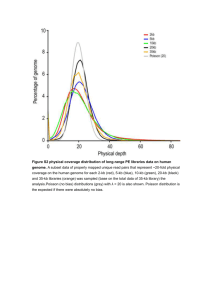
mec13327-sup-0001-supinfo
advertisement

Supporting Information for “Evolutionary history inferred from the de novo genome assembly of a non-model organism, the blue-eyed black lemur” Appendix S1. Sample information and DNA extraction for whole-genome sequenced samples For high-coverage sequencing, blood was obtained from one female blue-eyed black lemur (Harlow) and one female black lemur (Harmonia) during annual check-ups at the Duke Lemur Center. These individuals were chosen because of the potential availability of eye tissue for Harlow and the availability of other tissues for family members of Harmonia. For low-coverage population resequencing, blood or tissue was obtained from three blue-eyed black lemurs (one female: Lamour; and two males: Bogart and Monroe) and three black lemurs (one female: Blanche-Niege; and two males: Epimetheus and Zephyrus) during annual checkups or during necropsy. These individuals were chosen due to their limited relatedness to each other and to the individuals sequenced at high coverage (one low-coverage blue-eyed black lemur was a grandparent of the reference individual, but all other relationships were at least first cousins, once removed). We extracted DNA from all samples using a standard phenol-chloroform method. Appendix S2. Choice of sequencing libraries, library preparation, and sequencing for genome assemblies To generate a genome assembly with large contig and scaffold lengths (Salzberg et al. 2012), we sequenced libraries for several insert sizes, including paired-end (hereafter PE) libraries with mean target insert sizes of 180 bp, 500 bp and 1 kb, and mate-pair (MP) libraries with mean target insert sizes of 3 kb and 8 kb. The 180 bp library was chosen because the overlap of 100 bp reads from each direction can be used to create longer “superreads,” thus enabling the use of a larger value for k (Gnerre et al. 2011). We sequenced the PE libraries to high fold coverage and used the low-coverage sequencing reads from the MP libraries for scaffolding (Figure 2). PE libraries of target insert sizes 180 bp, 500 bp, and 1 kb were prepared using the recommended Illumina protocols at the University of Chicago and sequenced using an Illumina HiSeq2000 at Princeton University (Princeton, NJ). PE libraries with mean target insert size of 1 kb (blue-eyed black lemur) and 500 bp (black lemur) were prepared with Illumina’s TruSeq DNAseq Sample prep kit and MP libraries were prepared with Illumina’s Mate-pair version 2 Library prep kit at the Keck Sequencing Center of the University of Illinois (Champaign, IL); these libraries were also sequenced at the Keck Center. PE libraries were quantitated by qPCR and sequenced for 100 cycles from each end of the fragments on a HiSeq2000 machine using a TruSeq SBS sequencing kit version 3. Reads were analyzed with Casava 1.8 (pipeline 1.9) and subsequently stripped of adaptors. For low-coverage resequencing, PE libraries of target insert size 350 bp were generated using Illumina’s TruSeq DNA PCR-Free Sample Preparation Kit (cat no: FC-121-3001) and sequenced on a HiSeq2500 (2 lanes paired-end 100 bp in Rapid Run Mode) at the University of Chicago Genomics Core Facility (Chicago, IL). Appendix S3. Quality Control on Raw Reads Reads from each lane were screened to remove completely unusable reads (all bases read as ‘N’), and quality scores along the reads were visualized using the FASTX tool kit (http://hannonlab.cshl.edu/fastx_toolkit/). Because the ends of blue-eyed black lemur PE reads appeared to have highly variable quality scores, these reads were trimmed using the fastx_trimmer program to remove low quality suffixes prior to the stand-alone error correction programs (see S4). Black lemur reads were not trimmed using FASTX because their ends had higher quality scores and lower variance; however, any reads containing fewer than 70% bases with quality score > 20 were filtered out. MP reads were screened for redundancy by comparing the first 32 base pairs of reads, and only unique reads were retained. 24% of reads in the 3 kb library and 57% of reads in the 8 kb library were identified as redundant. Excluding them, usable read coverage from MP libraries dropped to 2X (Figure 2a). Appendix S4. Choice of pre-assembly correction method for PE reads Previous work suggests that the quality of the assembly is dependent upon both the quality of the data and the choice of assembler (Salzberg et al. 2012). Sequencing errors that are present in raw reads complicate the de Bruijn graph. Correcting these errors prior to assembly not only improves accuracy and contiguity but also simplifies the graph, which significantly lowers the memory overhead associated with the graph construction process (Schatz et al. 2010). In our own data, when assembling with raw reads only, the unitig (a unique contiguously assembled region ending at the boundaries of repeats) N50 was only 596 bp, and the corresponding scaffold N50 was approximately 8 kb. In order to increase these lengths, we used an external error correction program in addition to SOAPdenovo’s internal graph correction procedure. In order to assess how the choice of an error correction program affects assembly output, we compared the effect of two stand-alone assembly error correction programs; i) Quake (Kelley et al. 2010); and ii) a beta version of the then-in-development error correction program for SOAPdenovo, kindly provided to us by Ruibang Luo. The SOAP error correction package (version 0.04, compiled on March 6, 2012) is comparable to the method currently implemented in SOAPdenovo2 (Luo et al. 2012), with the exception that the version we implemented did not support long k-mers. Quake distinguishes between k-mers that are “trusted” to reflect genome sequence and those that are likely due to errors by evaluating the distribution of k-mer counts weighted by base quality values (“q-mers”). This process attempts to retain true, low-coverage k-mers while eliminating erroneous, high-coverage k-mers resulting from repetitive sequence (Kelley et al. 2010). To generate q-mer counts, we used the program Jellyfish (Marçais & Kingsford 2011), a fast and efficient k-mer or q-mer counter. The ‘jellyfish count’ program was run with ‘-m 19 –C –-quake ’ options, which indicates a q-mer size of 19, as recommended for the 3 Gb human genome (Kelley et al. 2010), and q-mers were counted on both strands. We used a hash size (-s) large enough to fit all q-mers in memory at once, since this minimizes the number of intermediate files written to disk. The counts output by the ‘jellyfish qdump’ command were used as input to the program ‘cov_model.py,’ a Quake accessory script that generates a histogram of q-mer counts. The program also calculates a coverage cutoff using, by default, the point at which the probability of the error distribution in a mixture model is 200 times the probability of the true distribution. To strike a balance between leaving many error k-mers uncorrected and over-correcting true k-mers, we chose a cutoff between that recommended by cov_model.py (1.25) and the local minimum of the q-mer distribution (5.25) for correction (c = 3; Figure S4b). q-mers were corrected with the ‘correct’ program using ‘–k 19 –c 3’ options. The SOAP error correction method makes use of the frequency spectra of both standard k-mers (“consecutive k-mers”) and k-mers incorporating a gap (“space k-mers”) to detect and correct error k-mers. It also uses overlapping PE information from short-insert libraries to avoid correcting true k-mers. For the SOAP error correction, we first ran the ‘k-merfreq’ program to count consecutive and space 17-mers (using k = 17, as recommended by Li et al. (2010), in the trimmed reads. We used the first local minimum in the resulting k-mer counts (c=14; Figure S4c) as the coverage cutoff, following Chaisson et al. (2009). The ‘ErrorCorrection correct’ program was then run in parallel with ‘-I –L -j -y’ options to account for overlapping reads from the 176bp insert library. For both error correction methods, reads that could not be corrected were discarded. The remaining reads were loaded to the assembly as PE if both mapped in the correct orientation to the same contig following error correction, or as single-end (SE) otherwise. Corrected read coverages were similar for Quake and SOAP (Figure 2a), and compared to Quake, SOAP maintained longer read lengths following correction (Table S1). Supporting Information Table S2 shows statistics for assemblies generated using SOAP-corrected reads and Quake-corrected reads (henceforth SCA and QCA, respectively). The contig N50 was comparable for both assemblies; however, QCA had a smaller proportion of chaff bases (bases within contigs less than 200 bp in length, which are discarded as unmapped reads), in addition to having larger contigs and scaffolds. We generated binary sequence alignment/map (BAM) files using the Burrows-Wheeler Aligner (BWA; Li & Durbin 2009) and SAMtools (Li et al. 2009) in order to assess the accuracy of read pair placement and to re-estimate the insert sizes for each library within the assembly. We indexed the scaffolds using ‘bwa index’, aligned all reads using ‘bwa aln’ command with default parameters, generated sequence alignment/map (SAM) files using ‘bwa sampe’ (for PE reads) and ‘bwa samse’ (for SE reads) commands, and converted the SAM files to BAM files using ‘samtools view.’ We filtered these alignments based on the mapping quality (MQ ≥ 20) and only included PE reads for which both reads aligned to the same scaffold. Filtered alignments were sorted using ‘samtools sort’ command, and PCR duplicates were removed using ‘samtools rmdup.’ For each target insert size, all BAM files corresponding to reads that were loaded as PE during the assembly were merged together, and the ‘TLEN’ fields were used to determine the insert size for each read pair. The merged BAM files were processed to retain only correctly oriented (-> <-) PE read pairs whose insert size was within three standard deviations of the mean (“happy pairs” according to Schatz et al. (2007)). The placement of PE reads in both assemblies appears to be highly accurate, with 98% of read pairs that map to the same scaffold identified as correctly oriented and happy (531,426,773 of 541,936,640 pairs for QCA, and 531,138,411 of 539,582,569 pairs for SCA). Only 0.024% of QCA mapped pairs and 0.027% of SCA mapped pairs mapped to different scaffolds. In choosing between QCA and SCA, our goal was to identify an assembly accurately representing the genome in as few contigs and scaffolds as possible, and in which the majority of read pairs were correctly oriented and happy. Although we did not detect any significant differences between the assemblies in proportion of happy pairs or in similarity to the black lemur bacterial artificial chromosome (BAC) sequences (see S9), our results indicated that QCA had more read pairs that were effectively merged into larger sequences. SCA had more than twice as many chaff contigs as QCA, in addition to a 71% increase in the number of unused reads, called singletons (Table S2). Such a high number of singletons could be due to assembly errors; for instance, when a tandem repeat has been incorrectly collapsed, the reads spanning the junction of the repeat cannot be mapped and become singletons (Treangen & Salzberg 2012). Singletons could also result from the presence of contaminant sequences, but such singletons should be present in both QCA and SCA. Chaff contigs, on the other hand, indicate the retention of low quality, repetitive sequences that are not able to be placed in scaffolds (Salzberg et al. 2012); such chaff contigs are more abundant in SCA than in QCA. These results suggest that more repeats may be unresolved in SCA than in QCA. We also observed a lower scaffold N50 and a lower percentage of scaffolds with no gaps in SCA than in QCA; this may be a consequence of using a higher depth threshold for classifying error k-mers, and thus eliminating a larger proportion of true k-mers. Alternatively, the use of a lower coverage cutoff in the QCA error correction may have led to the inclusion of some k-mers containing errors that would have been corrected in SCA, and to the assembler aggressively merging two erroneous sequences together (Salzberg & Yorke 2005). Based on the previous analyses and our preference for larger contigs and scaffolds, we chose QCA as our final assembly, and all analyses in the main text are done on this assembly. We also tried the strategy of retaining the error reads that could not be corrected by Quake or SOAPdenovo; however, this did not improve the contiguity of the assemblies. Instead, retention of error reads considerably increased the time to completion of each step of the assembly along with memory overhead, with peak memory exceeding 370 G during graph construction. Appendix S5. Choice of assembler At the time our assembly was generated, there were several genome assemblers capable of assembling mammalian-size datasets (Zerbino & Birney 2008; Miller et al. 2008; Simpson et al. 2009; Li et al. 2010; Gnerre et al. 2011; Simpson & Durbin 2012). In choosing among the available options, we prioritized ease of use, ability to run quickly and uninterrupted on a large dataset, ability to produce an assembly using primarily short insert library data, and performance in simulated datasets (Earl et al. 2011). Based on these criteria, we chose the de Bruijn graph (Idury & Waterman 1995; Pevzner et al. 2001) based short read assembler SOAPdenovo (Li et al. 2010). Appendix S6. Evaluation of insert size distributions and resolution of “bimodal” libraries The first step of scaffolding is to align the reads to the contig assembly, so that the paired end information can be used to join the contigs, and the size of gaps between contigs can be estimated (Li et al. 2010). The distribution of the true insert sizes for one library should in principle be normal, with mean μ approximately equal to the designed insert length for the library and variance σ2. However, the initial insert size estimates, inferred by comparing the migration of libraries within a gel to that of standardized DNA fragments, are often incorrect (Phillippy et al. 2008). Furthermore, the MP library construction protocol involves circularization and shearing of DNA fragments, thereby generating two populations of fragments: i) fragments that are truly separated by the MP distance and ii) contaminating short-insert fragments. If the library contains a large proportion of short-insert fragments, these reads provide little to no improvement in scaffolding. In order to assess the accuracy of the initial insert size estimates and to estimate the proportion of contaminating short-insert fragments within the MP libraries, we estimated PE and MP insert sizes by mapping pairs of reads to the assembly and determining the distance between them (see S4). We observed bimodal distributions for the 500 bp library and one of the two 1 kb libraries (Figure S6a and b). The assembly-based insert size distributions from a 1 kb library prepared at a different sequencing center did not appear bimodal (Figure S6c), which suggested that the additional peaks in the two distributions were artifacts caused by laboratory protocols, rather than assembly errors. Since SOAPdenovo assumes a normal distribution of insert sizes for each library, such bimodal distributions could be problematic for the assembler, leading to an incorrect estimation of library means, and thus potentially affecting the contiguity of contigs and scaffolds. Assuming that the bimodal distributions of insert sizes represented a mixture of two normal distributions, as visual inspection suggested, we estimated the two means and standard deviations using the R package ‘mixtools,’ which provides a set of functions based on expectation-maximization (EM) algorithms for analyzing finite mixture models (Benaglia et al. 2009). The normalmixEM() function was used to calculate the posterior probabilities of each insert size falling in each of two distributions, and the results were then used to assign read pairs to the distribution with the higher posterior probability for their mapped insert size. This resolved the distribution of insert sizes for the target 500 bp library into two component distributions, with mean insert sizes of 205 bp and 480 bp, and it resolved the 1 kb library distribution into distributions with mean insert sizes of 371 bp and 958 bp. These estimated insert sizes were then used to load the read pairs assigned to each distribution separately. This procedure resulted in a 25% increase in the contig N50 for QCA (16.3 kb) and a 28.9% increase for SCA (17.9 kb; Table S2), as well as a reduction in the percentage of chaff contig bases and singletons for both assemblies. Since MP libraries contain a combination of inward and outward facing reads, we used Stampy (Lunter & Goodson 2011) for MP alignments. Stampy trains insert size distributions separately for each orientation of read pairs and subsequently attempts the realignment of mates within each distribution. For re-estimation of insert sizes, we first mapped a small fraction of the data and used reads that fell within three standard deviations of the means as a training set for future mapping in Stampy. The resulting distributions were used as initial parameters, and the procedure was repeated until estimates approximately converged. We found approximately 7% and 17% of PE contamination in the 3 kb and 8 kb libraries, respectively. These PE reads (with mean insert sizes of 420 bp) were separated from the MP reads and loaded along with the other PE libraries for the assembly. From the reads determined to map as MP, we used the outward facing reads that mapped within three standard deviations of the estimated mean insert size for the final assembly. Figure 2b shows the increase in scaffold N50 values for QCA upon sequential addition of different PE and MP libraries. Appendix S7. Command line parameters for assembly generation All de novo assemblies were done with a k-mer size of 33 because we obtained the best N50 statistics with this k-mer size. We used the SOAPdenovo63mer executable for all computations. The first step of the de novo assembly is to load all the reads into a de Bruijn graph, which was constructed using the following command: ./SOAPdenovo63mer pregraph -s config_file -p 12 -K 33 -o Quake_K33 Contigs were constructed by merging similar sequences and resolving tiny repeats from read paths using the following command: ./SOAPdenovo63mer contig -g Quake_K33 -M 3 –R Reads were then mapped back to the contigs to transfer the paired-end information from the reads, using the following command: ./SOAPdenovo63mer map -s config_file -g Quake_K33 -p 12 Finally, scaffolds were constructed with an option to fill small gaps using the following command: ./SOAPdenovo63mer scaff -g Quake_K33 -F -p 12 At this point, we obtained scaffolds with masked repeats appearing as gaps in the assembled sequence. In order to disambiguate the repeats and fill in the gaps, we ran the ‘GapClosure’ module on the scaffolds. This program aligns the PE reads to the scaffolds and gathers the pairs in which one end has aligned and the other end falls in a gap region. A local assembly of the gap region is then done. The following command was used for gap closure: ./GapClosure –a Quake_K33.scafSeq –b config_file –o Quake_K33.gapClosed.scafSeq –p 31 –t 12 This procedure closed 79% of gaps. For the black lemur assembly, we used the recommended pipeline within BWA version 0.5.9 (Li & Durbin 2009). We first aligned reads separately for each end of the PE library and for the SE reads whose pair failed quality filters using the following command: ./bwa -aln -n 0.04 -o 1 -q 15 We then converted the alignments of the PE reads and the SE reads to separate SAM files using the following command: ./bwa sampe -a 1000 -n 1 and ./bwa samse -n 1 We filtered out reads with mapping quality less than 10 and converted to a BAM file in SAMtools version 0.1.17 using the following command: ./samtools view -S -h -q 10 -b To build a consensus genome sequence for the black lemur, we used the Genome Analysis Toolkit (GATK; McKenna et al. 2010; DePristo et al. 2011) to call polymorphic sites from the resulting BAM file and to produce an all-sites Variant Call Format (VCF) file including genotypes at all sites (see S11). We then used a custom python script to retain calls at all sites with at least three reads confidently mapped; at sites identified as polymorphic, the more common allele among the reads was selected as the consensus. Note that the minimum coverage for reporting a base pair within the consensus sequence is lower than that used for calling high confidence sites for the pairwise sequential Markovian coalescent (PSMC; Li & Durbin 2011) and other population genetic analyses. Custom scripts or modifications of available scripts used for processing these data are available at https://github.com/sorrywm/genome_analysis. Appendix S8. Details of memory usage and run times The peak memory consumption and run times for different stages of the assembly pipeline are shown in Figure S2. Note that this is for the error-corrected reads, whereas assembling the raw reads required a peak memory of 379 GB during the graph construction stage. This difference between memory usage for raw and corrected reads illustrates how, although in theory the size of the de Bruijn graph should depend primarily on the size of the reference genome, in practice sequencing errors create their own nodes in the graph and thus require substantial memory resources. Peak memory consumption was during the final step of the assembly, the Gap (gap closure) stage, whereas the maximum run time was for error correction. We performed all memoryintensive computations on two computers: one with 16 quadcore Intel Xeon 2.4 Ghz CPUs with 288 Gb memory installed and the other with 48, 12-core AMD Opteron Processors with 512 Gb memory installed. Using these machines, both QCA and SCA required approximately 5 days from the start of k-mer counting to the completion of the gap closure step. Similar figures were observed for each assembly step in SCA, and the error correction for SCA required 21 GB of RAM. Appendix S9. Aligning blue-eyed black lemur contigs to black lemur bacterial artificial chromosomes (BACs) We aligned the contigs of the blue-eyed black lemur genome assembly to the eight genomic regions (4.86 Mb total sequence) of the black lemur that have