Welcome to CSE 527:
Computational Biology
Lecture 1 – Sep 27, 2011
CSE 527 Computational Biology, Fall 2011
Instructor: Su-In Lee
TA: Christopher Miles
Monday & Wednesday 12:00-1:20
Johnson Hall (JHN) 022
1
Who is the instructor?
Prof. Su-In Lee
Assistant Professor
A joint faculty member
Computer Science & Engineering, Genome Sciences
Office hours: Wednesday 1:30-2:30
Research interests
Developing machine learning techniques applied to
Computational Biology (genetics, systems biology)
Predictive Medicine, Translational Medicine
2
Teaching assistant
Christopher Miles (CSE PhD student)
Office: TBA
Office hours: Monday 1:30-2:30
Email: cmiles@uw.edu
3
What is the Coolest Thing a
Computational/Mathematical Scientist
Can Do?
Curing cancer.
Understanding how the blue print of life (DNA)
determines important traits (e.g. diseases)?
Predicting your disease susceptibilities based on your
biological information including DNA sequence.
Predicting sudden changes in the condition of patients
at ICU (intensive care unit).
Determining the order of A,G,C and T in my 3-billion
long DNA sequence.
:
CSE 527 will provide you with basic
concepts and ML/statistical techniques
that you can use to realize these goals.
A cell’s biological
state can
beBiology
described byis
millions of
More
and
More
of
numbers!
What biological discoveries we can make highly depends on
Becoming
an method
Information
Science
the computational
we use to analyze
the data.
Machine learning techniques provide very effective tools.
Gene
(~30,000 in human)
Gene regulation
AGATATGTGGATTGTTAGGATTTATGCGCGTCAGTGACTACGCATGTTACGCACCTACGACTAGGTAATGATTGATC
DNA
RNA
Protein
Gene expression
AUGUGGAUUGUU
AUGCGCGUC
AUGCGCGUC
AUGUUACGCACCUAC
RNA
degradation
MWIV
MRV
MRV
MLRTY
Gene interaction map
Biological information (data)
DNA sequence information
RNA levels of 30K genes
Protein levels of 30K genes
DNA molecule’s 3D structure
:
AUGAUUGAU
AUGAUUAU
MID
Cell: The basic
unit of life
Outline
Course logistics
A zero-knowledge based introduction to biology
Potential project topics
6
Goals of this course
Introduction to Computational Biology
Basic concepts and scientific questions
Basic biology for computational scientists
In-depth coverage of ML techniques
Current active areas of research
Useful machine learning (ML) algorithms
Probabilistic graphical models, clustering, classification
Learning techniques (MLE, EM)
7
Topics in CSE 527
Part 1: Basic ML algorithms
Introduction to probabilistic models
Bayesian networks, Hidden Markov models
Representation and learning
Part 2: Topics in computational biology and
areas of active research
Genetics, systems biology, predictive medicine,
sequence analysis
Finding genetic factors for complex biological traits
Inferring biological networks from data
Comparative genomics
DNA/RNA sequence analysis
8
Course responsibilities
Class participation and attendance (10%)
Good answers to the questions asked in class
Initiating a productive discussion.
Homework assignments (40%)
Four problem sets
Collaboration allowed
Due at beginning of class
Up to 3 late days (24-hr period) for the quarter
Teams of 2 or 3 students
Individual writeups
Final project (50%)
A group of up to two students.
9
Project overview (1/2)
Topic
Choose from the list of project topics on the course
website, or come up with your own.
Open-ended
Project deliverables
Project proposal (due 10/19)
Midterm report (due 11/16)
Final report (due 12/14)
Final presentations or poster session (12/7)
10
Project overview (2/2)
Final report
Short report (up to 10 pages)
Conference-style presentation
Successful project reports can be submitted to
computational biology/ ML conferences (ISMB,
RECOMB, NIPS, ICML)
Or journals (PLoS journals, Nature journals, PNAS,
Genome Research and so on)
11
Reading material
Lecture notes
Biological background
The Cell, a molecular approach by Copper
Genetics, from genes to genomes by Hartwell and more
Principles of Population genetics by Hartl & Clark
Computational background
Mostly based on recent papers & old seminar papers
Probabilistic graphical models by Profs. Daphne Koller &
Nir Friedman
Prof. Andrew Ng’s machine learning lecture note
(cs229.stanford.edu)
No textbook required for the course
12
Class resources
Course website – cs.washington.edu/527
Lecture notes, assignments, project topics
Deadlines of assignments and projects
Mailing list
cse527@cs.washington.edu
13
Outline
Course logistics
A zero-knowledge based introduction to biology
Prepared by George Asimenos (PhD student,
Stanford) for CS262 Computational Genomics by
Prof. Serafim Batzoglou (Stanford).
