MEC_3905_sm_Supporting
advertisement

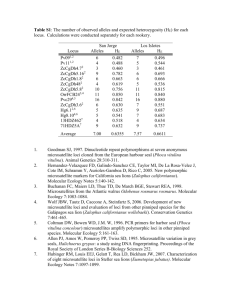
ONLINE SUPPORTING MATERIAL TITLE: RANGE-WIDE GENETIC HOMOGENEITY IN THE CALIFORNIA SEA MUSSEL (MYTILUS CALIFORNIANUS): A COMPARISON OF ALLOZYME, NUCLEAR DNA MARKERS, AND MITOCHONDRIAL DNA SEQUENCES. AUTHORS: Jason A. Addison, Brian S. Ort, Kathryn A. Mesa, and Grant H. Pogson Supporting Material 1. Summary of studies comparing the magnitude of population structuring between allozymes and nuclear DNA markers. The mean divergence column indicates the concordance between mean FST or RST using the method of Pogson et al. (1995). Species names are listed alphabetically. Common name Species name Mean Divergence Reference River mangrove Aegiceras corniculatum Mottled duck Anas fulvigula Herbaceous perennial Antirrhinum microphyllum allozymes = RAPDs Torres et al. 2003 Am. J. Bot. 90:85-92 Alder-leaf birch Betula alnoides allozymes = RAPDs Zeng et al. 2003. Ephytica 134:33-41 Cranberry fritillary Boloria aquilonaris allozymes = RAPDs Vandewoestijne and Baguette. 2002. Heredity 89:439-445 allozymes = ISSRs1 Ge and Sun. 1999. Mol. Ecol. 8:2061-2069 allozymes = microsatellites Williams et al. 2005. Condor 107:155-161 Asian water buffalo Bubalus bubalis Buffalo grass Buchloe dactyloides Common toad Bufo bufo Hairy edible crab Cancer setosus Perennial herb Centaurea corymbosa Flowering quince Chaenomeles cathayensis allozymes = RAPDs Bartish et al. 2000. Theor. Appl. Genet. 101:554-563 Japanese quince Chaenomeles japonica allozymes = RAPDs Bartish et al. 2000. Theor. Appl. Genet. 101:554-563 Chinese flowering quince Chaenomeles speciosa allozymes = RAPDs Bartish et al. 2000. Theor. Appl. Genet. 101:554-563 Indian major carp Cirrhinus mrigala allozymes = microsatellites Chauhan et al. 2007. Aquaculture 269 : 135-149 Atlantic herring Clupea harengus allozymes = microsatellites Larsson et al. 2007. Mol. Ecol. 16:1135-1147 Zebrafish Danio rerio allozymes = microsatellites Gratton et al. 2004. J. Zool. Syst. Evol. Research 42:54-62 European sea bass Dicentrarchus labrax allozymes > microsatellites Lemaire et al. 2000. Mol. Ecol.9:457-467 Fibrous wheatgrass Elymus fibrosus Dusky grouper Epinephelus marginatus allozymes > microsatellites De Innocentiis et al. 2001. Mol. Ecol. 10:2163-2175 Tsetse fly Glossina pallidipes allozymes = microsatellites Krafsur. 2002. Insect Mol. Biol. 11:37-45 S. African abalone Haliotis midae Wild barley Hordeum spontaneum allozymes = RAPDs Volis et al. 2005. Heredity 95:466-475 Alfalfa Medicago sativa allozymes = RAPDs Jenczewski et al. 1999. Mol. Ecol. 8:1317-1330 Red mullet Mullus surmuletus allozymes = RAPDs Mamuris et al. 1999. Heredity 83:30-38 Wood stork Mycteria americana microsatellites > allozymes Rocha et al. 2004. Biotropica 36:248-258 Chum salmon Oncorhynchus keta allozymes = microsatellites Scribner et al. 1998. Can. J. Fish. Aquat. Sci. 55:1748-1758 allozymes = microsatellites allozymes = RAPDs allozymes > minisatellites allozymes > AFLPs allozymes = microsatellites allozymes = RAPDs allozymes = microsatellites Barker et al. 1997. Animal Genetics 28:103-115 Peakall et al. 1995. Mol. Ecol. 4:135-147 Scribner et al. 1994. Mol. Biol. Evol. 11:737-748 Gomez-Uchida et al. 2003. J. Crust Biol. 23:486-494 Freville et al. 2001. Mol. Ecol. 10:879-889 Diaz et al. 2000. Genet. Resour. Crop Evol. 47:11-24 Evans et