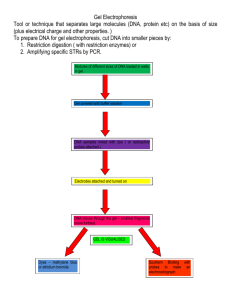
RESTRICTION ENZYMES AND RESTRICTION MAPPING
advertisement

RESTRICTION ENZYMES Objectives 1) To know common problems related to the use of restriction enzymes and how to detect and overcome them. 2) To be able to evaluate the accuracy of sizing DNA fragments on an electrophoretic gel. 3) To know what a Southern Blot is. 4) To know about infrequently cutting restriction endonucleases. General References: Geoffrey G. Wilson, Noreen E. Murray (1991) Ann. Rev. Genet. 25: 585-627. Robert Yuan (1981) Ann. Rev. Bioch. 50: 285-315 Every restriction enzyme manufacturer supplies a wealth of information about its enzymes in its catalogue and in specification sheets supplied with the enzymes. A running compendium of all available restriction enzymes is maintained at http://rebase.neb.com/rebase/. Background about restriction enzymes that you are expected to have: Type I restriction enzymes K & B as an impediment to cloning, and alternative designations of them (ie. hsdR, hsdM, hsdS, rK, mK) Type II enzymes (like EcoRI), use in cloning, interaction with methylases, isoschizomers, types of ends (blunt, 5' recessed, 3' recessed), compatible ends. Dephosphorylation of ends as a means of preventing ligation. Use of synthetic linkers. Expected frequency of 4 base sites, 6 base sites, etc. Modern purposes of restriction digestion Until recently, new clones were often subjected to restriction mapping by double digestion as a means of sorting out which features were on which fragments and merited what kinds of characterization. More recently, clones are usually of known sequence, so inferences from sequence searches are used to answer those kinds of questions. However, it is important to measure restriction patterns for clones and compare them to the patterns expected from the sequence. This reveals rearrangements that happen during clone growth, mislabeled clones, and sometimes errors in the original sequence assembly. Of course restriction enzymes are still used in clone construction. But clone construction often is facilitated with PCR tricks that place fairly light demands on presence of restriction sites. One of the most important purposes of restriction digestions is to verify that a clone has been properly constructed. One of the most common problems encountered is for an unexpected stray fragment to get ligated into a clone in addition to the targetted insert. Similarly, restriction digestion is often a convenient way to determine the orientation of an insert. Practical aspects of using restriction enzymes (Type II restriction endonucleases). Problems related to not enough activity: Restriction enzymes, like other enzymes, have an activity defined by cleaving a certain amount of substrate to completion in a specified time and under specified conditions. Typically a "unit" is defined as cleaving 1 ug of DNA per hour under optimal reaction conditions. Usually the vendor supplies a 10x buffer concentrate for composing your reactions in the optimal conditions. The "1 ug of DNA" contains a trap in that it is the concentration of sites, not the concentration of nucleotides that matters. For example, EcoRI is typically assayed on lambda DNA that has 1 site per 10 kb. You might use it on the PCR product of 100 bp having 1 site engineered into each primer. You will need 200 x as much enzyme as you might have expected. Typically, one plans for 5-10 times more activity than calculated from the advertised units to cover various causes of slow cleavage. These include certain sites that intrinsically cleave slowly, suboptimal conditions (carry over of salt, wrong pH, carry over of competing polyanions like tRNA, etc), misestimated DNA concentration, sites too close to the ends of a fragment, and enzyme preparations that have partially lost activity. If at all possible, one should digest, then withdraw some material to check on a gel, and have enough left over to digest some more, check on a gel again, and still have enough to do the planned experiment. The amount of activity applied can be increased by increasing time instead of just adding more enzyme. The "fold excess" should be calculated as hours times units/ug (adjusted for concentration of sites if appropriate). This, of course, assumes that the enzyme will survive long times under reaction conditions. Stability varies by enzyme, and by the amount of other protein present. One could hope that the manufacturer's specifications sheet that comes with each enzyme would warn if the enzyme is particularly unstable. Long term stability of nearly all enzymes benefit by the presence of carrier protein. Manufacturers generally supply BSA and recommend 1 ug/ul for stabilizing the enzymes. Another good option is 0.5 ug/ul of