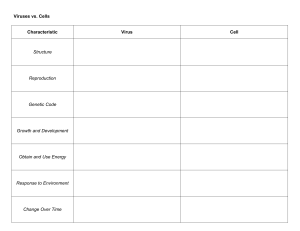
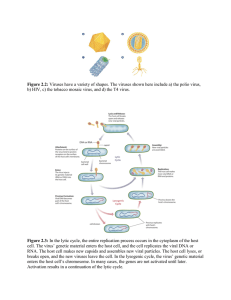
Republic of the Philippines UNIVERSITY OF EASTERN PHILIPPINES University Town, Catarman, Northern Samar COLLEGE OF EDUCATION Secondary Teacher Education Department 2nd Semester SY: 2020-2021 Module in Major 7a: GENETICS This module is prepared by: Christine M. Adlawan, LLB, MPA STEd Faculty Module in GENETICS Module Prof. Christine M. Adlawan 7 VIRUSES OVERVIEW ‗Viruses are everywhere‘ describes the ‗virosphere‘ — the virus biomass of enormous variety and complexity in the environment. Viruses are the most numerous microbes on Earth with an estimated 100 million different types. The largest virus, mimivirus, is thought to fall into an evolutionary position at a point before the animal and plant kingdoms split and indicate the long history of viruses. Humans have been battling viruses since before our species had even evolved into its modern form. For some viral diseases, vaccines and antiviral drugs have allowed us to keep infections from spreading widely, and have helped sick people recover. For one disease — smallpox — we've been able to eradicate it, ridding the world of new cases. This module will discuss everything that we want to know about virus including its prevention and treatment. . LEARNING PLAN At the completion of this lesson, you should be able to: 1. Describe how viruses were first discovered and how they are detected; 2. Identify the general characteristics of viruses as pathogens; 3. Discuss viral genomes; 4. Explain the general characteristics of viral life cycles; 2 Module in GENETICS Prof. Christine M. Adlawan 5. Describe the characteristics used to identify viruses as obligate intracellular parasites; 6. Explain the detailed steps of viral replication; and 7. Describe how vaccines are used in prevention and treatment of viral diseases ACTIVITY Clinical Focus: Joaquim Joaquim, a 45-year-old journalist, has just returned to the U.S. from travels in Russia, China, and Africa. He is not feeling well, so he goes to his general practitioner complaining of weakness in his arms and legs, fever, headache, noticeable agitation, and minor discomfort. He thinks it may be related to a dog bite he suffered while interviewing a Chinese farmer. He is experiencing some prickling and itching sensations at the site of the bite wound, but he tells the doctor that the dog seemed healthy and that he had not been concerned until now. The doctor ordered a culture and sensitivity test to rule out bacterial infection of the wound, and the results came back negative for any possible pathogenic bacteria. In your own understanding, based on this information above, what additional tests should be performed on the patient? What type of treatment should the doctor recommend? 3 Module in GENETICS Prof. Christine M. Adlawan ABSTRACTION A virus can be simply defined as an obligate intracellular parasite. Each viral particle, or virion, consists of a single nucleic acid, RNA or DNA, encoding the viral genome surrounded by a protein coat, and is capable of replication only within the living cells of bacteria, animals or plants. Viruses are classified into different orders and families by consideration of the type of nucleic acid present (RNA or DNA), whether the nucleic acid is single- or double-stranded, and the presence or absence of an envelope. History of Viruses 4 Module in GENETICS Prof. Christine M. Adlawan No one knows exactly when viruses emerged or from where they came, since viruses do not leave historical footprints such as fossils. Modern viruses are thought to be a mosaic of bits and pieces of nucleic acids picked up from various sources along their respective evolutionary paths. Viruses are acellular, parasitic entities that are not classified within any domain because they are not considered alive. They have no plasma membrane, internal organelles, or metabolic processes, and they do not divide. Instead, they infect a host cell and use the host‘s replication processes to produce progeny virus particles. Viruses infect all forms of organisms including bacteria, archaea, fungi, plants, and animals. Living things grow, metabolize, and reproduce. Viruses replicate, but to do so, they are entirely dependent on their host cells. They do not metabolize or grow, but are assembled in their mature form. Viruses are diverse. They vary in their structure, their replication methods, and in their target hosts or even host cells. While most biological diversity can be understood