emi412145-sup-0002-si
advertisement

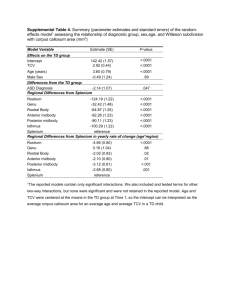
Supporting Information Appendix S1: More information on the robust regression method used in this study Figure S1: SIGI-HMM predictions in 54 Escherichia coli strains The left panel shows SIGI-HMM predictions (cyan regions) from the genomes of 54 E. coli strains, colored blue according to genomic %AT from light blue (most AT rich) to dark blue (most GC rich). The horizontal axis designates the chromosomal position in base pairs (bp), while the vertical axis to the right indicates total size of predicted GIs (bp). The right panel shows more detailed annotations of E. coli O26:H11 strain 11368, the E. coli with the largest genome, and E. coli K-12 substrain MG1655, a non-pathogenic model organism with one of the smallest E. coli genomes. Genes ending with an underscore indicate multiple variants of the gene. ECO26_P_ designates multiple variants of E. coli O26:H11 prophages. More details regarding these annotations can be found in tables S5 and S6 for E. coli K-12 and E. coli O26:H11, respectively. Graphical representation dissimilarities of HT regions between the genomes in the left and right panels are due to scale differences. Table S1: Results from phylum level genome size versus AT content robust regression analysis The table includes regression estimates (column 2) for the bacteria in each phylogenetic group and Proteobacteria subphyla (column 1) as well as standard error (column 3), tstatistic (column 4), p values (column 5) and number n of strains included in the analysis (column 6). Group/subphyla Actinobacteria Estimate Std. Error -17,54 1,24 Firmicutes -0,75 Tenericutes 0,25 Bacteroides-Chlorobi T p n -14,2 0,0001 214 0,75 -1 0,32 423 0,57 0,44 0,66 54 -5,55 2,16 -2,57 0,01 93 Alpha-proteobacteria -7,86 0,39 -20,02 0,0001 215 Beta-proteobacteria -8,26 0,76 -10,81 0,0001 120 Delta-proteobacteria -4,36 1,69 -2,58 0,01 43 0,43 0,19 2,2 0,03 74 -10,88 0,48 -22,77 0,0001 429 Epsilon-proteobacteria Gamma-proteobacteria Chlamydiae-Verrucombia 5,75 0,01 673,46 0,0001 49 Cyanobacteria -3,83 2,29 -1,67 0,1 44 Spirochaetes -2,56 1,66 -1,54 0,15 47 Table S2: Result from species level genomic size vs AT content robust regression analysis The table shows the results from regression analyses between genome size and genomic %AT in the strains of the designated species in column 1. Column 2 shows the regression estimates, column 3 - standard error, column 4 - t statistic, column 5 - p value and column 6 the number n of strains included in each regression model. Species Estimate Std. Error t p n Chlamydia trachomatis 20,56 1,34 15,34 0,0001 21 Neisseria meningitides 17,1 5,29 3,34 0,006 14 Helicobacter pylori 11,45 3,76 3,04 0,004 38 149,64 22,21 6,74 0,0001 54 -2,7 44,59 -0,06 0,95 25 -103,5 45,82 -2,26 0,04 12 Acinetobacter baumannii 2,17 31,89 0,07 0,95 10 Francisella tularensis 2,88 0,31 9,34 0,0001 10 Buchnera aphidicola 0,02 0,01 2,23 0,05 11 Bacillus cereus 0,48 8,6 0,06 0,95 18 Escherichia coli Salmonella enterica Yersinia pestis Clostridium botulinum -28,02 8,31 -3,37 0,006 13 Listeria monocytogenes 47,27 7,56 6,25 0,0001 22 Staphylococcus aureus -11,8 12,58 -0,94 0,35 31 33,3 12,9 2,58 0,02 19 33,16 8,66 3,83 0,002 14 10,5 12,99 0,81 0,43 16 0,6 0,52 1,16 0,27 14 -19,05 11,65 -1,64 0,13 13 Corynebacterium pseudotuberculosis -50,1 8,79 -5,7 0,0001 15 Prochlorococcus marinus -3,59 0,32 -11,22 0,0001 12 Streptococcus pneumoniae Streptococcus suis Streptococcus pyogenes Mycobacterium tuberculosis Corynebacterium diphtheriae