been sequenced using a BAC-byBAC shotgun sequencing strategy (NISC Comparative Sequencing Initiative; NCBI trace archive library ID CHORI-273; GenBank accession numbers DP000024.1, DP0011503.3, DP000556.1, DP000490.3, DP000520.1, DP000468.3, DP000905.1, and DP001278.1). Contig alignments were created as described in Salzberg et al. (2012). Briefly, we used the ‘dnadiff’ wrapper script, which is a part of the MUMmer package (Kurtz et al. 2004), to create the alignments. We used the BAC sequence as the reference and blue-eyed black lemur contigs (excluding chaff contigs) as the query, and we aligned these using the ‘nucmer’ program. Use of the shorter sequence as the reference minimized the memory requirement for alignment. Using the supermap algorithm, optimal alignments were created, while retaining large scale re-arrangements (Dubchak et al. 2009). Alignments with <95% identity or >95% overlap with another alignment were discarded using the ‘delta-filter’ program. The resulting high-quality one-to-one alignment mappings were used to calculate divergence by parsing the one-to-one alignment blocks, considering substitutions where ≥20 bp matched exactly on either side of the alignment. Appendix S10. Core Eukaryotic Gene Mapping Approach (CEGMA) analysis We ran the CEGMA pipeline (Parra et al. 2007, 2009) against our draft scaffolds to estimate the proportion of conserved genes represented in our assembly. The set of 248 highly conserved core eukaryotic genes (CEGs) have been divided into four groups based on the degree of conservation. Group 1 contains the least conserved proteins, while Group 4 contains the most conserved proteins. ‘Complete’ refers to a predicted protein that yields an alignment to the profile hidden Markov model (HMM) for that protein that is at least 70% of the profile HMM length and exceeds a pre-computed minimum alignment score, obtained by calculating the maximum alignment score for all non-CEG genes. Matches that exceed this threshold are generally fulllength proteins (Parra et al. 2009). ‘Partial’ refers to proteins with alignments that are shorter but still exceed the minimum alignment score. These partial matches may correspond to shorter fragments of proteins, such as domains. While we are able to map 94.7% of the proteins as partial matches, we only map 59.2% of the proteins as complete. One explanation could be that even though the N50 of our scaffolds is high, 421 kb, contigs within scaffolds are separated by gaps. If the average gene length for CEGs in Eulemur flavifrons is 24.6 kb (the mean length for the most highly conserved CEGs in humans), some genes may extend into gaps within the assembly because only 12.5% of contigs exceed 70% of this length. In addition, some genes may overlap repetitive regions of the genome that have been excluded as singleton contigs. Appendix S11. SNP calling We called SNPs from the blue-eyed black lemur BAM files generated to assess PE correctness and insert size distribution (see S4), renaming the merged BAM files to assign all data to the same sample using SAMtools ‘reheader,’ and the black lemur BAM file used for the referencebased assembly (see Methods and S7). To call SNPs, we used the Genome Analysis Toolkit (GATK; McKenna et al. 2010; DePristo et al. 2011) version 2.2-16. PCR duplicates had already been removed from each library of the blue-eyed black lemur data separately (see S4). For the black lemur, we marked duplicates using the MarkDuplicates tool within Picard (http://picard.sourceforge.net/) version 1.81, with the following arguments: ASSUME_SORTED=TRUE VALIDATION_STRINGENCY=LENIENT MAX_FILE_HANDLES_FOR_READ_ENDS_MAP=1000 Because this duplicate marking process disrupts the read group information in the BAM file, we then replaced the read groups using Picard’s AddOrReplaceReadGroups, with the following arguments: SORT_ORDER=coordinate RGLB=Em500PE RGPL=illumina RGPU=unknown RGSM=EmHarmonia CREATE_INDEX=True VALIDATION_STRINGENCY=LENIENT We realigned reads locally around indels using GATK’s RealignerTargetCreator and IndelRealigner. Finally, we called SNPs using UnifiedGenotyper, with the following arguments: -out_mode EMIT_ALL_SITES -mbq 20 -rbs 50000000 The ‘EMIT_ALL_SITES’ option was used to make calls for both monomorphic and polymorphic sites, and only those bases that had a base quality of 20 or greater were initially called. The raw SNP calls contain many false positives due to sequencing artifacts, mismapped reads, and indels, as well as potential false negatives. We therefore performed filtering to eliminate potential false positive SNPs and to mask monomorphic sites at which a SNP could not have been confidently called using several recommended criteria (Abecasis et al. 2010; Auton et al. 2012), excluding: i) sites with base quality < 30 ii) sites with read depth < half the mean read depth, or > twice the mean depth iii) sites with MQ (root mean square of the mapping quality of all reads spanning a site) < 20 When calculating diversity within and divergence between species, we directly used the QUAL value (the Phred-scaled probability estimated by GATK of a type I or type II error at a site) provided in the VCF for the base quality cutoff. For the conversion of the VCF to the fastq files used for PSMC, we further weighted the QUAL value by the GQ (the Phred-scaled probability of an error in the called genotype) in cases in which the called genotype was not homozygous reference. We additionally modified the ‘vcf2fq’ function in the vcfutils.pl script provided by SAMtools in order to enable genotype calls at sites where both alleles were non-reference and to mask sites with insufficient data, rather than calling them homozygous reference (see ‘vcf2fqnonref’ in https://github.com/sorrywm/genome_analysis/vcfutils_mod.pl). When estimating diversity and divergence, we used a more stringent MQ filter, excluding sites with MQ < 30. Appendix S12. Sanger-based resequencing of additional samples within the scaffold containing the OCA2 ortholog We generated Sanger sequence data for subsections of the scaffold containing the OCA2 ortholog in a larger sample of individuals. To this end, we used DNA samples for five blue-eyed black lemurs and five black lemurs from a previous study of ours (Meyer et al. 2013). DNA samples from three additional blue-eyed black lemurs and one additional black lemur were obtained from liver samples provided by the Duke Lemur Center, Durham, NC, using the DNeasy Blood and Tissue Kit (Qiagen). We designed primers to amplify regions with a target size of 1.2 kb and a Tm of 60ºC, using a custom script employing Primer3 (https://github.com/sorrywm/genome_analysis/DesignOCA2Primers.py). We filtered the resulting primers to obtain one primer pair per 30 kb of sequence, with the properties that the amplicons captured the maximum number of “fixed differences” between reference genomes and that there were no polymorphisms in the primer sequence based on the four chromosomes already sequenced. Primers yielding single amplicons for all samples, along with additional internal sequencing primers, are listed in Table S3. We amplified 20 ng of DNA in a total volume of 20 µL with 1X PCR buffer, 1.5 mM MgCl2, 500 µM, 100 nM each primer, and 0.5 units Platinum® High Fidelity Taq (Invitrogen); to improve specificity, 1 µL each Betaine and DMSO were added to some reactions. The PCR conditions were as follows: 94ºC for 2 min, 30 cycles of 94ºC for 30 s, 55ºC for 30 s, and 68ºC for 75 s, 68ºC for 10 min. PCR products were purified using Exo-SAP. For sequencing, 1.2 μL of purified PCR product and 1.2 μL of 4 μM primer were added to 2 μL BigDye® v3.1 (Applied Biosystems) and 2 μL water, and this reaction mix was cycled 50 times for 5 sec at 96ºC, 10 sec at 50ºC, and 3 min at 60ºC. Water was added to approximately 25 μL, and reactions were purified by Sephadex® G50 Superfine (GE Healthcare Bio-Sciences AB) in a 96-well filter plate (Pall). These were directly placed on the Applied Biosystems 3730XL or 3130 capillary sequencer and run as a standard 2 hour run using POP-7™ polymer (Applied Biosystems). Resulting sequences were determined from trace files using a python script utilizing the abifpy module (https://github.com/bow/abifpy/), and any bases with quality score < 20 were checked by eye from the chromatogram. In total, we obtained sequence data for regions totaling 9.9 kb in length from the blue-eyed black lemur and 15.1 kb from the black lemur. Within these regions, we observed 88.6% concordance of Sanger calls with GATK-derived polymorphisms (out of 35 heterozygous sites called by GATK, there were four discordant sites, all of which occur within the same amplicon) and only three sites where GATK failed to identify a SNP present in Sanger sequencing. We visually inspected the chromatogram for the amplicon containing the four sites that had been called heterozygous in the whole genome sequencing but homozygous in Sanger sequencing, and in three of these cases, the Sanger call appeared to be in error. Assuming no additional errors, these results suggest that the probability of incorrectly calling a homozygous site heterozygous is approximately 0.00012, and the probability of incorrectly calling a heterozygous site homozygous is approximately 0.029. We therefore concluded that the initial GATK calls were of high quality, and we used these data without any further recalibration in subsequent analyses. In order to determine haplotypes and estimate FST, sequences obtained for each region were phased using SHAPEIT2 (Delaneau et al. 2013). All polymorphic sites identified from sequencing were combined into one map file for phasing, and a constant recombination rate of 1 cM/Mb was assumed for determining genetic distance between markers. Effective population size was estimated from pairwise differences as follows: Ne = ((π Ef + 2* π Ef_Em + π Em)/4)/4μ, where π Ef and π Em represent genome-wide within-species diversity for blue-eyed black lemur and black lemur, respectively, and π Ef_Em represents genome-wide divergence. SHAPEIT2 was run using the following parameters: Shapeit --input-bed AllAmpliconsMerged_bfile.bed AllAmpliconsMerged_bfile.bim AllAmpliconsMerged_bfile.fam -M AllAmpliconsMerged.gmap --exclude-ind UnsequencedLemurs.ind -W 0.5 --effective-size 35000 Appendix S13. Simulations of pairwise sequential Markovian coalescent (PSMC) performance on scaffold data and with a population split The PSMC method (Li & Durbin 2011) was developed for use on data from linear genomes and initially tested on whole human genome sequences. In contrast, our assembly contains much more missing data, and the sequences are present in several thousand scaffolds rather than a small number of chromosomes. In order to determine how these features could influence the estimates of PSMC, we simulated data with similar properties to those of our draft genome assembly. Specifically, for a given demographic history (similar to the demographic history of the blue-eyed black lemur, estimated by PSMC), we simulated a sequence of length 2.5 x 109 bp (the approximate length of the blue-eyed black lemur genome), using a modification of msHOT (Hudson 2002; Hellenthal & Stephens 2007) developed by Heng Li and available at https://github.com/lh3/foreign/tree/master/msHOT-lite. Using a