through evolutionary history, such as how species have adapted to conditions and environments, much about virus origins and evolution remains unknown. Despite their small size, which prevented them from being seen with light microscopes, the discovery of a filterable component smaller than a bacterium that causes tobacco mosaic disease (TMD) dates back to 1892? At that time, Dmitri Ivanovski, a Russian botanist, discovered the source of TMD by using a porcelain filtering device first invented by Charles Chamberland and Louis Pasteur in Paris in 1884. Porcelain Chamberland filters has a pore size of 0.1 µm, which is small enough to remove all bacteria ≥0.2 µm from any liquids passed through the device. An extract obtained from TMD-infected tobacco plants was made to determine the cause of the disease. Initially, the source of the disease was thought to be bacterial. It was 5 Module in GENETICS Prof. Christine M. Adlawan surprising to everyone when Ivanovski, using a Chamberland filter, found that the cause of TMD was not removed after passing the extract through the porcelain filter. So if a bacterium was not the cause of TMD, what could be causing the disease? Ivanovski concluded the cause of TMD must be an extremely small bacterium or bacterial spore. Other scientists, including Martinus Beijerinck, continued investigating the cause of TMD. It was Beijerinck, in 1899, who eventually concluded the causative agent was not a bacterium but, instead, possibly a chemical, like a biological poison we would describe today as a toxin. As a result, the word virus, Latin for poison, was used to describe the cause of TMD a few years after Ivanovski‘s initial discovery. Even though he was not able to see the virus that caused TMD, and did not realize the cause was not a bacterium, Ivanovski is credited as the original discoverer of viruses and a founder of the field of virology. Today, we can see viruses using electron microscopes and we know much more about them. Viruses are distinct biological entities; however, their evolutionary origin is still a matter of speculation. In terms of taxonomy, they are not included in the tree of life because they are acellular (not consisting of cells). In order to survive and reproduce, viruses must infect a cellular host, making them obligate intracellular parasites. The genome of a virus enters a host cell and directs the production of the viral components, proteins and nucleic acids, needed to form new virus particles called virions. New virions are made in the host cell by assembly of viral components. The new virions transport the viral genome to another host cell to carry out another round of infection. 6 Module in GENETICS Prof. Christine M. Adlawan Figure 1: (a) Tobacco mosaic virus (TMV) viewed with transmission electron microscope. (b) Plants infected with tobacco mosaic disease (TMD), caused by TMV. (credit a: modification of work by USDA Agricultural Research Service—scale-bar data from Matt Russell; credit b: modification of work by USDA Forest Service, Department of Plant Pathology Archive North Carolina State University) Characteristics of Viruses Infectious, acellular pathogens; Obligate intracellular parasites with host and cell-type specificity; DNA or RNA genome (never both); Genome is surrounded by a protein capsid and, in some cases, a phospholipid membrane studded with viral glycoproteins; Lack genes products successful for many needed for reproduction, requiring exploitation of hostcell genomes to reproduce 7 Module in GENETICS Prof. Christine M. Adlawan Hosts and Viral Transmission Viruses can infect every type of host cell, including those of plants, animals, fungi, protists, bacteria, and archaea. Most viruses will only be able to infect the cells of one or a few species of organism. This is called the host range. However, having a wide host range is not common and viruses will typically only infect specific hosts and only specific cell types within those hosts. The viruses that infect bacteria are called bacteriophages, or simply phages. The word phage comes from the Greek word for devour. Other viruses are just identified by their host group, such as animal or plant viruses. Once a cell is infected, the effects of the virus can vary depending on the type of virus. Viruses may cause abnormal growth of the cell or cell death, alter the cell‘s genome, or cause little noticeable effect in the cell. Viruses can be transmitted through direct contact, indirect contact with fomites, or through a vector: an animal that transmits a pathogen from one host to another. In humans, a wide variety of viruses are capable of causing various infections and diseases. Some of the deadliest emerging pathogens in humans are viruses, yet we have few treatments or drugs to deal with viral infections, making them difficult to eradicate. Viruses that can be transmitted from an animal host to a human host can cause zoonoses. For example, the avian influenza virus originates in birds, but can cause disease in humans. Reverse zoonoses are caused by infection of an animal by a virus that originated in a human. 