al. 2004. Mar. Ecol. Prog. Ser. 270:163-172 Sockeye salmon Oncorhynchus nerka allozymes > microsatellites Allendorf and Seeb. 2000. Evolution 54:640-651 Chinook salmon Oncorhynchus tshawytscha allozymes = microsatellites Smith et al., 2007. Trans. Am. Fish. Soc. 136:1674-1687 Chinook salmon Oncorhynchus tshawytscha SNPs > allozymes Smith et al., 2007. Trans. Am. Fish. Soc. 136:1674-1687 Brownbeard rice Oryza rufipogon allozymes = RAPDs Brownbeard rice Oryza rufipogon allozymes = microsatellites Gao et al. 2002. Theor. Appl. Genet. 106:173-180 Geoduck clam Panopea abrupta allozymes > microsatellites Vadopalas et al. 2004. J. Shellfish Res. 23:693-706 Clouded Apollo butterfly Parnassius mnemosyne allozymes = microsatellites Meglecz et al. 1998. Hereditas 128:95-103 White spruce Picea glauca allozymes = ESTPs2 Jaramillo-Correa et al. 2001. Mol. Ecol. 10:2729-2740 Black spruce Picea mariana RAPDs3 > allozymes Isabel et al. 1995. Proc. Nat. Acad. Sci. USA 92:6369-6373 Knobcone pine Pinus attenuata allozymes = RAPDs Wu et al. 1999. Genome 42:893-908 Bristlecone pine Pinus longaeva RAPDs4 > allozymes Lee et al. 2002. Am. J. Botany 89:566-577 Bishop pine Pinus muricata allozymes = RAPDs Wu et al. 1999. Genome 42:893-908 Monterey pine Pinus radiata allozymes = RAPDs Wu et al. 1999. Genome 42:893-908 Scots pine Pinus sylvestris allozymes = RAPDs Szmidt et al. 1996. Heredity 76:412-420 Saltmarsh beetle Pogonus chalceus Moss Pleurochate squarrosa (Brid.) Sand goby Pomatoschistus minutus Mesopsammic flatworm Pseudomonocelis ophiocephala allozymes = RAPDs Casu and Curini-Galletti. 2006. Biol. J. Linn. Soc. 87:553-576 Douglas-fir Pseudotsuga menziesii allozymes = RAPDs Aagaard et al. 1998. Heredity 81:69-78 Little red flying-fox Pteropus scapulatus allozymes = RAPDs Sinclair et al. 1996. Biol. Conserv. 76:45-50 allozymes > microsatellites allozymes = nuclear ITS allozymes = microsatellites Cai et al. 2004. Heredity 92:409-417 Dhuyvetter et al. 2004. Mol. Ecol. 13:1065-1074 Grundmann et al. 2007. Mol. Ecol. 16:709-722 Pampoulie et al. 2004. Heredity 92:434-445 1 Sessile oak Quercus petraea Bird cherry-oat aphid Rhopalosiphum padi allozymes = microsatellites Delmotte et al. 2002. Mol. Ecol. 11:711-723 Brown trout Salmo trutta allozymes = microsatellites Corujo et al. 2004. Hereditas 141:258-271 Brown trout Salmo trutta allozymes = microsatellites Colihueque et al. 2003. Aquaculture Res. 34:525-533 Brown trout Salmo trutta allozymes = RAPDs Brown trout Salmo trutta allozymes = microsatellites Estoup et al. 1998. Mol. Ecol. 7:339-353 Turbot Scophthalmus maximus allozymes = microsatellites Buoza et al. 2002. Can. J. Fish. Aquat. Sci. 59:1460-1473 Acorn barnacle Semibalanus balanoides allozymes > microsatellites Dufresne et al. 2002 Mol. Ecol. 11:113-123 Dover sole Solea solea RAPDs > allozymes Exadactylos et al. 1998. J. Sea Res. 40 :117-129 Exadactylos et al. 2003. Mar. Ecol. Prog. Ser. 246:253-264 Stoplight parrotfish Sparisoma viride RAPDs > allozymes Geertjes et al. 2004. Mar. Ecol. Prog. Ser. 279:225-235 Yellowfin tuna Thunnus albacares allozymes = RAPDs Diaz-Jaimes and Uribe-Alcocer. 2003. Fish. Bull. 101:769-777 Bolander mule ears Wyethia bolanderi allozymes = RAPDs Ayres and Ryan. 1999. Am. J. Bot. 86:344-353 El Dorado Co. mule ears Wyethia reticulata allozymes = RAPDs Ayres and Ryan. 1999. Am. J. Bot. 86:344-353 Maize Zea mays allozymes = RAPDs Dubreuil and Charcosset. 