Potential project topics
14
Cells: Building Blocks of Life
cell, nucleus, cytoplasm, mitochondrion
Eukaryots:
Plants, animals, humans
DNA resides in the nucleus
Contain other compartments
for other specialized
functions
Prokaryots:
Bacteria
Do not contain compartments
Little recognizable
substructure
© 1997-2005 Coriell Institute for Medical Research
15
DNA: “Blueprints” for a cell
Genetic information encoded in long
strings of double-stranded DNA
Deoxyribo Nucleic Acid comes in only
four flavors: Adenine, Cytosine,
Guanine, Thymine
16
Nucleotide
Deoxyribose, nucleotide, base, A, C, G, T, 3’, 5’
to previous nucleotide
O
O
P
O-
5’
H
O
H
C
H
Guanine (G)
Thymine (T)
Cytosine (C)
to base
O
C
Adenine (A)
C
H
H
C
C
3’
H
H
Let’s write “AGACC”!
to next nucleotide
17
“AGACC” (backbone)
18
“AGACC” (DNA)
deoxyribonucleic acid (DNA)
3’
5’
5’
3’
19
DNA is double stranded
strand, reverse complement
5’
3’
3’
5’
DNA is always written 5’ to 3’
AGACC or GGTCT
20
DNA Packaging
histone, nucleosome, chromatin, chromosome, centromere, telomere
telomere
centromere
nucleosome
DNA
chromatin
H1
~146bp
H2A, H2B, H3, H4
21
The Genome
The genome is the full set of hereditary
information for an organism
Humans bundle two copies of the genome into 46
chromosomes in every cell
= 2 x (1-22 + X/Y)
22
Building an organism
DNA
cell
Every cell has the same sequence of DNA
Subsets of the DNA sequence
determine the identity and
function of different cells
23
From DNA To Organism
?
Proteins do most of the work in biology,
and are encoded by subsequences of DNA,
known as genes.
24
RNA
ribonucleotide, U
to previous ribonucleotide
O
O
P
O-
5’
H
O
H
C
H
3’
Guanine (G)
Uracil (U)
Cytosine (C)
to base
O
C
Adenine (A)
C
H
H
C
C
OH
to next ribonucleotide
H
TU
25
Genes & Proteins
gene, transcription, translation, protein
Double-stranded DNA
5’
3’
TAGGATCGACTATATGGGATTACAAAGCATTTAGGGA...TCACCCTCTCTAGACTAGCATCTATATAAAACAGAA
ATCCTAGCTGATATACCCTAATGTTTCGTAAATCCCT...AGTGGGAGAGATCTGATCGTAGATATATTTTGTCTT
3’
5’
(transcription)
Single-stranded RNA
AUGGGAUUACAAAGCAUUUAGGGA...UCACCCUCUCUAGACUAGCAUCUAUAUAA
(translation)
protein
26
Gene Transcription
promoter
5’
3’
G A T T A C A . . .
C T A A T G T . . .
3’
5’
27
Gene Transcription
transcription factor, binding site, RNA polymerase
5’
3’
G A T T A C A . . .
C T A A T G T . . .
3’
5’
Transcription factors: a type of protein that binds to
DNA and helps initiate gene transcription.
Transcription factor binding sites: short sequences of
DNA (6-20 bp) recognized and bound by TFs.
RNA polymerase binds a complex of TFs in the promoter.
28
Gene Transcription
5’
3’
3’
5’
The two strands are separated
29
Gene Transcription
5’
3’
3’
5’
An RNA copy of the 5’→3’ sequence is
created from the 3’→5’ template
30
Gene Transcription
G A T T A C A . . .
5’
3’
3’
5’
C T A A T G T . . .
pre-mRNA
5’
G A U U A C A . . .
