III. Circadian Structure in Drosophila
advertisement

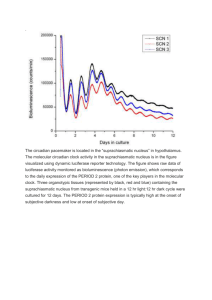
1 Circadian Systems: Progress towards Systemlevel Understanding of Circadian Rhythms Jeffrey J. Kang Abstract—Circadian rhythms manifest in the physiology and behaviour of many organisms, from bacteria to humans. Using the systems biology approach, cellular systems that generate circadian rhythms are systematically perturbed and characterized. Models of the relevant molecular mechanisms in Drosophila have been formulated and serve as roadmaps in the modeling of circadian systems in mammals. Numerical simulations are used to validate molecular models and provide insight into the behaviour of circadian systems. These models are constantly evolving as progress is made towards a system-level understanding of circadian rhythms. Index Terms—circadian, Drosophila, systems biology, clock genes. I. INTRODUCTION B iological clocks have been studied by scientists since the seventeenth century when it was observed that the leaves of the heliotrope plant open and close in adherence to a daynight rhythm even in the absence of light stimuli [1]. Since then, research has shown that endogenous rhythms with a period of approximately 24 hours manifest in almost all eukaryotic organisms as well as some bacteria. Named circadian clocks for the Latin term circa dies, meaning about one day, these biological rhythms effect changes in physiology and behaviour that allow organisms to anticipate and adapt to environmental changes brought about by the day-night cycle. In recent years, the application of systems biology techniques has provided new insight into the inner workings of circadian clocks, revealing a cellular oscillator comprised of a network of interlocking positive and negative transcriptional feedback loops [2]. Characterization of the clock circuitry has been performed by systematically perturbing circadian systems using genetic manipulation. By using targeted gene deletions, the behaviour of circadian rhythms in mutants can be observed to infer gene function and interaction. Advancements in highthroughput gene manipulation and gene expression measurement techniques have increased the efficiency of these experiments. These mutagenesis studies have led to the identification of clock genes and the roles they play in generating circadian rhythms. Models of circadian systems on the molecular levels for several organisms have been created and refined. Although these models are not yet complete, they will improve as the complexities of circadian circuitry are unravelled. Circadian systems are well suited to study using the systems biology approach. The emergence of systems biology theory and its associated tools have greatly benefited research in this field and aided progress towards a system-level understanding of circadian rhythms, whereby circadian systems may be controlled and designed. Such an understanding may lead to treatments for sleep-related ailments and certain neurological disorders in humans. II. THE CIRCADIAN SYSTEM In terms of classic systems theory, the circadian clock can be modelled as a system that receives input consisting of environmental light-dark signals and produces output in the form of a biological rhythm expressed in physiology and behaviour. The system can be thought of as an autonomous oscillator producing a signal of fixed frequency with phase changes determined by the input. Although some elements of the system’s molecular mechanisms have been discovered, the exact structure of the system is still unknown. The molecular basis of circadian systems has been studied in the most detail in the fruitfly Drosophila melanogaster, the organism in which the first clock gene per was isolated in 1984. It was observed that transcription of the per gene is induced when levels of its own product, the PER protein, are reduced [3]. This forms a negative autoregulatory feedback loop creating oscillations in transcription of the per gene, thus providing an early model for the circadian oscillator. However, as experimental advancements have been made, this model has undergone many changes, reflecting the need in systems biology to improve the system model iteratively. III. CIRCADIAN STRUCTURE IN DROSOPHILA Current models of circadian oscillators in Drosophila consist of multiple interlocking feedback loops. Both negative and positive feedback mechanisms are believed to exist. The negative feedback mechanism is based on two transcriptional activators, dCLK and CYC, and two repressors, PER and TIM. Fig. 1 shows a diagram of the molecular model proposed in [4]. The proteins dCLK and CYC form a heterodimer which bids to E-boxes in the regulatory regions of the per and tim genes, enhancing their transcription. After PER and TIM have reached a threshold concentration, they associate and enter the nucleus and bind to the dCLK-CYC heterodimers, thereby 2 IV. COMPUTATIONAL MODELING AND ANALYSIS Fig. 1. Molecular model of Drosophila circadian system. repressing per and tim transcription. Positive feedback is seen in the dCLK protein which represses dclk gene transcription. If dclk expression is slightly activated then the formation of dCLK-CYC heterodimers enhances per and tim transcription. As PER and TIM bind to dCLK-CYC, the repression of dclk by dCLK is relieved. This further increases dCLK levels, and hence a positive feedback loop is formed [5]. The circadian rhythm is produced by the alternating cycles of activation and repression of per