth. Variation in population size 1 Ne = " 1− ≈ Pt Qt k=1 t 1− 1 (k) 2Ne 1/t # (1) (2) 1 k=1 N (k) t (3) where Ne(k) is the effective size of the population in generation k. Note: If N = 1000 for k = 1, 2, · · · 9, but N = 10 for k = 10, Ne = 83 versus a mean of 901. Effective size is much more drastically affected by smallest population size than largest population size. Management implication — Populations subject to occasional crashes are more susceptible to random genetic changes than those with relatively stable populations. 6 If you’ve never heard of a negative binomial distribution before, don’t worry about it. All you need to know is that the variance of a negative binomial distribution is larger than that of a Poisson distribution with the same mean. 7 The reasons for this are technical and have to do with the difference between the variance and inbreeding effective size of a population. Ask me if you’re interested in the details. 5 Age-structured populations The stochastic dynamics of age-structured populations are complicated. To my mind we still don’t understand them very well. Age structure does tend to reduce effective population size still further (unless we’re dealing with annual plants that have a seed bank). Management implication — We have to trust that inferences about management practices without considering age structure aren’t qualitatively changed by age structure. Just be conservative, i.e., build in a large margin for error. Genetic changes during population bottlenecks There are two components to what is often loosely referred to as “genetic diversity:” additive genetic variance (the portion of the genetic differences among individuals that can respond to selection) and allelic diversity (the number of different types of alleles present at any locus). Even in the most extreme situation imaginable, a population reduced to one hermaphroditic individual for one generation, at most 50% of the additive genetic variance in quantiative traits will be lost if the population rebounds quickly to a large population size (Figure 3). A less extreme case, 2 males and 2 females would lead to only a 12% reduction in additive genetic variance. Moreover, selection against recessive lethal and deleterious alleles will retard the loss of variance at other loci, especially those to which they are linked. In short, several generations of greatly reduced population size are necessary to significantly deplete the aspect of genetic diversity that is responsible for adaptive responses to natural selection. The variety of alleles present, however, is greatly affected by population bottlenecks. Rare alleles are particularly susceptible to loss. In fact, population geneticists use the difference between the effect of bottlenecks on diversity and variety to detect the effect of selection or population size changes on nucleotide diversity. But for our purposes, the important observation is this: Population bottlenecks have little effect on the genetic variance responsible for adaptive responses to natural selection, even though they may have a dramatic effect on the number of alleles present. Short-term threats to persistence Drift in small populations has many of the same properties as inbreeding. In fact, 6 Figure 3: Loss and recovery of genetic diversity after a population bottleneck (from [18]; Figure 11.5 [7]). 1 (1 − ft ) = ft + 1 − 2Ne 1 = 2Ne 1 t = 1− 1− 2Ne ft+1 1 − ft+1 1 − ft ft (4) (5) (6) (7) where f is the inbreeding coefficient, a measure of the degree of inbreeding. By trial and error animal breeders have discovered how much inbreeding can be tolerated by domestic animals before the lines begin to decline in performance and fecundity [5, 19]. Their rule t+1 of thumb is that the per generation rate of inbreeding 1−f = 2N1 e should not be greater 1−ft than 2%–3%. If we want to be a little conservative, we might reduce this to 1%. Thus, we require8 1 < 0.01 2Ne 8 This calculation is the entire justification for the “50” part of the famous (infamous) 50/500 rule. 7 Ne > 50 An alternative approach is to observe that animal breeders have found a obvious effect on fecundity in small populations when the inbreeding coefficient approaches 0.5. If we want our population to be viable for 100 generations 0.5 > 1 − 1 − 1− 1 2Ne 1 2Ne 100 (8) 100 > 0.5 1 > (0.5)0.01 = 0.993 2Ne 1 0.007 > 2Ne Ne ≥ 73 1− (9) (10) (11) (12) (13) Still another approach is to assume that the deleterious effects noted in these small populations is primarily the result of the expression of recessive deleterious alleles. Then using the results of drift theory we can calculate the probability that a population has a particular allele frequency, given assumptions about the strength of selection and mutation rates. For a broad range of strentgths of selection and for what are thought to be typical per locus mutation rates (10−6 per generation) it is possible to calculate the effect on population mean fitness. This calculation suggests that populations will suffer noticeably (mean fitness reduction greater than 10%) if Ne < 100–300 [9]. Finally, we can consider a population-level version of Muller’s ratchet. As a result of genetic drift, there’s always a chance that a deleterious allele can be fixed as a result of genetic drift. If it does and if