gelatin. Gelatin can be autoclaved, so it is a good choice if contamination with stray activities is a concern. Reactions of duration greater than over night should be supplemented with a small amount of sodium azide to prevent bacterial growth. Problems related to too much activity. Usually one cannot add more than 10% enzyme stock solution to a reaction without the enzyme storage buffer disturbing the reaction conditions. A common phenomenon is to encourage relaxed specificity due to agents like glycerol in the storage buffer. This is sometimes called "star" activity. Commercial restriction enzymes are not pure, and are generally contaminated with non specific nucleases and phosphatases. Extensive overdigestion typically first damages DNA ends so that the fragments will not ligate, and eventually destroys the integrity of the fragments all together. Exonucleolytic damage produces the paradoxical effect that the fragments appear to smear upwards on electrophoretic gels. This is because single stranded DNA migrates slower than double stranded DNA. Most commercial enzymes are OK at 100 fold over digestion, but not at 1000 fold over digestion. Particularly contaminated enzymes should be ascertainable from the technical specifications sheet provided by the manufacturer. If they say they got complete digestion and > 95% religation at 10 fold overdigestion, that means that it was terrible at 100 fold overdigestion. Troubleshooting. If a target DNA will not cleave, then test the enzyme on a clean control substrate. If the enzyme is OK on its control, then mix experimental and control DNA together and try to cleave. If the experimental DNA causes the control to not cleave, your problem is purity of the experimental DNA. If the control cleaves while the experimental DNA does not, then the problem is with the covalent nature of the experimental DNA. Either it has no sites, or they are modified (eg. by methylation). In the case of modification, you may be able to find enzymes that cleave the same site but have a different sensitivity to modification. You may also remove modifications by conducting in vitro replication through PCR. Assay: Most typically restriction digests are assayed by electrophoresis on a small gel of 0.7 to 1% agarose. This technique is fast and simple. Although polyacrylamide gels have higher resolution for fragments below 500 bp, most people try to get by with higher percentage agarose, or certain brands of agarose engineered for resolution of smaller fragments. Capillary gel electrophoresis is also possible, but the equipment available within the department is configured for fluorescently labelled fragments, and is only applicable to special applications. Visualization: Although other staining agents are available, most commonly the DNA is visualized by staining in 0.5 ug/ml ethidium bromide, which can either be soaked into the gel or placed in the gel during its original formation. Visualization requires illumination with a UV light. A wave length of 265 nm is most effective, but also does more damage to the DNA and is a safety concern. Illumination at 300 nm is common, particularly if the DNA is to be recovered from the gel for cloning. For imaging the gel may be either illuminated from below or above. The investigator should never stare into a gel laying on top of a UV light source, at least not without a UV face shield. There are a number of high tech imaging devices ranging from specialized gel imagers to home-made CCD cameras mounted on laboratory computers. Many of us still use the ancient Poloroid camera setup in the C corridor common room. Sensitivity typically allows visualization down to 5-10 ng in a band. One should always record an image of every gel. The experiment depicted above is suboptimal for a number of reasons. Can you see why? There should be more markers, and there should be markers extending below the lowest experimental band of interest and markers better defining the onset of curvature. The gel could have been run longer to make better use of its revolving power. The major problem in using restriction fragment sizes for any purpose is the likelihood of confusing two fragments that are coincidentally close in size. This problem increases dramatically with the degree of sizing error and the numbers of fragments in the digest. Sizing error can be evaluated by the degree of deviation from a smooth curve within the standard, and by the deviation from the known size of some fragment in a second lane. The vector is often used for the latter. Error > 5% is due to poor technique. Accuracy down to 3% can be achieved with standards and unknowns loaded in the same lane to remove lane-to-lane variability. Lane-to-lane variability can be due to differing