and tim. A sustained oscillation is generated by separating the intervals of activation and repression [6]. The kinase DBT plays an important role in generating these delays by phosphorylating and destabilizing PER. Thus, DBT retards the accumulation of PER. Each activation-repression cycle lasts approximately 24 hours. Light input affects the system via the CRY protein which acts as a photoreceptor. When exposed to light, CRY associates with TIM, which promotes rapid TIM degradation. This reduction in TIM levels causes a phase delay in the circadian cycle. Two genes not shown in Fig. 1 are sgg and vr, whose roles are not fully understood, but appear to be ancillary. The heterodimer dCLK-CYC activates vri transcription in the same manner as it activates per and tim. High levels of VRI subsequently repress per and tim expression, though the mechanism by which this occurs is unknown. SGG plays a role in determining the timing of PER-TIM translocation to the nucleus by phosphorylating TIM. The multiple negative and positive transcription regulatory factors within circadian systems are necessary to ensure reliable oscillation with relatively constant periods in the presence of stochastic biochemical noise and under varying cellular conditions. The stochastic nature of reaction events is important when the number of molecules is small as is the case in many cellular systems. This behaviour can be regarded as internal noise of the system. Cellular conditions such as transcription and translation rates may vary with changes in temperature, nutrition or growth conditions [7]. The complex feedback loops present within the regulatory mechanisms create a robust periodicity, reducing the sensitivity of the system to noise and changes in cellular conditions. Computational models of circadian rhythms can be developed based on the molecular model. Computational analysis can then offer theoretical predictions based on simulations of the system’s molecular mechanisms. In [8], a procedure for developing a computational model of biological rhythms is proposed. This procedure can be summarized in four steps. 1) Variable identification: The key variables of the phenomenon to be modeled are identified. Additionally, the nature of the interactions between variables forming the relevant feedback loops is characterized. 2) Mathematical description: A system of differential equations describing the time evolution of the system is formulated. For spatially homogenous conditions, these equations are ordinary differential equations. Otherwise, as is the case in the presence of diffusion, partial differential equations are used to describe the system’s evolution in both the spatial and temporal domains. 3) Calculation of steady states: The steady states of the system, as determined by the differential equations, are solved using analytical methods or by numerical integration. 4) Steady state analysis: The stability properties of the steady states are determined using a method such as linear stability analysis. If a steady state is deemed to be unstable then the evolution of the system to either a stable steady state or sustained oscillations is calculated. In the case of sustained oscillations, the period and amplitude are described as functions of the system parameters. A computational model allows predictions of system behaviour to be obtained via numerical simulations using known or assumed parameter values. Theoretical predictions can then be compared to experimental observations to evaluate the accuracy of the model. The model is then modified if incongruity between prediction and experimental observations exists, or when additional information regarding the modeled variables and interactions is discovered. The first computation models for circadian rhythms were proposed for Drosophila. An early model based on the negative control of the PER protein on per expression consisting of only five kinetic equations was developed using the procedure detailed above [8]. Numerical simulations produced the desired sustained oscillations, but the steady state became unstable for some parameter values. As understanding of the system structure improved, a ten-variable model was proposed based on the negative control exerted by the PERTIM heterodimer. This model also accounted for the effects of light in entraining the circadian system. Simulations showed sustained oscillations in complete darkness with phase shifts induced by light pulses. However, models based on continuous differential equations are deterministic and only approximate reaction kinetics. If the number of molecules in a system is small, the model does not accurately portray the stochastic effects of biochemical noise. When the discrete nature of reaction events is taken into account, the simulated oscillations 3 Fig. 2. Comparison of simulated and experimental PER concentrations. persist but with widely fluctuating periods and amplitudes [7]. Thus a flaw is exposed in the molecular model upon which the computation model was formulated. In [7], an alternative model incorporating hysteresis in the negative regulation of transcription produces more accurate results when consideration is given to noise in reaction kinetics. Simulations show that even with as few as ten molecules per cell, the standard deviation of the period remains less than 10%. While this hysteresis model may not accurately portray the molecular mechanisms at work in the Drosophila circadian system, it illustrates the need for