Supporting Information S4.¾ Legend Phylogenetic analysis and
advertisement

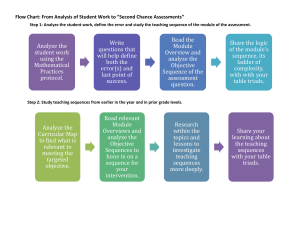
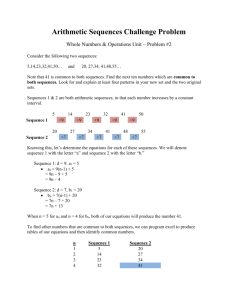
Supporting Information S4. Legend Phylogenetic analysis and divergence times inferred for cross-Andes and forestdependent xenarthrans. Trees were estimated from cytochrome oxidase I sequences of Bradypus (A) and Choloepus hoffmanni (B) using a Bayesian approach. Divergence estimates are indicated in the time scale and ranges representing 95% confidence interval for times estimates are represented as node bars. Numbers at the nodes indicate posterior probabilities. Terminals are identified according to subspecies (Table S2) and localities, indicated in the Neotropical region map containing the Andes extension and forested areas (Fig. 1, main text). For some C. hoffmanni (#) sequences the subspecies could not be determined because the exact geographical origin of the samples was unknown. For the C. h. hoffmanni sequence only the country of origin was known. Previously published DNA sequences are identified with *. For details on methodology, see Supporting Information S1, S2 and S3.