1998. Theor. Appl. Genet. 96:577-587 allozymes = RAPDs Le Corre et al. 1997. Mol. Ecol. 6:519-529 Cagigas et al. 1999. Mar. Biotechnol. 1:286-296 ISSRs = intersimple sequence repeat polymorphisms; 2 ESTPs = expressed sequence tag polymorphisms; 3,4 mean FST-value for RAPDs not significantly different from zero. LITERATURE CITED (SUPPORTING MATERIAL) 1 2 3 Aagaard JE, Krutovskii KV, Strauss SH (1998) RAPDs and allozymes exhibit similar 4 levels of diversity and differentiation among populations and races of Douglas-fir. 5 Heredity, 81, 69-78. 6 Allendorf FW, Seeb LB (2000) Concordance of genetic divergence among sokeye 7 salmon populations at allozyme, nuclear DNA, and mitochondrial DNA markers. 8 Evolution, 54, 640-651. 9 Ayres DR, Ryan FJ (1999) Genetic diversity and structure of the narrow endemic 10 Wyethia reticulata and its congener W. bolanderi (Asteraceae) using RAPD and 11 allozyme techniques. American Journal of Botany, 86, 344-353. 12 Barker JSF, Moore SS, Hetzel DJS, Evans D, Tan SG, Byrne K (1997) Genetic diversity 13 of Asian water buffalo (Bubalus bubalis): microsatellite variation and a comparison 14 with protein-coding loci. Animal Genetics, 28, 103-115. 15 Bartish IV, Garkava LP, Rumpunen K, Nybom H (2000) Phylogenetic relationships and 16 differentiation among and within populations of Chaenomeles Lindl. (Rosaceae) 17 estimated with RAPDs and isozymes. Theoretical and Applied Genetics, 101, 554- 18 563. 19 Bouza C, Presa P, Castro J, Sánchez L, Martínez P (2002) Allozyme and microsatellite 20 diversity in natural and domestic populations of turbot (Scophthalmus maximus) in 21 comparison with other Pleuronectiformes. Canadian Journal of Fisheries and 22 Aquatic Sciences, 59, 1460-1473. 23 Cagigas ME, Vázquez E, Blanco G, Sánchez JA (1999) Combined assessment of genetic 24 variability in populations of brown trout (Salmo trutta L.) based on allozymes, 25 microsatellites, and RAPD markers. Marine Biotechnology, 1, 286-296. 26 Cai H-W, Wang X-K, Morishima H (2004) Comparison of population genetic structures 27 of common wild rice (Oryza rufipogon Griff.), as revealed by analyses of 28 quantitative traits, allozymes, and RFLPs. Heredity, 92, 409-417. 29 Casu M, Curini-Galletti M (2006) Genetic evidence for the existence of cryptic species 30 in the mesopsammic flatworm Pseudomonocelis ophiocephala (Rhabditophora: 31 Proseriata). Biological Journal of the Linnean Society, 87, 553-576. 32 Chauhan T, Lal KK, Mohindra V, Singh RK, Punia P, Gopalakrishnan A, Sharma PC, 33 Lakra WS (2007) Evaluating genetic differentiation in wild populations of the 34 Indian major carp, Cirrhinus mrigala (Hamilton – Buchanan, 1882): evidence from 35 allozyme and microsatellite markers. Aquaculture, 269, 135-149. 36 Colihueque N, Vergara N, Parraguez M (2003) Genetic characterization of naturalized 37 populations of brown trout Salmo trutta L. in southern Chile using allozyme and 38 microsatellite markers. Aquaculture Research, 34, 525-533. 39 Corujo M, Blanco G, Vázquez E, Sánchez JA (2004) Genetic structure of northwestern 40 Spanish brown trout (Salmo trutta L.) populations, differences between 41 microsatellite and allozyme loci. Hereditas, 141, 258-271. 