it reduces the reproductive capacity of the population, the population size may get smaller, making it easier for new deleterious mutations to fix, which will reduce the population size further, and so on, and so on, and so on. Mike Lynch has called this process the “mutational meltdown” [6, 14]. The expected time to extinction increases rapidly in mutational meltdown models such that in populations with an effective size greater than a few hundred, persistence times are well into the hundreds or thousands of generations (Figure 4). In addition to the effects of inbreeding depression, which have been found in virtually every outbreeding organism — plant or animal — that has ever been studied9 , there has been 9 Except those that are known to have survived recent, severe population bottlenecks. 8 Figure 4: Mean time to extinction for monoecious populations with a constant carrying capacity (open circles and dashed line) and monogamous populations with variable carrying capacities (lognormal with the indicated coefficients of variation). Mutational parameters are close to those estimated for Drosophila melanogaster: genomic mutation rate = 1.5, selection coefficient against recessive homozygotes = 0.015, dominance coefficient = 0.35. Ten offspring are produced per individual (from [14].) 9 some suggestion that loss of genetic diversity may have an immediate impact on short-term survival. My (admittedly biased) reading of the evidence is that the evidence on this point is at best equivocal. The bottom line? Recall that to buffer the effects of environmental stochasticity we require populations consisting of thousands or tens of thousands of reproductive individuals. Ne is unlikely to be less than 0.1, except in rare circumstances, so populations large enough N to buffer environmental stochasticity are almost certainly large enough to buffer genetic stochasticity. On the other hand, when populations are critically endangered, genetic changes may pose an additional threat to population persistence. An example: Populations of the Flordia panther are extremely small. The high proportion of kinked tails and cowlicks among individuals in the population (about 90% in both cases) coupled with the poorest semen quality ever recorded suggests that the population is suffering from substantial inbreeding depression. As a result,10 Texas cougars were released into Florida in 1995 in the hope that individual fitness would rapidly improve. No kinked tails have been recorded from F1 and F2 offspring, and the proportion of F1 and F2 individuals with the cowlick are far less frequent (about 14%), suggesting that efforts to reduce the effects of inbreeding depression may have been successful [11]. Another example: Mountain gorillas survive as only two critically endangered populations, and the total population in the wild may consist of no more than 800 individuals. A recent whole-genome analysis of 13 gorillas, including 7 mountain gorillas, found that chromosomes were typically homozygous over more than 1/3 of their length [21]. Some tracts of homozygosity were as much as 10Mb long, suggesting recent inbreeding within the small remaining populations. Nonetheless, the authors conclude that These subspecies have survived for thousands of generations at low population levels and may have developed physiological and behavioral strategies to mitigate inbreeding, such as natal dispersal and gene flow between isolated populations. The purging of strongly deleterious mutations may also be an important factor. An exception: loss of self-incompatibility alleles in plants with genetically determined self-incompatibility. Hymenoxys acaulis: Illinois populations have a single compatibility type, Ohio populations have only 3-9. Long-term persistence of Illinois populations requires import of genotypes from Ohio. Reproductive capacity of all populations limited by availability of compatible mates. But “[T]he observed low genetic diversity in today’s population [of Tasmanian devil] preceded the Devil Facial Tumor Disease Outbreak by at least 100 y[ears]” [15], although a more recent analysis [17] argues that the loss of immunodiversity may leave the Tasmanian 10 And after considerable debate, as you can imagine. 10 devil “highly vulnerable to infectious disease.” If that’s the case, then loss of immunodiversity may be comparable to loss of self-incompatibility alleles. Long-term threats to persistence The threats to long-term persistence are those associated with loss of the genetic variability necessary for adaptation to evnironmental change. One way of assessing this threat is to ask how large the population must be to balance the loss of additive genetic variance through drift with the input of additive genetic variance through mutation. Several reviews suggest that the mutation rate for quantitative characters is on the order of 10−3 [12, 13]. The rate of loss, recall, is 2N1 e . This would suggest that an Ne of 500 is necessary to ensure longterm evolutionary potential [5].11 Again, it would appear that populations large enough to buffer environmental stochasticity will also be large enough to maintain the additive genetic variance necessary to respond adaptively to environmental change. When we recall that the main effect of population bottlenecks