python script, we further simulated 100 assembly-like sequences by selecting random segments from the whole data, with lengths drawn at random from the blue-eyed black lemur scaffold lengths and adding up to 2.0 x 109 bp (the approximate length of the assembly used for PSMC). In addition, we replaced blocks of these segments with missing values (N’s), where the lengths and distribution of these blocks were determined using the distribution of missing data from the blue-eyed black lemur assembly. The resulting plot of trajectories obtained by applying PSMC on simulated assemblies suggests that using scaffolds rather than the full data increases the variance of the inferred demographic trajectory, as might be expected, but does not lead to any marked biases (Figure S3b). Importantly, this increase in variance is negligible in the parameter range in which PSMC is expected to produce reliable estimates (i.e., excluding very recent or ancient times). The ms input for the simulated demographic history (Figure S3b) was: msHOT-lite 2 100 -t 50440 -r 6710 25000000 -l -eN 0.011 0.380 -eN 0.042 0.987 -eN 0.111 0.638 -eN 0.301 1.171 -eN 0.694 0.925 -eN 1.910 2.725 -eN 4.158 2.323 In addition, we observed that the time at which the PSMC-inferred trajectories for the blue-eyed black lemur and black lemur began to overlap was more ancient than the split time inferred from divergence and shared polymorphisms (see S14). To investigate whether this discrepancy might be due in part to PSMC smoothing of abrupt changes in the effective population size, we ran the method on simulated data generated with a sudden population split. We simulated demographies imitating the demographic histories of the two species initially inferred by PSMC, with a split time based on our estimates using diversity levels (Figure S3c). We further generated bootstrap resampling data (100 bootstraps of the total genome length using 500 kb intervals) using PSMC and obtained a distribution for the estimated time at which the two trajectories split based on the PSMC output (Figure S3c). As we suspected, the estimated split time tended to be greater than the simulated split time, consistent with the discrepancy between the PSMC output and the split time estimated from diversity and divergence data (Figure S3d; see also supporting information for Prado-Martinez et al. (2013) and Freedman et al. (2014)). The ms command for this simulation was: msHOT-lite 4 100 -t 14647.1163653362 -r 2084.10374328881 5000000 -I 2 2 2 0 -en 0.0334 1 1 en 0.06 1 3 -en 0.38 1 5 -ej 1.095 2 1 -en 1.095 1 3.5 -en 1.6335 1 3.5 -en 5 1 10 -en 10 1 5.8929 -en 0.0241 2 1.0000 -en 0.0958 2 3 -en 0.33 2 2 -en 1.095 2 3.5 Appendix S14. Estimating species split time The probability P(D) that the allele at a site differs between one chromosome from each of the species is given by the average divergence between lineages and the mutation rate. Under a simple isolation model and Wright-Fisher assumptions, the average time to the most recent common ancestor (TMRCA) of two lineages, one sampled in each of two species, is the time to the species split (t) plus the average time to coalescence in the ancestral population (2N a). Thus, P(D) = 2μ(t + 2Na), where μ is the mutation rate. Under the same assumptions, if the effective population size of both descendant species is Ne, then the probability P(S) of a polymorphism with both alleles shared identical by descent (IBD) in a sample of two diploid individuals (one per species) is: P(S) = (e-t/2Ne)2 x 2/3 x (2Na) x μ This equation represents the product of the mutation rate and three terms: 1) The probability of no coalescences occurring in either species prior to time t, which is (e -t/2Ne)2. 2) The probability of the four lineages in the ancestral population coalescing in an order that could lead to a shared polymorphism. Namely, the first coalescence must occur between lineages from different species, an event that has probability 2/3. 3) The total length of lineages on which a mutation leading to a shared polymorphism could occur. Once the first coalescent event has occurred (between lineages from different species), there are three possible orders for the second coalescent event. In one of these, the second coalescence does not involve the lineage resulting from the first coalescent event, and the expected branch length on which a mutation leading to a shared polymorphism could occur is two times the expected TMRCA for three lineages minus the expected time to the first coalescence in a sample of three, or 2 x (8Na/3) - 2Na/3 = 14Na/3. The other possibility is a coalescence between the branch resulting from the first coalescence and one of the other two lineages; in this case, the expected branch length on which a mutation leading to a shared polymorphism could occur is the time to the first coalescence in a sample of three, or 2Na/3. Thus, summing over all three cases, the expected branch length is 1/3 x (14N a/3) + 2/3 x (2Na/3) = 2Na. The product of these terms is the probability of a polymorphism shared IBD. Alternatively, we can write these equations in terms of τ, the split time scaled by 2Ne, and Na/Ne, the ratio of ancestral effective population size (Na) to current Ne, so the probability of a fixed difference between the two genomes is: P(D)=2 x (2Neτ + 2Na/Ne x Ne) x μ, and P(S)=(e-τ)2 x 2/3 x (2Na/Ne x Ne) x μ. These relationships can be written in terms of mean pairwise diversity (π = 4N eμ) as: P(D)=π x (τ + Na/Ne) P(S) = (e-τ)2 x 1/3 x (Na/Ne x π) We used these equations to estimate the split time and Na by a moment estimator. Specifically, we took the harmonic mean of the pairwise differences within each species to estimate π, and we solved for τ and Na/Ne by substituting the observed genome-wide divergence between species for P(D), and the observed fraction of shared polymorphic sites for P(S). We solved the system of equations using the uniroot function in R version 2.15.2 (Brent 1973). Appendix S15. Choice of parameters for PSMC and scaling of PSMC output and species split time Within the PSMC program, heterozygosity is summarized across a “window,” with a default size of 100 bp. Based on the estimated SNP density within the two lemur genomes, we chose a window size of 75 bp, in order to reduce the number of windows containing 2 or more SNPs to approximately 1%, and thus strike a balance between information loss and computational complexity. However, we additionally ran PSMC on the blue-eyed black lemur data with window sizes ranging from 20 to 500 bp, in order to assess what effect this choice might have on our conclusions. With the exception of 500 bp, the trajectory of ancestral population size appears highly similar across window sizes (Figure S3a). In order to translate the output of PSMC and the estimate of τ (see S14), both of which are given in terms of 2Ne (two times the current effective population size) generations, to a more interpretable temporal scale, we needed an estimate of the mutation rate. The genome-wide pergeneration mutation rate has not been directly estimated for lemurs; however, pedigree-based estimates are available for human data (1.2 x 10-8 per bp per generation; Kong et al. 2012), and estimates based on experimental mutation accumulation are available for mouse (Mus musculus: 3.8 x 10-8; Lynch 2010). Within primates, mutation rate has been shown to be inversely correlated with generation time, which may explain the “hominoid slowdown” in molecular divergence rates (Kim et al. 2006; Tsantes & Steiper 2009). This correlation may be due to DNA replication errorgenerated mutations, which accumulate over cell divisions; in contrast, mutations of CpG to T, which do not result from replication error, tend to behave in a clocklike manner across mammals (Hwang & Green 2004). This clocklike behavior suggests that the proportion of all mutations on a given branch that are CpG to T should negatively correlate with the per-generation mutation rate. Because the generation time of the blue-eyed black lemur is shorter than that of human (see below) and the proportion of CpG to T mutations is lower in the lemur than in the human branch (Hwang & Green 2004), we used a mutation rate estimate higher than that reported for humans (but lower than that for mouse), or 2.0 x 10-8 per bp per generation. The generation times of the blue-eyed black lemur and black lemur are also unknown; however, based on the reproductive ages of individuals in captivity at the Duke Lemur Center (1.5 - 25 years) and on the mean age of first reproduction of blue-eyed black lemurs in the wild (3 years; Volampeno et al. 2011), and assuming some reproductive senescence, we assumed the generation time for both species to be 5 years. The effects of using different point estimates within biologically plausible ranges (0.5 – 5.0 x 10-8 for mutation rate per generation and 2.5 - 10 years for generation time) are shown in Figure S3e-f. Because the black lemur was sequenced to lower coverage than the blue-eyed black lemur, we assumed that the ability to detect polymorphisms in this dataset might be reduced, leading to an under-estimate of overall diversity. To estimate the influence of coverage on SNP-calling ability, we called SNPs using a subset of the blue-eyed black lemur data containing approximately the same number of mapped reads as for the black lemur dataset and compared the results to those obtained using the full dataset. The number of “callable” bases for the thinned blue-eyed black lemur data turned out to be lower than that for the black lemur data, presumably due to differences in library preparation and sequencing between centers. We therefore estimated the effect of the reduced coverage by assuming a linear relationship between the number of callable bases and the proportion of true SNPs called as SNPs. We incorporated this effect into the PSMC output by reducing the mutation rate for the black lemur to 1.951 x 10-8 per generation per bp. The plots for the full blue-eyed black lemur dataset and the subset with adjusted mutation rate show strong concordance, indicating that this correction factor appears to adequately capture the effect of lower coverage (Figure S3g). We note that, because the two individuals we sequenced were female, some of the information may be derived from the X chromosome, which has Ne ¾ that of an autosome. This may lead to an underestimate of Ne by PSMC. Because karyotypic and chromosome painting work suggests that the X chromosomes of humans and black lemurs are homologous (Müller et al. 1997), the X should represent approximately 5.8% of the black lemur genome length, and thus the effect on N e inference should be small. Appendix S16. Identification of candidate regions for recent positive selection in one species Recent positive selection may be detectable as a reduction in diversity relative to divergence in one of the species. In the absence of a genetic map for these species, we initially assessed diversity and divergence on a physical scale of 100 kb, which is roughly the span over which a recent, strong selective sweep would impact genetic diversity at linked sites (assuming a selection coefficient of 1% and an average recombination rate similar to that of humans; Kaplan et al. 1989; Kong et al. 2004). In order to identify