42 De Innocentiis S, Sola L, Cataudella S, Bentzen P (2001) Allozyme and microsatellite 43 loci provide discordant estimates of population differentiation in the endangered 44 dusky grouper (Epinephelus marginatus) within the Mediterranean Sea. Molecular 45 Ecology, 10, 2163-2175. 46 Delmotte F, Leterme N, Gauthier J-P, Rispe C, Simon J-C (2002) Genetic architecture of 47 sexual and asexual populations of the aphid Rhopalosiphum padi based on allozyme 48 and microsatellite markers. Molecular Ecology, 11, 711-723. 49 Dhuyvetter H, Gaublomme G, Desender K (2004) Genetic differentiation and local 50 adaptation in the salt-marsh beetle Pogonus chalceus: a comparison between 51 allozyme and microsatellite loci. Molecular Ecology, 13, 1065-1074. 52 Díaz O, Sun G-L, Salomon B, von Bothmer R (2000) Levels and distribution of 53 allozyme and RAPD variation in populations of Elymus fibrosus (Schrenk) Tzvel. 54 (Poaceae). Genetic Resources and Crop Evolution, 47, 11-24. 55 Diaz-Jaimes P, Uribe-Alcocer M (2003) Allozyme and RAPD variation in the eastern 56 Pacific yellowfin tuna (Thunnus albacares). Fishery Bulletin, 101, 769-777. 57 Dubreuil P, Charcosset A (1998) Genetic diversity within and among maize populations: 58 a comparison between isozyme and nuclear RFLP loci. Theoretical and Applied 59 Genetics, 96, 577-587. 60 Dufresne F, Bourget E, Bernatchez L (2002) Differential patterns of spatial divergence 61 in microsatellite and allozyme alleles: further evidence for locus-specific selection 62 in the acorn barnacle, Semibalanus balanoides? Molecular Ecology, 11, 113-123. 63 Estoup A, Rousset F, Michalakis Y, Cornuet J-M, Adriamanga M, Guyomard R (1998) 64 Comparative analysis of microsatellite and allozyme markers: a case study 65 investigating microgeographic differentiation in brown trout (Salmo trutta). 66 Molecular Ecology, 7, 339-353. 67 Evans B S, Sweijd NA, Bowie RCK, Cook PA, Elliott NG (2004) Population genetic 68 structure of the perlemoen Haliotis midae in South Africa: evidence of range 69 expansion and founder events. Marine Ecology Progress Series, 270, 163-172. 70 Exadactylos A, Geffen AJ, Thorpe JP (1998) Population structure of the Dover sole, 71 Solea solea L., in a background of high gene flow. Journal of Sea Research, 40, 72 117-129. 73 Exadactylos A, Geffen AJ, Panagiotaki P, Thorpe JP (2003) Population structure of 74 Dover sole Solea solea: RAPD and allozyme data indicate divergence in European 75 stocks. Marine Ecology Progress Series, 246, 253-264. 76 Freville H, Justy F, Olivieri I (2001) Comparative allozyme and microsatellite 77 population structure in a narrow endemic plant species, Centaurea corymbosa 78 Pourret (Asteraceae). Molecular Ecology, 10, 879-889. 79 Gao L-Z, Schaal BA, Zhang C-H, Jia J-Z, Dong Y-S (2002) Assessment of population 80 genetic structure in common wild rice Oryza rufipogon Griff. using microssatellite 81 and allozyme markers. Theoretical and Applied Genetics, 106, 173-180. 82 Ge XJ, Sun M (1999) Reproductive biology and genetic diversity of a cryptoviviparous 83 mangrove Aegiceras corniculatum (Myrsinaceae) using allozyme and intersimple 84 sequence repeat (ISSR) analysis. Molecular Ecology, 8, 2061-2069. 85 Geertjes GJ, Postema J, Kamping A, van Delden W, Videler JJ, van de Zande L (2004) 86 Allozymes and RAPDs detect little genetic population substructuring in the 87 Carribean stoplight parrotfish Sparisoma viride. Marine Ecology Progress Series, 88 279, 225-235. 