8 Module in GENETICS Prof. Christine M. Adlawan Viral Structures In general, virions (viral particles) are small and cannot be observed using a regular light microscope. They are much smaller than prokaryotic and eukaryotic cells; this is an adaptation allowing viruses to infect these larger cells (see Figure 2). The size of a virion can range from 20 nm for small viruses up to 900 nm for typical, large viruses (see Figure 3). Recent discoveries, however, have identified new giant viral species, such as Pandoravirus salinus and Pithovirus sibericum, with sizes approaching that of a bacterial cell. Figure 2: (a) In this transmission electron micrograph, a bacteriophage (a virus that infects bacteria) is dwarfed by the bacterial cell it infects. (b) An illustration of the bacteriophage in the micrograph. (credit a: modification of work by U.S. Department of Energy, Office of Science, LBL, PBD) 9 Module in GENETICS Prof. Christine M. Adlawan Figure 3. The size of a virus is small relative to the size of most bacterial and eukaryotic cells and their organelles. In 1935, after the development of the electron microscope, Wendell Stanley was the first scientist to crystallize the structure of the tobacco mosaic virus and discovered that it is composed of RNA and protein. In 1943, he isolated Influenza B virus, which contributed to the development of an influenza (flu) vaccine. Stanley‘s discoveries unlocked the mystery of the nature of viruses that had been puzzling scientists for over 40 years and his contributions to the field of virology led to him being awarded the Nobel Prize in 1946. As a result of continuing research into the nature of viruses, we now know they consist of a nucleic acid (either RNA or DNA, but never both) surrounded by a protein coat called a capsid (see Figure 4). The interior of 10 Module in GENETICS Prof. Christine M. Adlawan the capsid is not filled with cytosol, as in a cell, but instead it contains the bare necessities in terms of genome and enzymes needed to direct the synthesis of new virions. Each capsid is composed of protein subunits called capsomeres made of one or more different types of capsomere proteins that interlock to form the closely packed capsid. Figure 4. Click for a larger image. (a) The naked atadenovirus uses spikes made of glycoproteins from its capsid to bind to host cells. (b) The enveloped human immunodeficiency virus uses spikes made of glycoproteins embedded in its envelope to bind to host cells (credit a ―micrograph‖: modification of work by NIAID; credit b ―micrograph‖: modification of work by Centers for Disease Control and Prevention) 11 Module in GENETICS Prof. Christine M. Adlawan There are two categories of viruses based on general composition. Viruses formed from only a nucleic acid and capsid are called naked viruses or nonenveloped viruses. Viruses formed with a nucleic-acid packed capsid surrounded by a lipid layer are called enveloped viruses (see Figure 4). The viral envelope is a small portion of phospholipid membrane obtained as the virion buds from a host cell. The viral envelope may either be intracellular or cytoplasmic in origin. Extending outward and away from the capsid on some naked viruses and enveloped viruses are protein structures called spikes. At the tips of these spikes are structures that allow the virus to attach and enter a cell, like the influenza virus hemagglutinin spikes (H) or enzymes like the neuraminidase (N) influenza virus spikes that allow the virus to detach from the cell surface during release of new virions. Influenza viruses are often identified by their H and N spikes. For example, H1N1 influenza viruses were responsible for the pandemics in 1918 and 2009, H2N2 for the pandemic in 1957, and H3N2 for the pandemic in 1968. 