regions that may have been subject to weaker selection, as well as to assess the genome-wide empirical significance of individual regions, we further considered non-overlapping windows of 20 kb. We calculated FST (1 – π w/ π b), but this statistic is extremely noisy when using a sample size of two chromosomes from each population. We also considered the following summary statistic: Ps1 = (hs1 + ss) / (hs1 + hs2 + ss + fd), where hs1 represents the number of sites heterozygous only within the focal species, hs2 the number of sites heterozygous only within the other species, ss the number of shared heterozygous sites, and fd the number of sites at which the two species are homozygous for different alleles. We expect recent selection to result in a decrease in hs1 and ss, and an increase in fd, but not affect hs2, so we are particularly interested in regions for which this statistic is unusually small. In order to delineate the boundaries of strongly selected regions for the annotation of genes based on the initial two-sample dataset, we calculated Ps1 in each window of 100 kb, sliding by 10 kb. Within all such windows with at least 80 kb of callable sites in both species, 3.9% had Ps1 = 0 for the blue-eyed black lemur (0.5% for the black lemur). We obtained an initial set of candidate regions to annotate for each species by identifying all windows that fell in the 3.9% tail of Ps1 for 100 kb sliding windows in that species and then combining any overlapping windows into larger regions. To determine the empirical percentile of individual regions described in the main text, as well as to identify regions subject to weaker selection in the larger dataset, we also calculated Ps1 and FST for 20 kb non-overlapping windows (Figure S7). We selected 20 kb windows in the 1% FST tail from the larger dataset, combining any adjacent windows, to generate a list of regions from the larger dataset to annotate for gene ontology analysis (see S21). Appendix S17. Annotation of orthologs of OCA2 and additional human iris pigmentation candidate genes within the blue-eyed black lemur genome We downloaded the sequence of exons from the longest human transcript of OCA2 from Ensembl GRCh37. We used BLASTN 2.2.26+ with default parameters to compare each human exon to the reference genome assembly of the blue-eyed black lemur. From the six exons that successfully mapped in this initial search, we determined that the coding region was likely to be within scaffold 2503. We downloaded the RefSeq-annotated human OCA2 sequence, including untranslated regions and all introns from the UCSC genome browser and used BLASTN to compare it to scaffold 2503. We used the overlap of the BLASTN hits with the positions of human exons from the RefSeq gene annotation to identify the positions of the remaining lemur exons within the scaffold. Part of the first exon, which lacks strong conservation in mammals, could not be annotated in this way. For this exon, we determined the first base pair of the sequence by finding the start codon that would maintain the same translation frame nearest to the predicted start position based on the human exon length. We obtained sequence for all exons for both species from the SNP-containing references, and we translated the resulting open reading frames using eBioX version 1.5.1 (http://www.ebioinformatics.org/). We found a single difference between the protein sequences of the two species, namely, that position 89 was L in blue-eyed black lemur, whereas in black lemur, it was heterozygous for L and P. Because this site is segregating within black lemurs, it cannot be solely responsible for the lemurs’ fixed iris pigmentation difference, yet it is possible that it could contribute to the phenotype. To determine whether this site might play a role in iris pigmentation, we aligned the inferred protein coding sequences of the two lemurs against coding sequences for nine other mammals (chimpanzee, dog, horse, human, macaque, marmoset, mouse, mouse lemur, and orangutan), downloaded from Ensembl. We observed that this site was not highly conserved, and notably both L and P alleles occurred in other brown-eyed mammals, and we thus inferred that this locus is likely not involved in iris color determination in lemurs. In addition to OCA2, a number of other genes have been much more weakly associated with natural iris pigmentation variation in humans or associated with disease or mutant phenotypes resulting in blue irises in humans or mice (Kanetsky et al. 2002; Loftus et al. 2002; Frudakis et al. 2003; Sulem et al. 2007; Sulem 2008; Sturm et al. 2008; Liu et al. 2009, 2010; Valenzuela et al. 2010; Pingault et al. 2010; Hellström et al. 2011). We identified the location of orthologs of 16 such pigmentation candidate genes (ASIP, CYP1A2, DCT, DSCR9, HGS, MITF, MYO5A, NPLOC4, PAX3, PMEL, RAB38, SLC24A4, SLC24A5, SLC45A2, TYR, and TYRP1) within the blue-eyed black lemur genome using BLAST, and we looked for evidence of selection within these regions. Specifically, we downloaded one DNA sequence for each gene from Ensembl GRCh37 and used BLASTN version 2.2.27+ (Altschul et al. 1990, 1997) with default parameters to identify the best match between each exon and the lemur reference genome. We examined the regions containing multiple exons for signatures of selection in the blue-eyed black lemur. In our initial, two-sample analysis, two of three 20 kb windows overlapping DCT and one of seven windows overlapping ASIP had Ps1 = 0. In the analysis of the larger dataset, the signals of selection at DCT and ASIP were somewhat reduced (DCT minimum Ps1 = 0.20, 9.1%-tile; maximum FST = 0.80, 18.7%-tile; ASIP minimum Ps1 = 0.17, 6.6%-tile; maximum FST = 0.88, 6.4%-tile). However, several other candidate genes overlapped windows in the tail of these test statistics in the larger dataset: MITF and TYR, which play known roles in the melanin biosynthesis pathway and have been robustly associated with human pigmentation variation (Sturm et al. 2008; Visser et al. 2012), and LYST and NPLOC4, which overlap regions associated with quantitative pigmentation variation in one recent study (Liu et al. 2010). When we corrected for the number of windows tested for each gene by comparing FST and Ps1 statistics in these windows to those in genome-wide regions matched for length (see Methods), only the signal at MITF remained unusual (corrected p-values: ASIP FST p = 0.269, Ps1 p = 0.287; DCT FST p = 0.383, Ps1 p = 0.212, LYST FST p = 0.155, Ps1 p = 0.177; MITF FST p = 0.029, Ps1 p = 0.014; NPLOC4 FST p = 0.061, Ps1 p = 0.107; TYR FST p = 0.351, Ps1 p = 0.132). We determined the amino acid sequences for three of these genes that showed potential signatures of selection in the larger dataset and whose function in melanin biosynthesis is wellestablished: ASIP, MITF, and TYR. We downloaded the sequence of exons from the two known transcripts of ASIP, five known transcripts of MITF, and one known transcript of TYR from Ensembl GRCh37. As in the annotation of OCA2, we used a BLASTN search with default parameters to find the scaffold or scaffolds most likely to contain the gene. We then extracted the sequence of those scaffolds and performed a discontiguous megablast of each human exon not found in the initial search to the scaffolds using the online BLASTN tool (http://www.ncbi.nlm.nih.gov/blast). In this way, we were able to identify the positions of all exons for known transcripts of these genes. We obtained sequence for all exons for both species from the SNP-containing references, and we translated the resulting open reading frames using eBioX version 1.5.1 (http://www.ebioinformatics.org/). Appendix S18. Identification of candidate regulatory changes within the scaffold containing the OCA2 ortholog We found no fixed amino acid differences between the blue-eyed black lemur and black lemur OCA2 sequences obtained by translating the coding sequence annotated from the two genomes (see S17). We therefore focused our search for candidate loci influencing iris pigmentation on regions with potential regulatory function. In humans, the causal variant disrupts a HLTF binding site; the blue-eyed allele is also associated with reduced binding of the transcription factors LEF1 and MITF at nearby motifs in comparison to binding at the brown-eyed allele, even though there are no sequence changes in these motifs (Eiberg et al. 2008; Visser et al. 2012). These findings suggest that all three factors (HLTF, LEF1, and MITF) may jointly regulate OCA2 expression in humans, and thus mutations that disrupt binding of one of these factors in blue-eyed black lemurs provide strong candidates for a causal mutation. We therefore searched for sequences predicted to bind HLTF, LEF1, or MITF strongly in the black lemur and weakly in the blue-eyed black lemur. Because in human all three of these factors bind near the causal site, we focused on inferred differential binding sites in lemur where another transcription factor also had a motif nearby. Previous research has demonstrated strong conservation of binding site preferences for some transcription factors across vertebrates (Schmidt et al. 2010). Assuming that binding site preferences for HLTF, LEF1, and MITF have been conserved between lemurs and humans, we used human-derived motifs to search for potential binding site changes in lemurs. We downloaded position weight matrices (PWMs) for RUSH (the mammalian ortholog of HLTF), LEF1, and E-box (the type of binding site recognized by MITF) from TRANSFAC (Wingender et al. 1996) or JASPAR (Sandelin et al. 2004). We used the PWMs to calculate "PWM scores," which measure how strongly observed sequences match a motif. Specifically, a PWM score is the likelihood of binding of a transcription factor given the observed sequence. Conditional on transcription factor binding at one allele, the difference in PWM scores between alleles has been shown to be a good predictor of a change in transcription factor binding (McVicker et al. 2013). We calculated PWM scores for each factor at each site within the scaffold 2503 sequence for black lemur and blue-eyed black lemur. For each factor, we identified sites where the black lemur and blue-eyed black lemur samples were fixed for different alleles, where the black lemur allele resulted in a PWM score within the top 10% of scores within the scaffold, and where the blueeyed black lemur allele resulted in a 10-fold lower predicted binding (a log-10 reduction in PWM score). For these sites, we determined which allele was derived using one of two outgroups: aye-aye (data downloaded from http://giladlab.uchicago.edu/data/AyeAyeGenome/) or mouse lemur (data downloaded from ftp://ftp.ensembl.org/pub/release-74/fasta/microcebus_murinus/dna). Specifically, we used BLASTN (Altschul et al. 1990, 1997) to identify the ortholog of the 100 bp surrounding each allele in aye-aye and/or mouse lemur, and we determined whether each sequence at the divergent site resulted in a match with the outgroup(s) in the top BLAST hit. Our candidates are cases in which the black lemur allele at the divergent site matched the outgroup allele and the blue-eyed black lemur allele did not (i.e., providing evidence that the black lemur allele was ancestral and the blue-eyed black lemur allele was derived). We required that