models to be robust to stochastic noise. It has been hypothesized that the addition of positive feedback elements to existing models may improve their sensitivity to noise [9]. A more recent 23variable model includes both positive and negative feedback loops. Simulation of this model suggests that positive feedback elements are not necessary to generate circadian rhythms [5], but no evaluation of their effect on the robustness of the model was made. The true importance of these positive feedback elements remains unclear. Recently proposed models provide simulated output of protein levels that closely match experimental data. Fig. 2 shows a comparison between simulated levels of gene product in the Drosophila circadian system obtained using the model proposed in [5] compared to experimental observations. In this diagram, “PER” refers to an average of PER and TIM protein levels. The peak, minimum and shape of the protein level progression over the course of the circadian cycle are similar in the simulated and experimental cases. The period of simulated cycle is approximately 24 hours. Simulations such as this one show that current understanding of the molecular mechanism governing circadian rhythms, while incomplete, accurately describes the key elements of the circadian system. V. CIRCADIAN RHYTHM IN MAMMALS The study of model organisms is useful in inferring the function of human genes. These organisms have been described as the Rosetta Stones for deciphering human biological systems [11]. In the study of human circadian systems, Drosophila serves as an excellent model organism. Understanding of the molecular mechanisms in the Drosophila circadian clock has greatly assisted the study of their counterparts in mammals. Homologues of most Drosophila clock genes have been identified in mammals [3]. The transcriptional feedback loops that generate oscillations in mammalian circadian systems are more complex than those found in Drosophila and the key proteins are different, but important similarities exist. Models of mammalian circadian systems have been developed based on various mutagenesis studies conducted on mice. As in Drosophila, the rhythm is based on a core negative feedback loop. The two transcription factors CLOCK and BMAL1 (MOP3) form a heterodimer that binds to the E-boxes in the promoter regions of mPer1, mPer2, mPer3, mCry1 and mCry2, thereby stimulating the transcription of the three mPer and two mCry genes [4]. This is analogous to the effect of the dCLK-CYC heterodimer on PER in Drosophila. The mPer and mCry genes are the respective homologues of per and cry. The mCRY proteins act as the transcription repressors instead of TIM and PER in Drosophila. By entering the nucleus and interfering with the activation of the CLOCK-BMAL1 heterodimer, the mCRY proteins repress the transcription of the mPer and mCry genes. Some mechanisms of the mammalian circadian system are still unclear. While mPER2 is believed to assist in the transcription of the bmal gene, the functions of mPER1 and mPER3 are unknown. To date, no homologue for tim has been discovered for mammals; however, genome-wide complex trait analysis in mice indicates the presence of undiscovered clock genes [2]. In addition to understanding how circadian rhythms are generated at the cellular level, it is also important to understand how the circadian system operates on an organism level to induce changes in physiology and behaviour. In organisms such as birds, reptiles and fish, the circadian clock is localized in specific areas of the central nervous system such as the pineal gland. However, the mammalian circadian system consists of a hierarchy of dispersed oscillators [1]. The center of this system is located in the suprachiasmatic nucleus (SCN) of the anterior hypothalamus of the brain. In mice, destruction of the SCN renders the animal behaviourally and physiologically arrhythmic. In humans with craniopharyngioma, the tumour can constrict the SCN region of the brain, leading to severe disruption of sleep-wake cycles [10]. The SCN can be regarded as a master clock that coordinates the timing of slave clocks in other areas of the brain and in peripheral organs such as the liver and the kidney. Synchronized slave clocks then regulate local rhythms that govern physiology and behaviour. Unlike the master clock, slave clocks are unable to produce self-sustaining oscillations and require periodic input from the master clock. SCN cells in culture produce circadian rhythm for weeks, but the rhythm produced by liver cells in culture dampen and disappear in under one week [2]. The differences 4 between the oscillation mechanisms within cells comprising the master clock and slave clocks are unknown. Genetic studies show that the molecular compositions of the master clock and slave clocks are very similar. One hypothesis states that the master clock benefits from global differences in the levels and kinetics of clock proteins rather than from the existence of dissimilar genes or proteins in the SCN [2]. The SCN contains approximately 10,000 to 20,000 neurons. Each neuron exhibits its own circadian rhythm with varying periods, though the mechanism of how individual oscillations are coordinated to generate a single overt rhythm is unknown. The SCN is often modeled as a single oscillator for the sake of simplicity. However, this simple model is not always valid as spatially inhomogeneous rhythms have been observed within regions of the SCN. One study has attempted to