12 Module in GENETICS Prof. Christine M. Adlawan Viruses vary in the shape of their capsids, which can be helical, polyhedral, or complex. A helical capsid forms the shape of tobacco mosaic virus (TMV), a naked helical virus, and Ebola virus, an enveloped helical virus. The capsid is cylindrical or rod shaped, with the genome fitting just inside the length of the capsid. Polyhedral capsids form the shapes of poliovirus and rhinovirus, and consist of a nucleic acid surrounded by a polyhedral (many-sided) capsid in the form of an icosahedron. An icosahedral capsid is a three-dimensional, 20-sided structure with 12 vertices. These capsids somewhat resemble a soccer ball. Both helical and polyhedral viruses can have envelopes. Viral shapes seen in certain types of bacteriophages, such as T4 phage, and poxviruses, like vaccinia virus, may have features of both polyhedral and helical viruses so they are described as a complex viral shape (see Figure 5). In the bacteriophage complex form, the genome is located within the polyhedral head and the sheath connects the head to the tail fibers and tail pins that help the virus attach to receptors on the host cell‘s surface. Poxviruses that have complex shapes are often brick shaped, with intricate surface characteristics not seen in the other categories of capsid. 13 Module in GENETICS Prof. Christine M. Adlawan Figure 5. Viral capsids can be (a) helical, (b) polyhedral, or (c) have a complex shape. (credit a ―micrograph‖: modification of work by USDA ARS; credit b ―micrograph‖: modification of work by U.S. Department of Energy) Classification and Taxonomy of Viruses Although viruses are not classified in the three domains of life, their numbers are great enough to require classification. Since 1971, the International Union of Microbiological Societies Virology Division has given the task of developing, refining, and maintaining universal virus taxonomy to the International Committee on Taxonomy of Viruses (ICTV). Since viruses 14 Module in GENETICS Prof. Christine M. Adlawan can mutate so quickly, it can be difficult to classify them into a genus and a species epithet using the binomial nomenclature system. Thus, the ICTV‘s viral nomenclature system classifies viruses into families and genera based on viral genetics, chemistry, morphology, and mechanism of multiplication. To date, the ICTV has classified known viruses in seven orders, 96 families, and 350 genera. Viral family names end in –viridae (e.g, Parvoviridae) and genus names end in −virus (e.g., Parvovirus). The names of viral orders, families, and genera are all italicized. When referring to a viral species, we often use a genus and species epithet such as Pandoravirus dulcis or Pandoravirus salinus. The Baltimore classification system is an alternative to ICTV nomenclature. The Baltimore system classifies viruses according to their genomes (DNA or RNA, single versus double stranded, and mode of replication). This system thus creates seven groups of viruses that have common genetics and biology. Aside from formal systems of nomenclature, viruses are often informally grouped into categories based on chemistry, morphology, or other characteristics they share in common. Categories may include naked or enveloped structure, single-stranded (ss) or double-stranded (ds) DNA or ss or ds RNA genomes, segmented or nonsegmented genomes, and positivestrand (+) or negative-strand (−) RNA. For example, herpes viruses can be classified as a dsDNA enveloped virus; human immunodeficiency virus (HIV) is a +ssRNA enveloped virus, and tobacco mosaic virus is a +ssRNA virus. Other characteristics such as host specificity, tissue specificity, capsid shape, and special genes or enzymes may also be used to describe groups of similar viruses. 15 Module in GENETICS Prof. Christine M. Adlawan The Table below lists some of the most common viruses that are human pathogens by genome type. Common Pathogenic Viruses Genome dsDNA, enveloped dsDNA, naked ssDNA, naked Family Poxviridae Example Virus Orthopoxvirus Poxviridae Herpesviridae Parapoxvirus Simplexvirus Adenoviridae Atadenovirus Papillomaviridae Papillomavirus Reoviridae Reovirus Parvoviridae Reoviridae Adeno-associated dependoparvovirus A Adeno-associated dependoparvovirus B Rotavirus Picornaviridae Picornaviridae Enterovirus C Rhinovirus Picornaviridae Togaviridae Hepatovirus Alphavirus Togaviridae Retroviridae Rubivirus Lentivirus Parvoviridae dsRNA, naked +ssRNA, naked +ssRNA, enveloped −ssRNA, enveloped Filoviridae Zaire Ebolavirus Orthomyxoviridae Influenzavirus A, B, C Rhabdoviridae Lyssavirus Clinical Features Skin papules, pustules, lesions Skin lesions Cold sores, genital