3’
31
RNA Processing
5’ cap, polyadenylation, exon, intron, splicing, UTR, mRNA
5’ cap
poly(A) tail
exon
intron
mRNA
5’ UTR
3’ UTR
32
Gene Structure
introns
5’
3’
promoter
5’ UTR
exons
3’ UTR
coding
non-coding
33
How many? (Human Genome)
Genes:
~ 20,000
Exons per gene:
~ 8 on average (max: 148)
Nucleotides per exon:
170 on average (max: 12k)
Nucleotides per intron:
5,500 on average (max: 500k)
Nucleotides per gene:
45k on average (max: 2,2M)
34
From RNA to Protein
Proteins are long strings of amino acids joined by
peptide bonds
Translation from RNA sequence to amino acid
sequence performed by ribosomes
20 amino acids 3 RNA letters required to
specify a single amino acid
35
Amino acid
amino acid
H
N
H
O
C
C
H
R
OH
Alanine
Arginine
Asparagine
Aspartate
Cysteine
Glutamate
Glutamine
Glycine
Histidine
Isoleucine
Leucine
Lysine
Methionine
Phenylalanine
Proline
Serine
Threonine
Tryptophan
Tyrosine
Valine
There are 20 standard amino acids
36
Proteins
N-terminus, C-terminus
to previous aa
N
H
O
C
C
to next aa
H
R
H
N-terminus
(start)
from 5’
OH
C-terminus
(end)
3’ mRNA
37
Translation
ribosome, codon
P site
A site
mRNA
The ribosome (a complex of protein and RNA) synthesizes a
protein by reading the mRNA in triplets (codons). Each codon is
translated to an amino acid.
38
The genetic code
Mapping from a codon to an amino acid
39
Translation
5’ . . . A U U A U G G C C U G G A C U U G A . . . 3’
UTR
Met
Start
Codon
Ala
Trp
Thr
Stop
Codon
40
Translation
amino acid
t-RNA
Met
Ala
Trp
5’ . . . A U U A U G G C C U G G A C U U G A . . . 3’
41
Errors?
mutation
What if the transcription / translation machinery makes
mistakes?
What is the effect of mutations in coding regions?
42
Reading Frames
reading frame
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
43
43
Synonymous Mutation
synonymous (silent) mutation, fourfold site
G
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
Ala
Cys
Leu
Arg
Ile
G C U U G U U U G C G A A U U A G
Ala
Cys
Leu
Arg
Ile
44
44
Missense Mutation
missense mutation
G
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
Ala
Cys
Leu
Arg
Ile
G C U U G G U U A C G A A U U A G
Ala
Trp
Leu
Arg
Ile
45
45
Nonsense Mutation
nonsense mutation
A
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
Ala
Cys
Leu
Arg
Ile
G C U U G A U U A C G A A U U A G
Ala
STOP
46
Frameshift
frameshift
G C U U G U U U A C G A A U U A G
G C U U G U U U A C G A A U U A G
Ala
Cys
Leu
Arg
Ile
G C U U G U
U A C G A A U U A G
Ala
Tyr
Cys
Glu
Leu
47
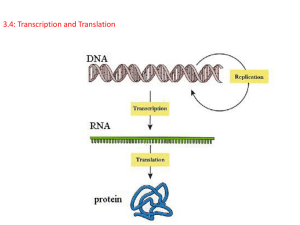
Transcription and translation
Let’s see how this happens!
Transcription: http://www.youtube.com/watch?v=DA2t5N72mgw
Translation: http://www.youtube.com/watch?v=WkI_Vbwn14g&feature=related
Illustration from Radboud University Nijmegen
48
Gene Expression Regulation
Regulation, signal transduction
When should each gene be expressed?
Regulate gene expression
Examples:
Make more of gene A when substance X is present
Stop making gene B once you have enough
Make genes C1, C2, C3 simultaneously
Why? Every cell has same DNA but each cell
expresses different proteins.
Signal transduction: One signal converted to
another
Cascade has “master regulators” turning on many proteins,
which in turn each turn on many proteins, ...
49
Gene Regulation
Gene expression is controlled at many levels
DNA chromatin structure
Transcription
Post-transcriptional modification
RNA transport
Translation
mRNA degradation
Post-translational modification
50
Transcription regulation
Much gene regulation occurs at
the level of transcription.
Primary players:
Binding sites (BS) in cis-regulatory
modules (CRMs)
Transcription factor (TF) proteins
RNA polymerase II
Primary mechanism:
TFs link to BSs
Complex of TFs forms
Complex assists or inhibits formation of
the RNA polymerase II machinery
51
Transcription Factor Binding Sites
Short, degenerate DNA sequences recognized by
particular TFs
For complex organisms, cooperative binding of
multiple TFs required to initiate transcription
Binding Sequence Logo
52
Summary
All hereditary information encoded in doublestranded DNA
Each cell in an organism has same DNA
DNA RNA protein
Proteins have many diverse roles in cell
Gene regulation diversifies protein products
within different cells
53
Outline
Course logistics
A zero-knowledge based introduction to biology
Potential project topics
54
Example project topic #1 (1/3)
Which Drug Patient X Should Be
Treated With?