is to reduce the number of alleles present, there is even more reason to suspect that this is a reasonable conclusion. • Most of the genetic variance responsible for an evolutionary response to natural selection (the additive genetic variation) is found in high-frequency alleles, precisely those least likely to be lost in small populations. • Many low-frequency alleles are probably unconditionally deleterious and are maintained in the population only by recurrent mutation. Their loss may actually benefit the population. • Low-frequency alleles have a short lifetime in populations. They are quickly lost through drift. Current low-frequency alleles are unlikely to provide the source of evolutionary novelties. They are more likely to arise as a result of mutation in the future. • Alleles that are selectively advantageous are unlikely to be lost from populations in a short time as a result of drift unless the populations are very small. But it is important to remember that when we manage species, they may respond not only through changes that are primarily ecological, they may adapt to the new management regime to which they are subject. This is most obvious when the managed environment is very different, as when captive populations are bred in zoos, aquaria, or botanical gardens, 11 The “500” part of the 50/500 rule. 11 but it may also happen in the wild if populations are restricted to a part of their former range.As Stockwell and his colleagues point out, refuge populations may diverge from those in other parts of their range making it more difficult to use them as source populations for re-introduction or restoration efforts. Surviving with low genetic diversity I argued from first principles that loss of genetic diversity is more likely to be a symptom of endangerment than a cause of endangerment. Milot et al. [16] present an example that is consistent with my argument. The authors compare levels of genetic diversity (as judged with AFLPs12 ) in the wandering albatross and the Amsterdam albatross. Based on molecular clock estimates, these two taxa diverged from one another about 800,000 years ago. The Amsterdam albatross is recovering from an extreme bottleneck. When first discovered in the early 1980s, only 5 breeding pairs were known. Approximately 130 adult birds now exist (http://en.wikipedia.org/wiki/Amsterdam Albatross). Only 5% of AFLP loci are polymorphic in wandering albatross, and only 2% are polymorphic in Amsterdam albatross. In contrast, other vertebrates show polymorphism at 17-99% of AFLP loci. Simulations suggest that the low levels of polymorphism are not the result of population bottlenecks in the species’ evolutionary history.13 Instead, it appears that they both inherited a low level of polymorphism from their ancestor. If that’s right, then both species have survived for 800,000 years with very low levels of genetic diversity, suggesting that lack of diversity is a reflection of their life history and that it did not contribute to very small population sizes in the Amsterdam albatross. A caveat There is one detail about AFLP markers that’s important to know. Differences among individuals at most AFLP loci probably have little or no impact on survival and reproduction. Lack of diversity at these loci provides information on levels of adaptively significant variation only through several steps of logic (low neutral diversity implies small effective population size means natural selection is less effective means loss of diversity even if it’s favored by selection). It’s possible that levels of adaptive diversity in wandering and Amsterdam albatrosses are 12 The molecular basis of this variation isn’t important for our purposes. If you’re interested, ask me. Remember, only the Amsterdam albatross shows evidence of recent recovery from a bottleneck. The wandering albatross has substantially larger populations. 13 12 markedly higher than at AFLPs. Nonetheless, it seems likely that they are still lower than levels of adaptive diversity in other vertebrates. Avoiding inbreeding depression I mentioned that animals from Texas were introduced into southern Florida in an attempt to alleviate problems associated with inbreeding depression in the Florida panther, and I mentioned that evidence suggests that the attempt was successful. I didn’t mention that beyond the obvious concern that many conservationists have about manipulating populations in this way, it isn’t always successful. You may remember from your study of evolutionary biology that barriers to reproduction between species may arise as a result of hybrid breakdown. Hybrid breakdown occurs when there are epistatic interactions among loci causing only certain combinations of alleles at different loci to work well together. A phenomenon like hybrid breakdown can occur when crosses are made between populations of the same species that have become differentially adapted — outbreeding depression. So there’s the risk that in attempting to alleviate problems associated with inbreeding depression, conservation managers could introduce new problems associated with outbreeding depression. The resulting advice is pretty simple in principle, if difficult to put into practice: “[M]anagers can minimize the risks of both inbreeding and outbreeding by using intentional hybridization only for populations clearly suffering from inbreeding depression, maximizing the genetic and adaptive similarity between populations, and testing the effects of hybridization for at least two generations whenever possible” [3, p. 463].14 References [1] W Amos and A Balmford. When does conservation matter. Heredity, 87:257–265, 2001. [2] J F Crow and M Kimura. An Introduction to Population Genetics Theory. Burgess Publishing Company, Minneapolis, Minn., 1970. [3] Suzanne Edmands. Between a rock and a hard place: evaluating the relative risks of inbreeding and outbreeding for conservation and management. Molecular Ecology, 16(3):463–475, 2007. 