Table S3 - Robust regression analysis of relative entropy versus genomic %AT species level The table shows the results from regression analyses between genome-based relative entropy and genomic %AT in the strains of the designated species in column 1. Column 2 shows the regression estimates, column 3 - standard error, column 4 - t statistic, column 5 - p value and column 6 the number n of strains included in each regression model. Species Estimate Std. Error T p n Chlamydia trachomatis 0,05 0,08 0,67 0,51 21 Neisseria meningitides -0,5 0,06 -7,72 0,0001 14 Helicobacter pylori -0,04 0,12 -0,34 0,74 38 Escherichia coli -0,67 0,06 -12,17 0,0001 54 Salmonella enterica -1,04 0,04 -23,91 0,0001 26 Yersinia pestis -0,03 0,14 -0,22 0,83 12 0,01 0,1 0,05 0,96 10 Francisella tularensis 0,2 0,07 2,94 0,02 10 Buchnera aphidicola 0,24 1,00E-03 158,37 0,0001 11 Acinetobacter baumannii Bacillus cereus -0,07 0,07 -1,12 0,28 18 0,72 0,06 12,62 0,0001 13 Listeria monocytogenes -0,55 0,04 -13,31 0,0001 22 Staphylococcus aureus 0,05 0,05 0,9 0,38 31 Streptococcus pneumoniae -0,24 0,07 -3,19 0,005 19 Streptococcus suis -0,12 0,09 -1,43 0,18 14 0,01 0,12 0,1 0,92 16 Mycobacterium tuberculosis -0,06 1,60E-03 -36,58 0,0001 14 Corynebacterium diphtheriae -0,04 0,09 -0,46 0,65 13 0,09 0,06 1,46 0,17 15 -0,03 0,01 -2,43 0,04 12 Clostridium botulinum Streptococcus pyogenes Corynebacterium pseudotuberculosis Prochlorococcus marinus Table S4 - Robust regression analysis of relative entropy versus genome size species level The table shows the results from regression analyses between genome-based relative entropy and genome size in the strains of the designated species in column 1. Column 2 shows the regression estimate, column 3 - standard error, column 4 - t statistic, column 5 - p value and column 6 the number n of strains included in each regression model. Species Estimate Std. Error T p Chlamydia trachomatis -7,78 0,79 -9,79 Neisseria meningitidis -32,69 8,52 Helicobacter pylori -20,69 4,84 -280,34 n 0,0001 21 -3,84 0,002 14 -4,27 0,0001 38 21,97 -12,76 0,0001 54 26,41 23,84 1,11 0,28 26 107,03 126,28 0,85 0,42 12 -133,97 108,25 -1,24 0,25 10 Francisella tularensis -1,63 6,43 -0,25 0,81 10 Buchnera aphidicola 0,05 3,00E-02 1,75 0,11 11 Bacillus cereus -245,54 32,23 -7,62 0,0001 18 Clostridium botulinum -127,68 3,45 -37,05 0,0001 13 Listeria monocytogenes -124,06 13,44 -9,23 0,0001 22 Staphylococcus aureus -203,96 14,61 -13,96 0,0001 31 Streptococcus pneumoniae -133,67 22,91 -5,83 0,0001 19 -82,33 43,49 -1,89 0,08 14 -87,4 11,58 -7,55 0,0001 16 0,27 2,30E-01 1,17 0,26 14 Escherichia coli Salmonella enterica Yersinia pestis Acinetobacter baumannii Streptococcus suis Streptococcus pyogenes Mycobacterium tuberculosis Corynebacterium diphtheriae Corynebacterium pseudotuberculosis Prochlorococcus marinus -25,97 48,16 -0,54 0,6 13 -335,35 13,55 -24,75 0,0001 15 -25,02 5,28 -4,74 0,0008 12 Table S5 - Annotations of SIGI-HMM predicted regions in Escherichia coli K-12 substrain MG1655 in Excel format NCBI name and chromosome position of SIGI-HMM predictions are found in the first column, with more detailed explanation of the predicted DNA in columns 2 and 3. All annotations are taken from the Islandviewer web-site. Table S6 - Annotations of SIGI-HMM predicted regions in Escherichia coli O26:H11 strain 11368 in Excel format NCBI name and chromosome position of SIGI-HMM predictions are found in the first column, with more detailed explanation of the predicted DNA in columns 2 and 3. All annotations are taken from the Islandviewer web-site.