concentrations of salt in the different samples. Below 3% uncertainty, the base composition of fragments, and the nature of the ends may become a factor, particularly on small fragments. Understanding circular DNA Plasmid DNA as isolated from cells is a mixture of forms: monomer covalently closed circle (ccc) (supercoiled), dimer and higher forms of ccc, and nicked circles (alias relaxed circles). The supercoils generally run as if they were about half the size of the corresponding linear, although the relationship between ccc and linear migration is dependent on gel conditions. The sizing curve for circles remains linear higher up on the gel. Quantitation of clones using restriction digests: A restriction digest of some standard molecule, such as bacteriophage lambda DNA, will serve as both a size marker and a quantitation standard curve. Each fragment is present in an amount in micrograms that is in proportion to its size in bp. Either by eye, or by a quantitative scan, one may quantitate an unknown against the resulting standard curve. Be careful about saturation. For example, a photographed band is often white. This is saturated, meaning that the film can't get any whiter. So one can't compare two saturated bands and know which one was "brighter". Intuitively, people tend to observe the band width in this case. But beware that bands also get broader for smaller fragments. Paying attention to the quantitation after digestion where there are bands of several different sizes to judge often gives the first warning that the initial estimate of DNA quantity was way off. Infrequently cutting enzymes for mammalian DNA: NotI BssHII MluI NruI SfiI SalI ClaI GC^GGCCGC G^CGCGC A^CGCGT TCG^CGA GGCCN4^NGGCC G^TCGAC AT^CGAT ~4x106 ~3x105 ~3x105 ~3x105 ~7x104 ~7x104 ~7x104 Infrequently cutting enzymes are sometimes included in cloning vectors as a site for recleavage that is unlikely to be present in the insert. Infrequently cutting enzymes are sometimes used for examination of genomic DNA to get a broad scale verification by restriction mapping of a sequence assembly, or to investigate the distribution of CG modifications. CG is more rare in mammalian DNA than one would expect in random sequence. The frequency of CG is depressed in mammalian DNA because methylation at the C promotes mutation to a T by deamination. Also, many of the sites are not cleaved because they are methylated. Most of the unmethylated CGs in the genome are clustered in "CG islands". Examination of large restriction fragments requires a special electrophoresis system called "pulsed field electrophoresis". In ordinary gel electrophoresis, small DNA fragments migrate as a random coil reflecting the length of the molecule. For large fragments, the random coil elongates in the direction of migration, such that increased size ceases to be a factor in its migration. In pulsed field electrophoresis, the direction of the voltage is frequently shifted to prevent this effect. There are several versions of the method. To get very large fragments into the gel without shearing them, the cells are often placed intact in an agarose plug, and lysed and cleaved in situ. For visualization of individual fragments within the context of an entire digested genome, a hybridization method called a Southern Blot is used. Southern blotting Alternatives: Southern blotting is nearly always substituted by some combination of PCR and DNA sequencing whenever possible these days. One common use remaining is to characterize transgenes in transgenic animals, particularly in untargetted constructs. Southern blotting is used to tell which bands on a gel share sequences with some hybridization probe. The DNA in the gel is denatured with alkali, and then transferred onto a piece of filter paper by one of a variety of methods. The hybridization probe is radiolabeled, denatured, and allowed to reanneal with the DNA affixed to the filter. The filter is washed under conditions that remove probe that is not annealed to DNA on the filter, and then the filter is subjected to autoradiography. If the probe is not perfectly matched to the target, or if there are related sequences that cross hybridize in the clone, then the washing conditions can be adjusted to try to include or exclude hybrids with some particular degree of mismatching. The degree of tolerance to mismatching is called "stringency". Stringency High Moderate Low Mismatch tolerated 0 - 8% 0 - 20% 0 - 30% Temp.(in 1.5 mM Na+) 62 52 42 The washing is a kinetic experiment, not an equilibrium one. The above conditions completely remove the signal from hybrids below stringency in about an hour of washing. The wash buffer should be frequently changed to keep reannealing from occurring. Last revised 2/21/05 - Steve Hardies