89 Gomez-Uchida D, Weetman D, Hauser L, Galleguillos R, Retamal M (2003) Allozyme 90 and AFLP analyses of genetic population structure in the hairy edible crab Cancer 91 setosus from the Chilean coast. Journal of Crustacean Biology, 23, 486-494. 92 Gratton P, Allegrucci G, Gallozzi M, Fortunato C, Ferreri F, Sbordoni V (2004) 93 Allozyme and microsatellite genetic variation in natural samples of zebrafish, Danio 94 rerio. Journal of Zoological Systematics and Evolutionary Research, 42, 54-62. 95 Grundmann M, Ansell SW, Russell S, Koch MA, Vogel JC. (2007) Genetic structure of 96 the widespread and common Mediterranean bryophyte Pleurochaete squarrosa 97 (Brid.) Lindb. (Pottiaceae) – evidence from nuclear and plastidic DNA sequence 98 variation and allozymes. Molecular Ecology, 16, 709-722. 99 Isabel N, Beaulieu J, Bousquet J (1995) Complete congruence between gene diversity 100 estimates derived from genotypic data at enzyme and random amplified 101 polymorphic DNA loci in black spruce. Proceedings of the National Academy of 102 Science USA, 92, 6369-6373. 103 Jaramillo-Correa JP, Beaulieu J, Bousquet J (2001) Contrasting evolutionary forces 104 driving population structure at expressed sequence tag polymorphisms, allozymes 105 and quantitative traits in white spruce. Molecular Ecology, 10, 2729-2740. 106 Jenczewski E, Prosperi JM, Ronfort J (1999) Differentiation between natural and 107 cultivated populations of Medicago sativa (Leguminosae) from Spain: analysis with 108 random amplified polymorphic DNA (RAPD) markers and comparison to 109 allozymes. Molecular Ecology, 8, 1317-1330. 110 Krafsur ES (2002) Population structure of the tsetse fly Glossina pallidipes estimated by 111 allozyme, microsatellite and mitochondrial gene diversities. Insect Molecular 112 Biology, 11, 37-45. 113 Larsson LC, Laikre L, Palm S, André C, Carvalho GR, Ryman N (2007) Concordance of 114 allozyme and microsatellite differentiation in a marine fish, but evidence of 115 selection at a microsatellite locus. Molecular Ecology, 16, 1135-1147. 116 Le Corre V, Dumolin-Lapègue S, Kremer A (1997) Genetic variation at allozyme and 117 RAPD loci in sessile oak Quercus petraea (Matt.) Leibl.: the role of history and 118 geography. Molecular Ecology, 6, 519-529. 119 Lee S-W, Ledig FT, Johnson DR (2002) Genetic variation at allozyme and RAPD 120 markers in Pinus longaeva (Pinaceae) of the White Mountains, California. 121 American Journal of Botany, 89, 566-577. 122 Lemaire C, Allegrucci G, Naciri M, Bahri-Sfar L, Kara H, Bonhomme F (2000) Do 123 discrepancies between microsatellite and allozyme vatiation reveal differential 124 selection between sea and lagoon in the sea bass (Dicentrarchus labrax)? Molecular 125 Ecology, 9, 457-467. 126 Mamuris Z, Stamatis C, Triantaphyllidis C (1999) Intraspecific genetic variation of 127 striped red mullet (Mullus surmuletus L.) in the Mediterranean Sea assessed by 128 allozyme and random amplified polymorphic DNA (RAPD) analysis. Heredity, 83, 129 30-38. 130 Meglécz E, Pecsenye K, Varga Z, Solignac M (1998) Comparison of differentiation 131 pattern at allozyme and microsatellite loci in Parnassius mnemosyne (Lepidoptera) 132 populations. Hereditas, 128, 95-103. 133 Pampoulie C, Gysels ES, Maes GE, Hellemans B, Leentjes V, Jones AG, Volckaert 134 FAM (2004) Evidence for fine-scale genetic structure and estuarine colonisation in 135 a potential high gene flow marine goby (Pomatoschistus minutus). Heredity, 92, 136 434-445. 