the sequence surrounding both alleles have BLAST hits for at least one of the two outgroups, and that neither outgroup be discordant (discordant cases were ones in which the black lemur allele was inferred to be derived or in which both alleles had perfect matches in the outgroup). We then identified the subset of these candidates residing within 200 bp of a site within the top 10% of PWM scores for at least one of the other transcription factors (Figure 5b). The scripts used to perform these searches are available at https://github.com/sorrywm/genome_analysis/regulatory_annotation. Appendix S19. Calculation of summary statistics from the combined sample using ANGSD and ngsTools Following raw read filtering, reads were aligned separately for each sample and lane using bwa and samtools, with the same alignment procedure as for the high coverage black lemur data (SI Text S7). Duplicates were marked with Picard, and reads were re-aligned locally around indels using GATK’s RealignerTargetCreator and IndelRealigner. The resulting alignment (.bam) files were used as input to ANGSD (http://popgen.dk/wiki/index.php/ANGSD). Initial site frequency spectrum likelihoods were estimated per site based on individual genotype likelihoods, assuming Hardy-Weinberg equilibrium (-doSaf 1), with a GATK model for genotype likeihoods (-GL 2). For this step, sites were filtered to require a minimum mapping quality of 1, minimum quality of 20, and a minimum of three individuals per species with data. The resulting likelihoods for 99% of all sites were used to estimate a genome-wide site frequency spectrum for each species separately, using emOptim2. This site frequency spectrum was used as a prior to estimate posterior probabilities for per-site allele frequencies (-pest). For this step, sites were filtered as previously, with the additional requirement of a minimum depth of 9. The posterior probabilities of per-site allele frequency spectra were provided as input to the ngsTools programs ngsStat and ngsFST (https://github.com/mfumagalli/ngsTools) (Fumagalli 2013; Fumagalli et al. 2013). These programs output summary statistics, which were then summarized over 20 kb windows to assess the genome-wide distribution of the statistics Ps1 and FST. Specifically, we calculated Ps1 for each species as ∑ss/∑Pvar, where ss represents the persite probability of a segregating site in individuals of that species, and Pvar represents the per-site probability of a segregating site in the whole sample. We used the per-species heterozygosity (2pq) and dXY (p1q2 + p2q1) output by ngsStat as HW and HB, respectively, to calculate single species FST as 1 – Hw/HB. Across regions, we summarized FST as 1 – ∑Hw/∑HB. For the calculation of both statistics, we excluded sites with Pvar < 0.8. We excluded regions in which <90% of sites passed the data filters above from analysis. Appendix S20. Assessment of admixture in the combined sample We used a subset of high confidence polymorphic sites (Pvar ≥ 0.8 as in S19) in the combined sample to perform principal components analysis (PCA) and an estimation of admixture proportions. To minimize linkage disequilibrium between sites, we randomly sampled sites from scaffolds longer than 100 kb, selecting at most one site per 100 kb. For PCA, we first called genotypes using ANGSD –doGeno 32, and subsequently estimated the covariance matrix among sites using ngsCovar (Fumagalli 2013; Fumagalli et al. 2013). We plotted the first two principal components, which represent 34.6% and 12.4% of the total variation, respectively (Figure S5a). For the estimation of admixture proportions, we generated Beagle format genotype likelihoods using ANGSD –doGlf 2, and then ran NGSadmix (Skotte et al. 2013) with k = 2 to infer the proportion of each sample derived from two source populations (Figure S5b). The command lines for angsd, ngsCovar, and NGSadmix were as follows: angsd -bam $BAMLIST -nInd 8 -doGeno 32 -doPost 1 -out $PCAOUT -doGlf 2 -P 5 -sites $SITESFILE -GL 1 -doMajorMinor 1 -doMaf 2 ngsCovar -probfile $PCAOUT.geno -outfile $PCAOUT.covar -nind 8 -nsites $N_SITES -call 0 NGSadmix -likes $PCAOUT.beagle.gz -K 2 -P 4 -o $PCAOUT.ngsadmix Appendix S21. Annotation of candidate selected regions and gene ontology analysis We annotated genes in the regions identified in SI Text S17 (from the two-sample dataset, using 100 kb sliding windows, and from the full dataset, using 20 kb non-overlapping windows) by aligning human protein sequences to the blue-eyed black lemur genome. We obtained protein sequences for human genome build hg18 and used TBLASTN version 2.2.22+ (Altschul et al. 1990, 1997), with an e-value threshold of 10-5 (two-sample dataset) or 5 x 10-5 (full dataset) to identify orthologs within the regions of the blue-eyed black lemur reference genome corresponding to the 3.9% PS1 tail (two-sample dataset) or 1% FST tail (full dataset). We then took the list of all human proteins with hits within candidate regions and performed TBLASTN for these proteins against the entire lemur genome. We retained proteins whose best genome-wide match (containing the lowest e-value or maximum mean percent identity) for any subset of the protein sequence overlapped the candidate region. In cases in which multiple proteins mapped to the same location (>50% protein length overlapping, presumably representing multiple transcripts of the same gene or multiple genes in the same family), we retained the protein with the largest total length spanned by initial TBLASTN hits or the largest mean percent identity. If multiple proteins were equivalent by these two tests (generally representing multiple transcripts of the same gene), we selected one at random for further testing. All transcripts mapped to regions in the 1% tail of FST from the full dataset in either species are provided in Dataset S1 (available on Dryad at doi:10.5061/dryad.rn745). We converted the candidate lists to unique Ensembl gene IDs for gene ontology (GO) enrichment analysis. We tested for an enrichment of specific GO categories among genes annotated within candidate regions for the blue-eyed black lemur from the full dataset using the Database for Annotation, Visualization and Integrated Discovery (DAVID) v6.7 (Huang et al. 2008, 2009). In order to focus on genes with evidence for selection specific to the blue-eyed black lemur lineage, we excluded any genes mapping to regions with black lemur FST in the 20% tail. We performed functional annotation with default settings in DAVID using this candidate gene list, subsampled to include only one gene per region, with the background gene list generated from the hg18 unique Ensembl gene IDs. We report fold enrichment and EASE score (adjusted Fisher Exact Test pvalue) from the functional annotation chart. Our initial analysis indicated an enrichment of genes related to melanocyte development (Biocarta, 50.5-fold enrichment, p = 0.034) and within the melanogenesis pathway (KEGG, 7.3-fold enrichment, p = 0.057); however, both of these categories have median gene lengths that are substantially longer than those of the background gene set. To correct for this, we ranked all genes in the background set by longest transcript length and selected the sets of genes that would result in the same median gene length as those of the categories of interest, starting with the longest genes; we then used these background sets to re-run the GO enrichment analysis. Melanocyte development was the most enriched category in our initial analysis. Other GO categories with initial enrichments at least as strong as those for melanogenesis were U2A'/phosphoprotein 32 family A, C-terminal (Interpro, 41.8-fold enrichment, p = 0.046); Spectrin repeat (Interpro, 23.9-fold enrichment, p = 0.079) and repeat: Spectrin 1, 2, 3, 4, and 5 (UP_SEQ_FEATURE, 18.9 – 35.0-fold enrichment, p = 0.055 – 1); LRRcap (SMART, 34.6-fold enrichment, p = 0.055); single fertilization and fertilization (GOTERM_BP_FAT, 12.8- and 9.7-fold enrichment, p = 0.022 and 0.037, respectively); and transcription factor (SP_PIR_KEYWORDS, 11.2-fold enrichment, p = 0.029). Appendix S22. Genome size estimation One way to estimate the size of a genome is by dividing the total sequence contained within kmers of a given length by the peak depth of such k-mers. To do this, we calculated the total sequence length of 17-mers within trimmed reads from the blue-eyed black lemur data (Ktotal). The 17-mers present only once are unique k-mers, which are likely sequencing errors, and considering them can lead to overestimation of the genome size. Thus, we subtracted Kunique (the total sequence length of unique 17-mers) from Ktotal and divided the result by the peak 17-mer sequencing depth to obtain the estimated genome size. Ktotal = 130.259 Gb Kunique = 1.554 Gb Peak sequencing depth = 48X The genome size is thus estimated to be (130.259 – 1.554) / 48 = 2.681 Gb. This estimate corresponds well to molecular weight-based genome size estimates for the black lemur, which range from 2.62 to 3.51 Gb (http://www.genomesize.com). In particular, the most recent molecular estimate based on flow cytometry indicates a genome size of approximately 2.62 Gb (Krishan et al. 2005). Appendix S23. Identification of neighboring scaffolds to the scaffold containing the OCA2 ortholog In order to investigate patterns of divergence and diversity beyond the 600 kb in scaffold 2503, where the orthologs of HERC2 and OCA2 reside, we identified the neighboring scaffolds using two different methods. First, we used BLASTN to identify the regions of the human genome (build hg19) orthologous to the first and last 100 kb of scaffold 2503. We then downloaded these sequences, along with the 100 kb beyond the mapping position of the end of scaffold 2503, from the UCSC genome browser (http://genome.ucsc.edu/), and used BLASTN to compare each region to the lemur assembly. Assuming synteny had been conserved to human, we inferred that the scaffold with the highest scoring match to the 100 kb beyond the mapping position of scaffold 2503 was likely the adjacent scaffold. The BLASTN results indicated that the first 60 kb of scaffold 3393 “preceded” scaffold 2503 (i.e., was adjacent to the 0 end) with approximately 1.6 kb of unmappable sequence separating the two, and that scaffold 2174 “followed” scaffold 2503 (i.e., was adjacent to the 600 kb end), separated by approximately 7 kb. We additionally used information about the mapping of MP paired reads to provide further evidence that the first 60 kb of scaffold 3393 mapped adjacent to the 0 end of scaffold 2503. We used SAMtools to identify read pairs from the unfiltered BAM file that did not map as “proper pairs” with both reads mapping in the appropriate orientation to the same scaffold (using “view –f 2”), and in which one read mapped to scaffold 2503. Reads within the first 3 kb and 8 kb of scaffold 2503 had pairs mapping to 56.6 kb and 51.5 kb to 59.2 kb of scaffold 3393 for 3 kb and 8 kb MP libraries, respectively. No reads within the last 8 kb of scaffold 2503 had pairs that mapped to different scaffolds, suggesting that the “following” scaffold was separated by too long a region of unmappable DNA (presumably repetitive DNA) to be identified in this way. The patterns of diversity and divergence across scaffold 2174 suggested that the orientation of this scaffold was not the same as that of scaffold 2503, relative to the human genome. We hypothesize that an inversion may have occurred within this region since the ancestor of lemurs and humans, and we represent scaffold 2174 in Figure 4a (the scaffold to the right of scaffold 2503) assuming such an inversion. Supporting References Abecasis GR, Altshuler D, Auton A et al. (2010) A map of human genome variation from population-scale sequencing. Nature, 467, 1061–73. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. Journal of Molecular Biology, 215, 403–10. Altschul SF, Madden TL, Schäffer AA et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids research, 25, 3389–402. Auton A, Fledel-Alon A, Pfeifer S et al. (2012) A fine-scale chimpanzee genetic map from population sequencing. Science (New York, N.Y.), 336, 193–8. Benaglia T, Chauveau D, Hunter DR, Young DS (2009) mixtools: An R Package for Analyzing Mixture Models. J Stat Softw, 32, 1–29. Brent RP (Richard P. (1973) Algorithms for minimization without derivatives. Prentice-Hall, Englewood Cliffs ;[Hemel Hempstead. Chaisson MJ, Brinza D, Pevzner PA (2009) De novo fragment assembly with short mate-paired reads: Does the read length matter? Genome research, 19, 336–46. Delaneau O, Zagury J-F, Marchini J (2013) Improved whole-chromosome phasing for disease and population genetic studies. Nature methods, 10, 5–6. DePristo M a, Banks E, Poplin R et al. (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genetics, 43, 491–8. Dubchak I, Poliakov A, Kislyuk A, Brudno M (2009) Multiple whole-genome alignments without a reference organism. Genome research, 19, 682–9. Earl D, Bradnam K, St John J et al. (2011) Assemblathon 1: a competitive assessment of de novo short read assembly methods. Genome research, 21, 2224–41. Eiberg H, Troelsen J, Nielsen M et al. (2008) Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Human Genetics, 123, 177–87. Freedman AH, Gronau I, Schweizer RM et al. (2014) Genome Sequencing Highlights the Dynamic Early History of Dogs (L Andersson, Ed,). PLoS Genetics, 10, e1004016. Frudakis T, Thomas M, Gaskin Z et al. (2003) Sequences associated with human iris pigmentation. Genetics, 165, 2071–2083. Fumagalli M (2013) Assessing the effect of sequencing depth and sample size in population genetics inferences. (L Orlando, Ed,). PLOS ONE, 8, e79667. Fumagalli M, Vieira FG, Korneliussen TS et al. (2013) Quantifying population genetic differentiation from next-generation sequencing data. Genetics, 195, 979–92. Gnerre S, Maccallum I, Przybylski D et al. (2011) High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proceedings of the National Academy of Sciences of the United States of America, 108, 1513–8. Hellenthal G, Stephens M (2007) msHOT: modifying Hudson’s ms simulator to incorporate crossover and gene conversion hotspots. Bioinformatics (Oxford, England), 23, 520–1. Hellström AR, Watt B, Fard SS et al. (2011) Inactivation of Pmel alters melanosome shape but has only a subtle effect on visible pigmentation. (IJ Jackson, Ed,). PLoS genetics, 7, e1002285. Huang DW, Sherman BT, Lempicki RA (2008) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protocols, 4, 44–57. Huang DW, Sherman BT, Lempicki R a (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Research, 37, 1–13. Hudson RR (2002) Generating samples under a Wright-Fisher neutral model of genetic variation. Bioinformatics (Oxford, England), 18, 337–8. Hwang DG, Green P (2004) Bayesian Markov chain Monte Carlo sequence analysis reveals varying neutral substitution patterns in mammalian evolution. Proceedings of the National Academy of Sciences of the United States of America, 101, 13994–14001. Idury RM, Waterman MS (1995) A new algorithm for DNA sequence assembly. Journal of computational biology : a journal of computational molecular cell biology, 2, 291–306. Kanetsky P a, Swoyer J, Panossian S et al. (2002) A polymorphism in the agouti signaling protein gene is associated with human pigmentation. American journal of human genetics, 70, 770– 5. Kaplan NL, Hudson RR, Langley CH (1989) The “hitchhiking effect” revisited. Genetics, 123, 887–99. Kelley DR, Schatz MC, Salzberg SL (2010) Quake: quality-aware detection and correction of sequencing errors. Genome Biology, 11, R116. Kim S-H, Elango N, Warden C, Vigoda E, Yi S V (2006) Heterogeneous genomic molecular clocks in primates. PLoS genetics, 2, e163. Kong A, Barnard J, Gudbjartsson DF et al. (2004) Recombination rate and reproductive success in humans. Nature Genetics, 36, 1203–6. Kong A, Frigge ML, Masson G et al. (2012) Rate of de novo mutations and the importance of father’s age to disease risk. Nature, 488, 471–5. Krishan A, Dandekar P, Nathan N et al. (2005) DNA index, genome size, and electronic nuclear volume of vertebrates from the Miami Metro Zoo. Cytometry - Part A, 65, 26–34. Kurtz S, Phillippy A, Delcher AL et al. (2004) Versatile and open software for comparing large genomes. Genome biology, 5, R12. Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics (Oxford, England), 25, 1754–60. Li H, Durbin R (2011) Inference of human population history from individual whole-genome sequences. Nature, 475, 493–6. Li H, Handsaker B, Wysoker A et al. (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics (Oxford, England), 25, 2078–9. Li R, Zhu H, Ruan J et al. (2010) De novo assembly of human genomes with massively parallel short read sequencing. Genome Research, 20, 265–72. Liu F, van Duijn K, Vingerling JR et al. (2009) Eye color and the prediction of complex phenotypes from genotypes. Current biology : CB, 19, R192–3. Liu F, Wollstein A, Hysi PG et al. (2010) Digital quantification of human eye color highlights genetic association of three new loci. PLoS genetics, 6, e1000934. Loftus SK, Larson DM, Baxter LL et al. (2002) Mutation of melanosome protein RAB38 in chocolate mice. Proceedings of the National Academy of Sciences of the United States of America, 99, 4471–6. Lunter G, Goodson M (2011) Stampy: a statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome research, 21, 936–9. Luo R, Liu B, Xie Y et al. (2012) SOAPdenovo2: an empirically improved memory-efficient shortread de novo assembler. GigaScience, 1, 18. Lynch M (2010) Evolution of the mutation rate. Trends in genetics : TIG, 26, 345–52. Marçais G, Kingsford C (2011) A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics (Oxford, England), 27, 764–70. McKenna A, Hanna M, Banks E et al. (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome research, 20, 1297–303. McVicker G, van de Geijn B, Degner JF et al. (2013) Identification of genetic variants that affect histone modifications in human cells. Science (New York, N.Y.), 342, 747–9. Meyer WK, Zhang S, Hayakawa S, Imai H, Przeworski M (2013) The convergent evolution of blue iris pigmentation in primates took distinct molecular paths. American Journal of Physical Anthropology, 151, 398–407. Miller JR, Delcher AL, Koren S et al. (2008) Aggressive assembly of pyrosequencing reads with mates. Bioinformatics (Oxford, England), 24, 2818–24. Müller S, O’Brien PCM, Ferguson-Smith MAF, Wienberg J (1997) Reciprocal chromosome painting between human and prosimians (Eulemur macaco macaco and E. fulvus mayottensis). Cytogenetic and Genome Research, 78, 260–271. Parra G, Bradnam K, Korf I (2007) CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics (Oxford, England), 23, 1061–7. Parra G, Bradnam K, Ning Z, Keane T, Korf I (2009) Assessing the gene space in draft genomes. Nucleic Acids Research, 37, 289–97. Pevzner PA, Tang H, Waterman MS (2001) An Eulerian path approach to DNA fragment assembly. Proceedings of the National Academy of Sciences of the United States of America, 98, 9748–53. Phillippy AM, Schatz MC, Pop M (2008) Genome assembly forensics: finding the elusive misassembly. Genome biology, 9, R55. Pingault V, Ente D, Dastot-Le Moal F et al. (2010) Review and update of mutations causing Waardenburg syndrome. Human Mutation, 31, 391–406. Prado-Martinez J, Sudmant PH, Kidd JM et al. (2013) Great ape genetic diversity and population history. Nature, 499, 471–5. Salzberg SL, Phillippy AM, Zimin A et al. (2012) GAGE: A critical evaluation of genome assemblies and assembly algorithms. Genome research, 22, 557–67. Salzberg SL, Yorke JA (2005) Beware of mis-assembled genomes. Bioinformatics (Oxford, England), 21, 4320–1. Sandelin A, Alkema W, Engström P, Wasserman WW, Lenhard B (2004) JASPAR: an openaccess database for eukaryotic transcription factor binding profiles. Nucleic acids research, 32, D91–4. Schatz MC, Delcher AL, Salzberg SL (2010) Assembly of large genomes using secondgeneration sequencing. Genome research, 20, 1165–73. Schatz MC, Phillippy AM, Shneiderman B, Salzberg SL (2007) Hawkeye: an interactive visual analytics tool for genome assemblies. Genome biology, 8, R34. Schmidt D, Wilson MD, Ballester B et al. (2010) Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science (New York, N.Y.), 328, 1036– 40. Simpson JT, Durbin R (2012) Efficient de novo assembly of large genomes using compressed data structures. Genome research, 22, 549–56. Simpson JT, Wong K, Jackman SD et al. (2009) ABySS: a parallel assembler for short read sequence data. Genome research, 19, 1117–23. Skotte L, Korneliussen TS, Albrechtsen A (2013) Estimating Individual Admixture Proportions from Next Generation Sequencing Data. Genetics, 195, 693–702. Sturm R a, Duffy DL, Zhao ZZ et al. (2008) A single SNP in an evolutionary conserved region within intron 86 of the HERC2 gene determines human blue-brown eye color. American Journal of Human Genetics, 82, 424–31. Sulem P (2008) Two newly identified genetic determinants of pigmentation in Europeans. Nature genetics, 40, 835. Sulem P, Gudbjartsson DF, Stacey SN et al. (2007) Genetic determinants of hair, eye and skin pigmentation in Europeans. Nature genetics, 39, 1443–52. Treangen TJ, Salzberg SL (2012) Repetitive DNA and next-generation sequencing: computational challenges and solutions. Nature reviews. Genetics, 13, 36–46. Tsantes C, Steiper ME (2009) Age at first reproduction explains rate variation in the strepsirrhine molecular clock. Proceedings of the National Academy of Sciences of the United States of America, 106, 18165–70. Valenzuela RK, Henderson MS, Walsh MH et al. (2010) Predicting phenotype from genotype: normal pigmentation. Journal of forensic sciences, 55, 315–22. Visser M, Kayser M, Palstra R-J (2012) HERC2 rs12913832 modulates human pigmentation by attenuating chromatin-loop formation between a long-range enhancer and the OCA2 promoter. Genome Research, 22, 446–55. Volampeno MSN, Masters JC, Downs CT (2011) Life history traits, maternal behavior and infant development of blue-eyed black lemurs (Eulemur flavifrons). American journal of primatology, 73, 474–84. Wingender E, Dietze P, Karas H, Knüppel R (1996) TRANSFAC: a database on transcription factors and their DNA binding sites. Nucleic acids research, 24, 238–41. Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome research, 18, 821–9. Supporting Information Figures Pipeline for generation and analysis of genomic data Contig: contiguous sequence of DNA obtained by merging overlapping k-mers Blue-eyed black lemur (Eulemur flavifrons) Black lemur (Eulemur macaco) k-mer: sequences of k bp in length, derived from sequencing reads and used in generating a de novo assembly Mate pair: outward-facing pairs of short (32 bp) reads sequenced from the ends of large (2 – 10 kb) circularized amplicons N50: The size of the smallest contig/scaffold such that 50% of the genome is contained in scaffolds of that size or larger. Paired-end: inward-facing pairs of short (50 – 150 bp) reads sequenced from the ends of small (200 – 1000 bp) amplicons Scaffold: sets of contigs merged using paired-end or mate pair data, with Nʼs separating contiguous sequence 1) Prepare genomic DNA libraries Additional individuals resequenced to low coverage Photos courtesy D. Haring 180, 500, 1000 bp insert size paired-end 500 bp insert size paired-end 350 bp insert size paired-end 3 kb + 8 kb mate-pair 2) Sequence 9 lanes Illumina HiSeq (achieved 52x mean coverage) 1 lane Illumina HiSeq (achieved 21x mean coverage) 1 lane Illumina HiSeq (all 6 samples barcoded and pooled) (achieved 4 - 15x coverage) 3) Perform quality control Filter reads with FASTX-Toolkit Filter reads with FASTX-Toolkit Filter reads with FASTX-Toolkit Align to assembly using bwa (Li and Durbin 2009) and SAMtools (Li et al. 2010) Align to assembly using bwa (Li and Durbin 2009) and SAMtools (Li et al. 2010) Trim reads with bwa (Li and Durbin 2009) Error correction with Quake (Kelley et al. 2010) 4) Assemble/ align de novo assembly using SOAPdenovo (Li et al. 2009) (scaffold N50: 420 kb) Generate reference-based assembly using custom python script 5) Identify variable sites Identify polymorphic and divergent sites using GATK (DePristo et al. 2011) Identify polymorphic and divergent sites using GATK (DePristo et al. 2011) Estimate regional heterozygosity and divergence Estimate Ne and species split time, and identify signatures of selection Use PSMC (Li and Durbin 2011) to estimate historic Ne Generate genotype likelihoods using ANGSD and ngsTools (Fumagalli et al. 2013) Estimate regional heterozygosity and divergence Identify signatures of selection Figure S1. Overview of assembly and analysis pipeline. A schematic highlighting the steps in the assembly and variant detection pipelines for the two individuals sequenced to high coverage, as well as the additional samples sequenced to low coverage for further population genetic analyses. In the upper left is a brief glossary of key assembly-related terms. Figure S2. Peak memory consumption and time to completion for genome assembly steps. Jellyfish: q-mer counting; Quake: error correction; Graph: de Bruijn graph construction; Contig: merging sequences into contigs; Map: mapping reads to contigs; Scaffold: using PE information to construct scaffolds; Gap: using local assemblies to close scaffold gaps. Figure S3. Simulations to assess impact of window size, scaffolds, generation time, mutation rate, adjustment for coverage, and population split on output of PSMC. A) Different window sizes produce similar PSMC output. B) PSMC output for scaffolds is comparable to that for full data. The simulated trajectory was simulated using the parameters in SI Text S13. Figure S3, continued. C) True and inferred trajectories including a population split. Red: PSMC estimate, Purple: True (simulated) trajectories; Green: Bootstrap estimates. D) Histogram of the bootstrapped PSMC estimates of the population split time, using the same scaling parameters as in Figure 4. The red dashed line denotes the simulated split time. E) Influence of mutation rate estimates on inferred demographic history. F) Influence of generation time estimates on inferred demographic history. G) Adjusted mutation rate provides an adequate correction for the reduced ability to call SNPs in lower coverage data. 0 20 40 60 Mapped read coverage 80 0.020 0.000 0.010 Proportion of all k−mers 0.020 0.000 0.010 Proportion of all q−mers 2.5 2.0 1.5 1.0 0.5 0.0 Percent of genome C B A 0 10 20 30 40 Q−mer coverage 50 60 0 50 100 150 200 250 K−mer coverage Figure S4. Coverage distributions for mapped reads, q-mers, and k-mers. A) Distribution of mapped read coverage for blue-eyed black lemur assembly. B) Distribution of coverage weighted by quality score for 17-mers, which was used to choose the coverage cutoff of 3 (red line) for Quake error correction. The mean of the distribution excluding all 17-mers with coverage ≤ 3 (assumed to represent true kmers) is 23.35, and the variance is 60.72. This shows that k-mer coverage distribution before error correction does not correspond to the expectation of a single Poisson, likely due in part to sequencing errors. C) Distribution of frequency of 17-mers in the unfiltered dataset. Peak k-mer coverage is at 48X. The first valley in k-mer frequency is at 14, shown by the red line. We used this coverage cutoff to correct the reads using SOAP’s error correction. For A) and B), the plotted region represents 99.5% of the total. For C), the line at 250 represents k-mers from repetitive regions that are present more than 250 times. Figure S5. Principal components analysis (PCA) and estimation of admixture proportions indicate the absence of admixture in the combined sample. A) The samples’ projections along the first and second principal component are displayed, with blue-eyed black lemurs in blue and black lemurs in orange. The majority of variation separates the two species. B) Shown are inferred proportions of ancestry from each of two source populations (arbitrarily colored in purple and green), with Ef1 – 4 representing blue-eyed black lemur samples and Em1 – 4 black lemur samples. None of the samples appears to have admixed ancestry. B 0 200 400 600 800 1000 1200 insert size C 30000 10000 0 500 1000 1500 2000 2500 insert size 2e+05 4e+05 good 1Kb library 0e+00 frequency bad 1Kb library 0 100000 frequency 250000 bad 500bp library 0 frequency A 0 200 400 600 800 1000 1200 1400 insert size Figure S6. Bimodal distributions of estimated insert sizes indicate the presence of artifacts in some library preparations. The mean insert sizes estimated from fitting a mixture model with two normal distributions using mixtools (Benaglia et al. 2009) in R were 205 and 480 for the 500 bp library (A). For the first 1 kb library (B), the mean insert sizes from the mixture model were 371 and 958 bp, respectively. The second 1 kb library (C) did not show evidence of an artifactual inclusion of smaller insert sizes. 3000 0 1000 2000 Frequency 6000 B 0 Frequency A 0.0 0.2 0.4 0.6 0.8 1.0 0.0 0.2 Blue−eyed black lemur PS1 (two samples) 0.8 1.0 3000 0 1000 Frequency 4000 2000 0 Frequency 0.6 D 6000 C < −1.0 −0.5 0.0 0.5 < −1.0 1.0 Blue−eyed black lemur FST (two samples) −0.5 0.0 0.5 1.0 Blue−eyed black lemur FST (two samples) F 0 600 0 200 200 400 Frequency 600 1000 800 E Frequency 0.4 Black lemur PS1 (two samples) 0.0 0.2 0.4 0.6 0.8 0.0 0.2 Blue−eyed black lemur PS1 (full dataset) 0.4 0.6 0.8 1.0 Black lemur PS1 (full dataset) H 600 0 200 400 Frequency 400 200 0 Frequency 600 800 800 G < 0.0 0.2 0.4 0.6 0.8 Blue−eyed black lemur FST (full dataset) 1.0 < 0.0 0.2 0.4 0.6 0.8 1.0 Black lemur FST (full dataset) Figure S7. Distributions of summary statistics from scans for selection in two-sample and full datasets. Histograms of Ps1 and FST for each species, summarized across 20 kb windows. A) Ps1 for the blue-eyed black lemur from the two-sample dataset. B) Ps1 for the black lemur from the two-sample dataset. C) FST for the blue-eyed black lemur from the two-sample dataset. Figure S7, continued. D) FST for the black lemur from the two-sample dataset. E) Ps1 for the blue-eyed black lemur from the full dataset. F) Ps1 for the black lemur from the full dataset. G) FST for the blue-eyed black lemur from the full dataset. H) FST for the black lemur from the full dataset. Supporting Information Tables Table S1. Read counts for each blue-eyed black lemur library Libraries (Fragment Sizes) # of Raw Reads (read length) # of lanes # of Quake corrected Reads (read length) # of SOAP corrected reads (read length) 180bp 1 157441980 (100) 155080642 (80.76) 130918792 (99) 500bp 6 1211058780 (100) 961903196 (79.13) 851845445 (84) 1Kb 3 574221050 (100) 564925769 (86.74) 514601130 (91.5) 3 Kb / 8Kb 1 318329630 (35) 194896994 (35) 194896994 (35) Total 11 2261051440 1876806601 1692262361 Average read lengths are shown in parentheses below the read counts. Although SOAP discards more reads as errors, it produces a longer corrected read length and achieves comparable coverage to that from Quake. Table S2. Statistics for QCA and SCA S2a Assembly statistics prior to resolution of bimodal insert size libraries Estimate d Genome Size (Gb) QCA 2.6813 SCA 2.6813 Largest contig %chaf contig scaffold # of N50 # of f size N50 scaffold (kb) contigs bases (kb) (Mb) s 13.0 398910 0.88 178.1 328.6 28413 13.9 385282 1.38 156.5 197.0 34879 % Largest single scaffold tons size 2.1 2423633 2.7 1708178 %bases assemble d from scaffolds 80.46 79.8 S2b Assembly statistics following resolution of bimodal insert size libraries QCA 2.6813 SCA 2.6813 16.3 17.9 280211 0.39 263463 0.86 222.3 195.1 421 323.1 21210 22730 0.6 1.1 3296881 79.8 2934864 79.6 Table S3 Primers Name Sequence EmEf_12197_F GCCATCCTGTTTTCATTTCG Position on scaffold 2503 12417 EmEf_12197_R GTGTCTGAAAGCCCATCTCC 13653 EO-1R EmEf_35470_F CAGAGTCTTTCCCGACCTTG 35725 EO-2F EmEf_35470_R GGAAGGGTGAGAAAGGTGGT 36928 EO-2R EmEf_67146_F TGCACTTGAGTGAAGGACCTAA 67308 EO-3F EmEf_3int_R ATCCCATCATTCTGCCTCTG 67937 EO-3intR EmEf_67146_R AAAGGTCTTCTGCGACTTGC 68551 EO-3R EmEf_142843_F TCTCCTTTCCCTGCTCTGTG 143079 EO-5F EmEf_142843_R GGACCAGTGAAGGCAAGATT 144299 EO-5R EmEf_192282_F GGGTTTTGGTTGTAGGTCCA 192539 EO-7F EmEf_192282_R CCAAAGGAGAAAAGCAGGAG 193761 EO-7R EmEf_211317_F ATTTAAGTGGGTGCCAATGC 211500 EO-8F EmEf_211317_R CCACTGAATAGGAGAAAACATCC 212730 EO-8R EmEf_263789_F TAGTGGGAATGGGAAGCAAA 263979 EO-9F EmEf_263789_R TGTTTCCAAACTGCGGTCTA 265211 EO-9R EmEf_335375_F AGGGGCAACCTAATGCTCTC 335554 EO-12F EmEf_12int_R CCTTTTACAGTGGCCTGTAGC 336282 EO-12intR EmEf_335375_R GATGTGGGGGCAGAGTGTAG 336791 EO-12R EmEf_357225_F ATTTTCCTGAGCCCTTCTGG 357419 EO-12.5F EmEf_357225_R CCTCACGGCAGATTCTTAGC 358647 EO-12.5R EmEf_371731_F CAGGCCATTTCTTTCCCTTT 371917 EO-13F EmEf_13int_F CTGCAGGAAAACTCGTGGAT 372462 EO-13intF EmEf_371731_R TAGACATGTCCCAGCTCCTG 373168 EO-13R EmEf_405684_F GATTGCTGGCCAGAGTTTTT 405885 EO-14bF EmEf_405684_R ATCAAAGCTAGCACCCCAAA 407113 EO-14bR EmEf_447752_F TCCACGGTCTATTTGTTTGG 447978 EO-15bF EmEf_447752_R CCTTCAAGCGTGACAATTCC 449223 EO-15bR EmEf_450326_F CGTTGGCACATCTCCACTTA 450549 EO-16bF EmEf_450326_R AGCTGAGCAATCCCTGATGT 451782 EO-16bR EmEf_492021_F CATTTTTCATCTCCGCCAGT 492262 EO-17bF EmEf_492021_R ACCCTGAGTTAAGCAAAGATTG 493491 EO-17bR EmEf_509600_F GGAAAGAGCTGGAGGAACAA 509841 EO-17F EmEf_17int_F GCAGCATGCACTGTCTTGAT 510724 EO-17intF EmEf_509600_R AGTGGCTGAAAGCAGAGTCC 511059 EO-17R EmEf_513958_F AGGGGTTCTTGAGCTCTGTG 514179 EO-18F EmEf_18int_R TGCTTAATGGCCTTCAGAGG 514908 EO-18intR SeqName Notes EO-1F internal primer for amplicon 3 internal primer for amplicon 12 internal primer for amplicon 13 internal primer for amplicon 17 internal primer for amplicon 18 EmEf_513958_R GGGGGTTATGGCTTCAACTT 515411 EO-18R EmEf_591196_F GACAGTACTGGGGGCTCAAA 591291 EO-20F EmEf_20int_F GTATCCGGGAGCAGTTCTCA 591829 EO-20intF EmEf_591196_R TGGAGGTCATGCCTCTTTTC 592616 EO-20R internal primer for amplicon 20