reconcile this observation by modelling the SCN as a system of 10,000 neurons behaving as locally-coupled, self-sustained oscillators [12]. Simulation of this system showed that a global rhythm emerged due to local coupling between oscillators leading to synchronization. The period of the global output corresponded to the average periods of the individual oscillators. The period of the rhythm was found to be stable for low amplitudes as well as amplitude transients. This model presents one possible explanation for SCN rhythm generation, though molecularlevel understanding of neuron synchronization is lacking. The input and output pathways of the mammalian circadian system have also been a focus of study. Input pathways provide external stimulation to the system in the form of environmental inputs that entrain the circadian rhythm. In humans, it has been observed that the daily cycles of sleeping, waking, activity and hunger conform to a period of about 23 hours if the body is deprived of environmental cues associated with the day-night cycle [1]. Therefore, environmental information, primarily taking the form of observed light and dark, is required to extend this period to 24 hours. In mammals, the signalling pathway that carries entraining signals to the SCN has been identified as the retinohypothalamic tract (RHT) [2]. The RHT delivers photic input from retinal ganglion cells to SCN neurons. These visual cues are believed to be necessary and sufficient to synchronize circadian rhythms to 24 hour day-night cycles. The output pathways by which the SCN master clock synchronizes slave clocks are believed to the hormonal in nature. Neurons secrete several neuropeptides such as the hormone vasopressin based on a circadian rhythm [1]. Additionally, neurons send signals to the pineal gland, resulting in production of melatonin, another hormone. VI. CONCLUSION Circadian rhythms and their manifestation in physiology and behaviour have long been observed. In recent years, advances in genetics have allowed the molecular mechanisms—the so called clockwork—of circadian rhythms to be studied. Models of circadian systems of organisms such as Drosophila emerged first from these studies. These models have been invaluable in the modeling of circadian systems of more complex organisms such as mammals thanks to similarities in the circadian systems of disparate species at the molecular level. Current models are constantly evolving as new experimental observations are made and fresh insight is gained. The role played by systems biology in the study of circadian systems is crucial. Systems biology provides a strategy to deal with the immense complexity of circadian systems. Through the use of incrementally improving models and systematic perturbations of circadian systems, an understanding of the relationships between the various genes and proteins that form the core of the circadian systems has been gained. Computational models and simulations have been used to confirm or invalidate hypotheses regarding the structure and kinetics of the molecular mechanisms of circadian systems. Simulations are also used to make predictions of system behaviour, providing insights that are sometimes counterintuitive and difficult to obtain by other means. The culmination of these studies will be a system-level understanding of circadian systems in not just Drosophila and mice, but also humans. Such an understanding would enable circadian systems to be characterized, controlled and perhaps even created. This may lead to new avenues for pharmacological treatments for conditions such as jetlag and ailments afflicting shift workers, as well as sleep-related psychiatric disorders and SCN-based neurological disorders. REFERENCES P. Sassone-Corsi, “Molecular clocks: mastering time by gene regulation,” Nature, vol. 392, pp. 871-874, April 1998. [2] S. M. Reppert and D. R. Weaver, “Coordination of circadian timing in mammals,” Nature, vol. 418, pp. 935-941, August 2002. [3] H. Okamura, S. Yamaguchi, and K. Yagita, “Molecular machinery of the circadian clock in mammals,” Cellular Tissue Research, vol. 309, pp. 47-56, July 2002. [4] J. A. Ripperger and U. Schibler, “Circadian regulation of gene expression in animals,” Current Opinion in Cell Biology, vol. 13, pp.357-362, June 2001. [5] P. Smolen, D. A. Baxter, and J. H. Bryne, “Modeling circadian oscillations with interlocking positive and negative feedback loops,” Journal of Neuroscience, vol. 21, pp. 6644-6656, September 2001. [6] M. W. Young and S. A. Kay, “Time zones: a comparative genetics of circadian clocks,” Nature Reviews Genetics, vol. 2, pp. 702-715, September 2001. [7] N. Barkai and S. Leibler, “Circadian clocks limited by noise,” Nature, vol. 403, pp. 267-268, January 1999. [8] A. Goldbeter, “Computational approaches to cellular rhythms,” Nature, vol. 420, pp. 238-245, November 2002. [9] M. H. Hastings, “Circadian clockwork: two loops are better than one,” Nature Reviews Neuroscience, vol. 1, pp.143-146, November 2000. [10] H. D. Piggins, “Human clock genes,” Annals of Medicine, vol. 34, pp. 394-400, May 2002. [11] T. Ideker, T. Galitski, and L. Hood, “A new approach to decoding life: systems biology,” Annual Review of Genomics and Human Genetics, vol. 2, pp. 343-372, September 2001. [12] H. Kunz and P. Achermann, “Simulation of circadian rhythm generation in the suprachiasmatic nucleus with locally coupled self-sustained oscillators,” Journal of Theoretical Biology, vol. 224, pp 63-78, September 2003. [1] 5