MEC_4909_sm_FigS1-Table-S1-2
advertisement

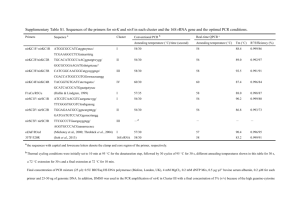
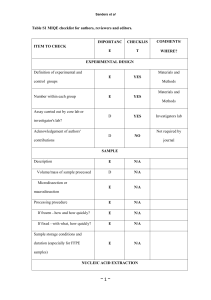
Supplementary information Cyt b diversity from nested PCR Sequences obtained from the nested PCR contained 12 unique Plasmodium cyt b lineages and 3 unique Haemoproteus lineages. Four Plasmodium lineages were common in this dataset, in line with previous findings (Wood et al., 2007); among single lineage infections, pSGS1 was most abundant (45% infections), followed by pTURDUS1 (36%), pBT7 (14%) and pGRW11 (3%). The other previously documented lineages found were pBLUTI1 (n=3 samples), pBLUTI2 (n=3), pBLUTI3 (n=1), pBLUTI4 (n=1) and pBLUTI5 (n=1), hBLUTI1 (n=1) and H. majoris (n=1) Four previously unreported cyt b lineages were found, in one sample each: three Plasmodium lineages (pBLUTI6, pBLUTI7, pBLUTI8) and one Haemoproteus lineage (hBLUTI2). These sequences have been deposited in Genbank (Accession numbers HQ123330 to HQ123333) and the MalAvi database (Bensch et al. 2009). Species diagnosis by qPCR Based on published comparisons of morphological and molecular data, we denoted pSGS1 and pGRW11 as belonging to P. relictum, and pTURDUS1 and pBT7 as belonging to P. circumflexum (Palinauskas et al. 2007). Agreement in species diagnosis by qPCR and sequence analysis was high; in 96% of infections for which nested PCR gave a single lineage diagnosis (n=302), the species assigned by cyt b sequence and qPCR product melting temperature was identical. In the twelve cases of disagreement, 10 involved assignment of P. circumflexum by qPCR where nested PCR assigned P. relictum, and 2 involved the reverse situation. Thus there is a small degree of error in species-assignment by this method (4% infections, assuming nested PCR assignment is 100% accurate). However, it has the benefit of allowing many samples for which a sequence is difficult to obtain because of low parasitaemia to be assigned to species. Three of four samples diagnosed as containing Haemoproteus by nested PCR yielded qPCR products, suggesting the assay may also detect this parasite. These samples were excluded from analysis of Plasmodium prevalence and parasitaemia. Of the samples known to contain a Plasmodium lineage other than pSGS1, pGRW11, pTURDUS1 or pBT7, only two samples with lineage pBLUTI2 produced a qPCR signal, and their melting temperatures clustered 1 with P. circumflexum, as is expected from their high sequence similarity to pTURDUS1 and pBT7 (Wood et al. 2007). Comparative sensitivity of qPCR and nested PCR Infection prevalence by qPCR was significantly higher than by nested PCR, for all Plasmodium infections combined and for P. relictum, but not P. circumflexum (Table S1), indicating that sensitivity of the qPCR assay is considerably higher than that of nested PCR. However, since during nested PCR each sample was tested only once whereas in qPCR each sample was tested in triplicate we also compared sensitivity of these two methods when only one replicate was considered. When the number of replicates is held constant there is little evidence for a difference in sensitivity of the two assays, with a tendency towards greater detection by nested PCR (χ21=2.99, p=0.084, n=342). Since increased detection by qPCR is apparently due to greater replication during screening, we examined among samples tested twice by this method (and thus for which 6 replicates were available), how prevalence changed as the number of replicates increased. For both Plasmodium species, prevalence increased dramatically from one to three replicates and began to plateau after this (Fig. S1). However, the total percentage increase in prevalence from a single replicate to six was strikingly large (with several clean negative controls per plate), between 59-67% depending on the species, which highlights the difficulty of detecting chronic Plasmodium infections even with sensitive molecular diagnostics. qPCR appeared to be less sensitive at detecting mixed-species infections than nested PCR. Of the 7% (23/334) infections identified as containing both P. relictum and P. circumflexum by clear double peaks on electropherograms, only 3 contained two appropriate double melting peaks in the qPCR, and such double melting peaks were also observed in 6 infections not diagnosed as mixed by sequence analysis. 