herpes, sexually transmitted disease Respiratory infection (common cold) Genital warts, cervical, vulvar, or vaginal cancer Gastroenteritis severe diarrhea (stomach flu) Respiratory tract infection Respiratory tract infection Gastroenteritis Poliomyelitis Upper respiratory tract infection (common cold) Hepatitis Encephalitis, hemorrhagic fever Rubella Acquired immune deficiency syndrome (AIDS) Hemorrhagic fever Flu Rabies 16 Module in GENETICS Prof. Christine M. Adlawan How Viruses Replicate Viral replication is the term used that indicates the formation of biological viruses during the infection process in the target host cells. Viruses must first penetrate and enter the cell before viral replication can occur. From the perspective of the virus, the purpose of viral replication is to allow reproduction and survival of its kind. By generating abundant copies of its genome and packaging these copies into viruses, the virus is able to continue infecting new hosts. Replication between viruses is varied and depends on the type of genes involved. Most DNA viruses assemble in the nucleus; most RNA viruses develop solely in cytoplasm. Viral populations do not grow through cell division, because they are acellular. Instead, they hijack the machinery and metabolism of a host cell to produce multiple copies of themselves, and they assemble inside the cell. The life cycle of viruses differs greatly between species but there are six (6) common basic stages: 1. Attachment is a specific binding between viral capsid proteins and specific receptors on the host cellular surface. This specificity determines the host range of a virus. For example, HIV can infect only a limited range of human leukocytes. Its surface protein, gp120, specifically interacts only with the CD4 molecule – a chemokine receptor – which is most commonly found on the surface of CD4+ T-Cells. This mechanism has evolved to favor those viruses that infect only cells within which they are capable of replication. Attachment to the receptor can fore the viral envelope protein to undergo either changes that 17 Module in GENETICS Prof. Christine M. Adlawan result in the fusion of viral and cellular membranes, or changes of nonenveloped virus surface proteins that allow the virus to enter. 2. Penetration follows attachment. Virions enter the host cell through receptor-mediated endocytosis or membrane fusion. This is often called viral entry. The infection of plant and fungal cells is different from that of animal cells. Plants have a rigid cell wall made of cellulose, and fungi one of chitin, so most viruses can get inside these cells only after trauma to the cell wall. However, nearly all plant viruses (such as tobacco mosaic virus) can also move directly from cell to cell, in the form of single-stranded nucleoprotein complexes, through pores called plasmodesmata. Bacteria, like plants, have strong cell walls that a virus must breach to infect the cell. However, since bacterial cell walls are much less thick than plant cell walls due to their much smaller size, some viruses have evolved mechanisms that inject their genome into the bacterial cell across the cell wall, while the viral capsid remains outside. 3. Uncoating is a process in which the viral capsid is removed: This may be by degradation by viral or host enzymes or by simple dissociation. In either case the end-result is the release of the viral genomic nucleic acid. 4. Replication - Replication of viruses depends on the multiplication of the genome. This is accomplished through synthesis of viral messenger RNA (mRNA) from ―early‖ genes (with exceptions for positive sense RNA viruses), viral protein synthesis, possible assembly of viral proteins, then viral genome replication mediated by early or regulatory protein expression. This may be followed, for complex viruses with larger genomes, by one or more further rounds of mRNA synthesis: ―late‖ gene expression is, in general, of structural or virion proteins. 18 Module in GENETICS Prof. Christine M. Adlawan Hepatitis C virus: A simplified diagram of the Hepatitis C virus replication cycle. 5. Assembly - Following the structure-mediated self-assembly of the virus particles, some modification of the proteins often occurs. In viruses such as HIV, this modification (sometimes called maturation) occurs after the virus has been released from the host cell. 6. Release - Viruses can be released from the host cell by lysis, a process that kills the cell by bursting its membrane and cell wall if present. This is a feature of many bacterial and some animal viruses. Some viruses undergo a lysogenic cycle where the viral genome is incorporated by genetic recombination into a specific place in the host‘s chromosome. The viral genome is then known as a provirus or, in the case of bacteriophages a prophage. Whenever the host divides, the viral genome is also replicated. 