Say that a cancer patient X undergoes a chemotherapy.
There are >200 drugs patient X can be treated with.
How do doctors choose which drug to use in
chemotherapy treatment ?
Follicular lymphoma
Diffuse large B cell
lymphoma
A few histologic
features
Patient X
How can we improve this?
Chemotherapy drugs
5-Iodotubercidin
Acrichine
ARQ-197
Arsenic trioxide
AS101
AS-703026
AT-7519
Axitinib
Azacitidine
:
Example project topic #1 (1/3)
Which Drug Patient X Should Be
Treated With?
…ACGTAGCTAGCT
AGCTAGCTGATGC
TAGCTACGTGCT…
A few histologic
features
Epigenetics
(Methylation)
DNA sequence
RNA levels
of genes
Follicular lymphoma
Diffuse large B cell
lymphoma
A few histologic
features
Patient X
Protein levels
of genes
Chemotherapy drugs
5-Iodotubercidin
Acrichine
ARQ-197
Arsenic trioxide
AS101
AS-703026
AT-7519
Axitinib
Azacitidine
:
Doctors cannot handle millions of numbers!
How about computers?
Example project topic #1 (3/3)
Let’s Build a Prediction Model
This is a pure machine learning problem!
Transfer learning,
Feature reconstruction
~100 patients at UWMC
30,000 genes
…
g1
g2
g4
RNA levels
of genes in
cancer cells
g7
g10
g13
g3
g5
g6
g15
g16
g30,000
g
g9
g14
g
g3
g
g
30,000
features!
g
(feature
selection)
160 drugs
Drug i
Drug 2
Drug
sensitivity
test
Publicly available
RNA level data
g
g11
g
g
e8
g12
>3000 patients
Patient X
Drug 3
Drug 4
Drug 6
Drug 5
Goal: realizing
personalized
cancer treatment
Prior knowledge
on drugs’ targets
Drug 160
In collaboration with Tony Blau, Pam Becker, Ray Monnat, David Hawkins (Medicine)
Example project topic #2 (1/2)
How Well Can We Predict Diseaserelated Traits Based on DNA?
DNA sequence
Athin, T fat
Nresearch
instances
One of the most important
Individual
A
…ACCCGGTAGACCTTTATTCGGCCCGG…
…ACCCGGTAGACCTTTATTCGGCCCGG…
problems
in this areaIndividual
is to develop new
environmental factors
A
…ACCCGGTAGACCTTAATTCGGCCGGG…
…ACCCGGTAGACCTTAATTCGGCCGGG…
Individual
computational
methods
that
can
:
represent
more complicated
interaction
Individual
T
…ACCCGGTAGTCCTATATTCGGCCCGG…
…ACCCGGTAGTCCTATATTCGGCCCGG…
cell,
aand
complex
system
between
sequence variation
trait.
Individual
T
…ACTCGGTAGTCCTATATTCGGCCGGG…
…ACTCGGTAGTCCTATATTCGGCCGGG…
…ACTCGGTAGACCTAAATTCGGCCCGG…
A
…ACTCGGTAGACCTAAATTCGGCCCGG…
1
2
3
:
N-1
N
p≈106 !
s1
s2
…
too weak to
be detected
?
Causality?
obesity
?
sp
?
Standard approach
Obesity
Find a simple rule!
Failed to detect the DNA affecting
many important traits.
Example project topic #2 (2/2)
How Well Can We Predict Diseaserelated Traits Based on DNA?
~2000 subjects
…ACTCGGACCTAAATCCCG…
…ACCCGGACCTTAATGCGG…
…ACCCGGACCTATATGCCG…
…ACCCGGACCTTTATGCCG…
:
…ACCCGGACCTTAATGCGG…
…ACCCGGTCCTATATGCCG…
…ACTCGGTCCTTAATGCGG…
…ACTCGGTCCTATATGCGG…
…
s1
Sequence
Information
Year
0
Phenotype
Data
:
Year
25
Phenotype
Data
Longitudinal study
Environmental factors
Age, sex, smoking status
s2
s3
s4
p≈106
sP
!
(feature selection)
Cholesterol
Fatty acid
Glucose
Insulin
Structural
learning
:
Age-specific
genetic influence
Cholesterol
Fatty acid
Glucose
Insulin
In collaboration with Alex Reiner (Epidemiology)
More project topics at the
course website!
Questions?