137 Peakall R, Smouse PE, Huff DR (1995) Evolutionary implications of allozyme and 138 RAPD variation in diploid populations of dioecious buffalograss Buchloë 139 dactyloides. Molecular Ecology, 4, 135-147. 140 141 Rocha CD, Del Lama SN, Regitano LCD (2004) Lack of genetic structuring among 142 tropical Brazilian wood stork populations and low genetic differentiation from 143 North American populations. Biotropica, 36, 248-258. 144 Scribner KT, Arntzen JW, Burke T (1994) Comparative analysis of intra- and 145 interpopulation genetic diversity in Bufo bufo, using allozyme, single-locus 146 microsatellite, minisatellite, and multilocus minisatellite data. Molecular Biology 147 and Evolution, 11, 737-748. 148 Scribner KT, Crane PA, Spearman WJ, Seeb LW (1998) DNA and allozyme markers 149 provide concordant estimates of population differentiation: analyses of U.S. and 150 Canadian populations of Yukon River fall-run chum salmon (Oncorhynchus keta). 151 Canadian Journal of Fisheries and Aquatic Sciences, 55, 1748-1458. 152 Sinclair EA, Webb NJ, Marchant AD, Tidemann CR (1996) Genetic variation in the 153 little red flying-fox Pteropus scapulatus (Chiroptera: Pteropodidae): implications 154 for management. Biological Conservation, 76, 45-50. 155 Smith CT, Antonovich A, Templin WD, Elfstrom CM, Narum SR, Seeb LW (2007) 156 Impacts of marker class bias relative to locus-specific variability on population 157 inferences in Chinook Salmon: a comparison of single-nucleotide polymorphisms 158 with short tandem repeats and allozymes. Transactions of the American Fisheries 159 Society, 136, 1674-1687. 160 161 Szmidt AE, Wang X-R, Lu M-Z (1996) Empirical assessment of allozyme and RAPD variation in Pinus sylvestris (L.) using haploid tissue analysis. Heredity 76:412-420. 162 Torres E, Iriondo JM, Pérez C (2003) Genetic structure of an endangered plant, 163 Antirrhinum microphyllum (Scrophulariaceae): allozyme and RAPD analysis. 164 American Journal of Botany, 90, 85-92. 165 Vadopalas B, Leclair LL, Bentzen P (2004) Microsatellite and allozyme analyses reveal 166 few genetic differences among spatially distinct aggregations of geoduck clams 167 (Panopea abrupta, Conrad 1849). Journal of Shellfish Research, 23, 693-706. 168 Vandewoestijne S, Baguette M (2002) The genetic structure of endangered populations 169 in the Cranberry Fritillary, Boloria aquilonaris (Lepidoptera, Nymphalidae): 170 RAPDs vs allozymes. Heredity, 89, 439-445. 171 Volis S, Yakubov B, Shulgina I, Ward D, Mendlinger S (2005) Distinguishing adaptive 172 from nonadaptive genetic differentiation: comparison of QST and FST at two spatial 173 scales. Heredity, 95, 466-475. 174 Williams CL, Fedynich AM, Pence DB, Rhodes OE (2005) Evaluation of allozyme and 175 microsatellite variation in Texas and Florida mottled ducks. Condor, 107, 155-161. 176 Wu J, Krutovskii KV, Strauss SH (1999) Nuclear DNA diversity, population 177 differentiation, and phylogenetic relationships in the California closed-cone pines 178 based on RAPD and allozyme markers. Genome, 42, 893-908. 179 Zeng J, Zou Y, Bai J, Zheng H (2003) RAPD analysis of genetic variation in natural 180 populations of Betula alnoides from Guangxi, China. Euphytica, 134, 33-41. 181