19 Module in GENETICS Prof. Christine M. Adlawan The viral genome is mostly silent within the host; however, at some point the provirus or prophage may give rise to active virus, which may lyse the host cells. Enveloped viruses (e.g., HIV) typically are released from the host cell by budding. During this process the virus acquires its envelope, which is a modified piece of the host‘s plasma or other internal membrane. The genetic material within virus particles, and the method by which the material is replicated, varies considerably between different types of viruses. COVID-19 and inhibiting viral replication COVID-19 has become a recent focus of viral replication studies. So far, over 300 human proteins have been found to interact with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) during infection. It is believed that by blocking certain interactions between human and viral proteins, the viral replication can be stopped, thereby stopping transmission. Similar therapeutic methods have been suggested for other diseases, such as HIV. When similar studies on interactions between human and viral proteins were done for HIV, they took a few years. Despite this, the interaction mapping for COVID-19 is progressing much quicker and many existing drugs that may be useful in stopping or slowing COVID-19 have been identified. However, many of these drugs have now been found to be ineffective. There is some opposition to this method. Drugs targeting human protein function, which would then stop viral replication, might also produce side effects that are highly deleterious to already vulnerable patients. Some short-term drugs are seen as more applicable for COVID-19, but the wisdom and efficacy of this are still disputed. 20 Module in GENETICS Prof. Christine M. Adlawan Steps of Virus Infections A virus must ―take over‖ a cell to replicate. The viral replication cycle can produce dramatic biochemical and structural changes in the host cell, which may cause cell damage. These changes, called cytopathic effects, can change cell functions or even destroy the cell. Some infected cells, such as those infected by the common cold virus (rhinovirus), die through lysis (bursting) or apoptosis (programmed cell death or ―cell suicide‖), releasing all the progeny virions at once. The symptoms of viral diseases result from the immune response to the virus, which attempts to control and eliminate the virus from the body, and from cell damage caused by the virus. Many animal viruses, such as HIV (human immunodeficiency virus), leave the infected cells of the immune system by a process known as budding, where virions leave the cell individually. During the budding process, the cell does not undergo lysis and is not immediately killed. However, the damage to the cells that HIV infects may make it impossible for the cells to function as mediators of immunity, even though the cells remain alive for a period of time. Most productive viral infections follow similar steps in the virus replication cycle: attachment, penetration, uncoating, replication, assembly, and release. Attachment A virus attaches to a specific receptor site on the host cell membrane through attachment proteins in the capsid or via glycoproteins embedded in the viral envelope. The specificity of this interaction determines the host (and the cells within the host) that can be infected by a particular virus. This can be illustrated by thinking of several keys and several locks where each key will fit only one specific lock. 21 Module in GENETICS Prof. Christine M. Adlawan Entry The nucleic acid of bacteriophages enters the host cell naked, leaving the capsid outside the cell. Plant and animal viruses can enter through endocytosis, in which the cell membrane surrounds and engulfs the entire virus. Some enveloped viruses enter the cell when the viral envelope fuses directly with the cell membrane. Once inside the cell, the viral capsid is degraded and the viral nucleic acid is released, which then becomes available for replication and transcription. Pathway to viral infection: In influenza virus infection, glycoproteins attach to a host epithelial cell. As a result, the virus is engulfed. RNA and proteins are made and assembled into new virions. 