2 Table S1: Comparison of Plasmodium detection by the nested PCR method of Waldenström et al. (2004), and a novel quantitative PCR assay, among samples tested by both methods. Prevalence estimates and 95% confidence intervals are shown. All Plasmodium Year N 2005 P. relictum P. circumflexum nested PCR qPCR nested PCR qPCR nested PCR qPCR 405 0.264 (0.223, 0.309) 0.483 (0.434, 0.531) 0.117 (0.089, 0.152) 0.296 (0.253, 0.342) 0.162 (0.129, 0.201) 0.211 (0.174, 0.254) 2006 416 0.351 (0.307, 0.398) 0.394 (0.348, 0.442) 0.173 (0.140, 0.212) 0.188 (0.153, 0.228) 0.200 (0.164, 0.241) 0.204 (0.168, 0.246) 2007 436 0.314 (0.272, 0.359) 0.443 (0.397, 0.490) 0.195 (0.160, 0.235) 0.296 (0.255, 0.340) 0.152 (0.121, 0.189) 0.133 (0.104, 0.168) All years 1254 0.310 (0.285, 0.336) 0.439 (0.412, 0.467) 0.163 (0.143, 0.184) 0.260 (0.236, 0.285) 0.171 (0.151, 0.193) 0.182 (0.161, 0.204) 3 Table S2: Results from GLMM modelling of Plasmodium infection status (prevalence) variation across the Wytham blue tit population. All variables included in the starting model are shown, with effects in the minimal model in bold. For significant terms (p<0.05) statistics are from the minimal model; for non-significant terms, statistics are those at the point that factor left the model. Effect sizes (Pearson’s r) are given for all effects with a single degree of freedom. Predictor Minimum age Minimum age2 Year x y x*y Sex (F) Species (P. relictum) x*Species y*Species Sex*Species Minimum age*Species Minimum age2 *Species Year*Species x*y*Species †Denominator df Den df† F p r 1 1 3 1 1 1 1 1 1 1 1 1 1 3 1 1824 1824 1824 1824 1824 1824 1824 1824 1824 1824 1824 1822 1821 1824 1823 10.37 4.59 5.83 12.17 8.35 2.17 0.21 39.64 10.44 116.44 6.45 3.24 0.66 18.55 3.56 0.001 0.032 <0.001 <0.001 0.004 0.141 0.646 <0.001 0.001 <0.001 0.011 0.072 0.416 <0.001 0.059 0.075 -0.050 -0.081 0.068 -0.035 -0.011 0.146 0.075 0.245 0.059 0.042 0.019 0.044 degrees of freedom 4 Fig. S1: Increase in cumulative Plasmodium prevalence as more qPCR tests are performed. Means and 95% confidence intervals are shown. Analysis of age-related changes in parasitaemia Mixed models are commonly used to test for within-individual changes in traits with age. However, a significant age effect in such a model does not definitively indicate withinindividual effects when datasets contain missing values, i.e. where not all individuals are present at all ages, as in this study. In this case, selective appearance or disappearance of individuals can also contribute towards the age effect. We used the mixed model approach of van de Pol & Verhulst (2006) to distinguish age-related within-individual increases in parasitaemia from selective appearance or disappearance of individuals with either low or high parasitaemia with age. To do this we modelled parasitaemia among individuals infected in more than one year, including fixed effects of minimum age, minimum age at which an individual was first present in the dataset (first age), minimum age at which an individual was last present in the dataset (last age) and year. Individual identity was included as a random effect. Here minimum age represents the within-individual effect, whilst selective appearance and disappearance are tested for by including first age and last age. We found no evidence for selective appearance or disappearance of individuals with respect to parasitaemia (first age: F1,94=0.47, p=0.497; last age: F1,94=0.01, p=0.914). Of these two effects, only the very weak negative effect of last age is in a direction that could contribute to the overall positive 5 relationship between age and parasitaemia detected at the population level. Whilst controlling for these two effects, there was a non-significant effect of minimum host age in the expected direction (parasitaemia increasing within individuals as they age; F1,169=3.00, p=0.085). When first age and last age were sequentially excluded from the model, the effect of minimum age became significant (F=1,121=4.21, p=0.043). This increase in significance is likely because age, first age and last age are to an extent collinear. References Bensch, S., Hellgren, O., & Pérez-Tris, J. (2009) MalAvi: A public database of malaria parasites and related haemosporidians in avian hosts based on mitochondrial cytochrome b lineages. Molecular Ecology Resources 9, 1353–1358. Palinauskas, V., Kosarev, V., Shapoval, A., Bensch, S., & Valkiūnas, G. (2007) Comparison of mitochondrial cytochrome b lineages and morphospecies of two avian malaria parasites of the subgenera Haemamoeba and Giovannolaia (Haemosporida: Plasmodiidae). Zootaxa, 1626, 39-50. Wood, M.J., Cosgrove, C.L., Wilkin, T.A., Knowles, S.C.L., Day, K.P., & Sheldon, B.C. (2007) Withinpopulation variation in prevalence and lineage distribution of avian malaria in blue tits, Cyanistes caeruleus. Molecular Ecology, 16, 3263-3273. 6