14 For a more recent analysis, see [4] for a more recent meta-analysis arguing that we may have been too cautious in avoiding attempts at genetic rescue, and an accompanying commentary [20] that remains skeptical. 13 [4] Richard Frankham. Genetic rescue of small inbred populations: meta-analysis reveals large and consistent benefits of gene flow. Molecular Ecology, 24(11):2610–2618, 2015. [5] I R Franklin. Evolutionary change in small populations. In M E Soulé and B A Wilcox, editors, Conservation Biology: An Evolutionary-Ecological Perspective, pages 135–150. Sinauer Assoc., Sunderland, MA, 1980. [6] W Gabriel, M Lynch, and R Brger. Muller’s ratchet and mutational meltdowns. Evolution, 47(6):1744–1757, 1993. [7] M Groom, G K Meffe, and C R Carroll. Principles of Conservation Biology. Sinauer Associates, Sunderland, MA, 3rd edition, 2005. [8] D L Hartl and A G Clark. Principles of Population Genetics. Sinauer Associates, Sunderland, MA, 3rd editio edition, 1997. [9] Kent E Holsinger and L D Gottlieb. Conservation of rare and endangered plants: principle and prospects. In Donald A Falk and Kent E Holsinger, editors, Genetics and Conservation of Rare Plants, pages 195–208. Oxford University Press, New York, NY, 1991. [10] R R Hudson. Gene genealogies and the coalescent process. In D J Futuyma and J Antonovics, editors, Oxford Surveys in Evolutionary Biology, volume 7, pages 1–44. Oxford Univ. Press, Oxford, 1990. [11] Warren E Johnson, David P Onorato, Melody E Roelke, E Darrell Land, Mark Cunningham, Robert C Belden, Roy McBride, Deborah Jansen, Mark Lotz, David Shindle, JoGayle Howard, David E Wildt, Linda M Penfold, Jeffrey A Hostetler, Madan K Oli, and Stephen J OÄôBrien. Genetic Restoration of the Florida Panther. Science, 329(5999):1641–1645, 2010. [12] R Lande. The maintenance of genetic variability by mutation in a polygenic character with linked loci. Genetical Research, 26:221–235, 1975. [13] M Lynch. The rate of polygenic mutation. Genet. Res. (Camb.), 51:137–148, 1988. [14] M Lynch, J Conery, and R Bürger. Mutation accumulation and the extinction of small populations. American Naturalist, 146(4):489–518, 1995. [15] Webb Miller, Vanessa M Hayes, Aakrosh Ratan, Desiree C Petersen, Nicola E Wittekindt, Jason Miller, Brian Walenz, James Knight, Ji Qi, Fangqing Zhao, Qingyu 14 Wang, Oscar C Bedoya-Reina, Neerja Katiyar, Lynn P Tomsho, Lindsay McClellan Kasson, Rae-Anne Hardie, Paula Woodbridge, Elizabeth A Tindall, Mads Frost Bertelsen, Dale Dixon, Stephen Pyecroft, Kristofer M Helgen, Arthur M Lesk, Thomas H Pringle, Nick Patterson, Yu Zhang, Alexandre Kreiss, Gregory M Woods, Menna E Jones, and Stephan C Schuster. Genetic diversity and population structure of the endangered marsupial Sarcophilus harrisii (Tasmanian devil). Proceedings of the National Academy of Sciences, 108(30):12348–12353, 2011. [16] E Milot, H Weimerskirch, P Duchesne, and L Bernatchez. Surviving with low genetic diversity: the case of albatrosses. Proceedings of the Royal Society B: Biological Sciences, 274(1611):779–787, 2007. [17] Katrina M Morris, Belinda Wright, Catherine E Grueber, Carolyn Hogg, and Katherine Belov. Lack of genetic diversity across diverse immune genes in an endangered mammal, the Tasmanian devil (Sarcophilus harrisii). Molecular Ecology, 24(15):3860–3872, 2015. [18] M Nei. Molecular Population Genetics and Evolution. North Holland Publishing Company, Amsterdam, 1975. [19] Michael E Soulé. Thresholds for survival: maintaining fitness and evolutionary potential. In Michael E Soulé and Bruce A Wilcox, editors, Conservation Biology: An Evolutionary-Ecological Perspective, pages 151–169. Sinauer Associates, Sunderland, MA, 1980. [20] Donald M Waller. Genetic rescue: a safe or risky bet? Molecular Ecology, 24(11):2595– 2597, 2015. [21] Yali Xue, Javier Prado-Martinez, Peter H Sudmant, Vagheesh Narasimhan, Qasim Ayub, Michal Szpak, Peter Frandsen, Yuan Chen, Bryndis Yngvadottir, David N Cooper, Marc de Manuel, Jessica Hernandez-Rodriguez, Irene Lobon, Hans R Siegismund, Luca Pagani, Michael A Quail, Christina Hvilsom, Antoine Mudakikwa, Evan E Eichler, Michael R Cranfield, Tomas Marques-Bonet, Chris Tyler-Smith, and Aylwyn Scally. Mountain gorilla genomes reveal the impact of long-term population decline and inbreeding. Science, 348(6231):242–245, 2015. Creative Commons License These notes are licensed under the Creative Commons Attribution-NonCommercial-ShareAlike License. To view a copy of this license, visit 15 http://creativecommons.org/licenses/by-nc-sa/3.0/ or send a letter to Creative Commons, 559 Nathan Abbott Way, Stanford, California 94305, USA. 16