22 Module in GENETICS Prof. Christine M. Adlawan Replication and Assembly The replication mechanism depends on the viral genome. DNA viruses usually use host cell proteins and enzymes to make additional DNA that is transcribed to messenger RNA (mRNA), which is then used to direct protein synthesis. RNA viruses usually use the RNA core as a template for synthesis of viral genomic RNA and mRNA. The viral mRNA directs the host cell to synthesize viral enzymes and capsid proteins, and to assemble new virions. Of course, there are exceptions to this pattern. If a host cell does not provide the enzymes necessary for viral replication, viral genes supply the information to direct synthesis of the missing proteins. Retroviruses, such as HIV, have an RNA genome that must be reverse transcribed into DNA, which then is incorporated into the host cell genome. To convert RNA into DNA, retroviruses must contain genes that encode the virus-specific enzyme reverse transcriptase, which transcribes an RNA template to DNA. Reverse transcription never occurs in uninfected host cells; the needed enzyme, reverse transcriptase, is only derived from the expression of viral genes within the infected host cells. The fact that HIV produces some of its own enzymes not found in the host has allowed researchers to develop drugs that inhibit these enzymes. These drugs, including the reverse transcriptase inhibitor AZT, inhibit HIV replication by reducing the activity of the enzyme without affecting the host‘s metabolism. This approach has led to the development of a variety of drugs used to treat HIV and has been effective at reducing the number of infectious virions (copies of viral RNA) in the blood to non-detectable levels in many HIVinfected individuals. 23 Module in GENETICS Prof. Christine M. Adlawan Egrees The last stage of viral replication is the release of the new virions into the host organism, where they are able to infect adjacent cells and repeat the replication cycle. Some viruses are released when the host cell dies and other viruses can leave infected cells by budding through the membrane without directly killing the cell. Vaccines for Prevention While we do have limited numbers of effective antiviral drugs, such as those used to treat HIV and influenza, the primary method of controlling viral disease is by vaccination, which is intended to prevent outbreaks by building immunity to a virus or virus family. A vaccine may be prepared using weakened live viruses, killed viruses, or molecular subunits of the virus. In general, live viruses lead to better immunity, but have the possibility of causing disease at some low frequency. Killed viral vaccine and the subunit viruses are both incapable of causing disease, but in general lead to less effective or long-lasting immunity. Weakened live viral vaccines are designed in the laboratory to cause few symptoms in recipients while giving them immunity against future infections. Polio was one disease that represented a milestone in the use of vaccines. Mass immunization campaigns in the U.S. in the 1950s (killed vaccine) and 1960s (live vaccine) essentially eradicated the disease, which caused muscle paralysis in children and generated fear in the general population when regional epidemics occurred. The success of the polio vaccine paved the way for the routine dispensation of childhood vaccines against measles, mumps, rubella, chickenpox, and other diseases. 24 Module in GENETICS Prof. Christine M. Adlawan Live vaccines are usually made by attenuation (weakening) of the ―wild-type‖ (disease-causing) virus by growing it in the laboratory in tissues or at temperatures different from what the virus is accustomed to in the host. For example, the virus may be grown in cells in a test tube, in bird embryos, or in live animals. The adaptation to these new cells or temperature induces mutations in the virus‘ genomes, allowing them to grow better in the laboratory while inhibiting their ability to cause disease when reintroduced into the conditions found in the host. These attenuated viruses thus still cause an infection, but they do not grow very well, allowing the immune response to develop in time to prevent major disease. The danger of using live vaccines, which are usually more effective than killed vaccines, is the low but significant risk that these viruses will revert back to their disease-causing form by back mutations. Back mutations occur when the vaccine undergoes mutations in the host such that it readapts to the host and can again cause disease, which can then be spread to other humans in an epidemic. This happened as recently as 2007 in Nigeria where mutations in a polio vaccine led to an epidemic of polio in that country. Some vaccines are in continuous development because certain viruses, such as influenza and HIV, have a high mutation rate compared to other viruses or host cells. With influenza, mutation in genes for the surface molecules helps the virus evade the protective immunity that may have been obtained in a previous influenza season, making it necessary for individuals to get vaccinated every year. Other viruses, such as those that cause the childhood diseases measles, mumps, and rubella, mutate so little that the same vaccine is used year after year. 25 Module in GENETICS Prof. Christine M. Adlawan SUMMARY Viruses are acellular entities that can usually only be seen with an electron microscope. Their genomes contain either DNA or RNA, and they replicate using the replication proteins of a host cell. Viruses are diverse, infecting archaea, bacteria, fungi, plants, and animals. Viruses consist of a nucleic-acid core surrounded by a protein capsid with or without an outer lipid envelope. Viral replication within a living cell always produces changes in the cell, sometimes resulting in cell death and sometimes slowly killing the infected cells. There are six basic stages in the virus replication cycle: attachment, penetration, uncoating, replication, assembly, and release. A viral infection may be productive, resulting in new virions, or nonproductive, meaning the virus remains inside the cell without producing new virions. Viruses cause a variety of diseases in humans. Many of these diseases can be prevented by the use of viral vaccines, which stimulate protective immunity against the virus without causing major disease. Viral vaccines may also be used in active viral infections, boosting the ability of the immune system to control or destroy the virus. Antiviral drugs that target enzymes and other protein products of viral genes have been developed and used with mixed success. Combinations of anti-HIV drugs have been used to effectively control the virus, extending the lifespan of infected individuals. 26 Module in GENETICS Prof. Christine M. Adlawan APPLICATION Name each labeled part of the illustrated bacteriophage. ASSESSMENT 1. Discuss the geometric differences among helical, polyhedral, and complex viruses. 2. What was the meaning of the word ―virus‖ in the 1880s and why was it used to describe the cause of tobacco mosaic disease? 3. In terms of evolution, which do you think arises first? The virus or the host? Explain your answer. 27 Module in GENETICS Prof. Christine M. Adlawan 4. Do you think it is possible to create a virus in the lab? Imagine that you are a mad scientist. Describe how you would go about creating a new virus? FEEDBACK Do you have any question relative to our topic? Write them below. ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ___________________________________________________________ REFERENCES Goulding, J., 2020. Virus Replication. [online] Immunology.org. Available at: <https://www.immunology.org/public- information/bitesized-immunology/pathogens-and-disease/virusreplication>. H. Lecoq. "[Discovery of the First Virus, the Tobacco Mosaic Virus: 1892 or 1898?]." Comptes Rendus de l‘Academie des Sciences – Serie III – Sciences de la Vie 324, no. 10 (2001): 929– 933. 28 Module in GENETICS Prof. Christine M. Adlawan US Department of Health and Human Services, Centers for Disease Control and Prevention. "Antibiotic Resistance Threats in the United States, 2013." http://www.cdc.gov/drugresistance/pdf/ar-threats-2013-508.pdf (accessed September 22, 2015). M. Clokie et al. "Phages in Nature." Bacteriophage 1, no. 1 (2011): 31–45. ↵ Marks, R., 2020. Unveiling How Coronavirus Hijacks Our Cells to Help Rush New Drugs to Patients. [online] UCSF.edu. Available at: <https://www.ucsf.edu/news/2020/03/416986/unveiling-how- coronavirus-hijacks-our-cells-help-rush-new-drugs-patients>. US Food and Drug Administration. "FDA Approval of Listeriaspecific Bacteriophage Preparation on Ready-to-Eat (RTE) Meat and Poultry Products." Reece, JB., Campbell, N.A, Urry, L.A, et.al., Campbell Biology. 11th edition. Wang, W., Wu, C., Amarasinghe, G., and Leung, D., 2019. Ebola Virus Replication Stands Out. Trends in Microbiology, 27(7), pp. 565-566. 29