12596019_Visuals.ppt (1.913Mb)
advertisement

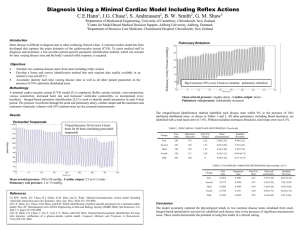
Simulating Cardiac Disease From Onset with a Minimal Cardiac Model Including Reflex Actions THE 12th INTERNATIONAL CONFERENCE ON BIOMEDICAL ENGINEERING C. E. Hann1, J. G. Chase1, S. Andreassen2, B.W. Smith2, G. M. Shaw3, 1Department of Mechanical Engineering, University of Canterbury, Christchurch, New Zealand for Model-based Medical Decision Support, Aalborg University, Aalborg, Denmark 3 Department of Intensive Care Medicine, Christchurch Hospital, Christchurch, New Zealand 2Centre Diagnosis and Treatment • Cardiac disease state difficult to diagnose - Limited data - Reflex actions • Minimal Cardiac Model - primary parameters - common ICU measurements • Increased resistance in pulmonary artery – pulmonary embolism, atherosclerotic heart disease • Optimize drug treatment on computer -e.g. Warfarin • Require fast parameter ID Heart Model D.E.’s and PV diagram V Q1 Q2 P P Q1 R1 Q1 1 2 L1 P P3 Q2 R2 Q 2 2 L2 P2 e(t ) Ees (V Vd ) (1 e(t )) P0 (e (V V0 ) 1), e(t ) e 80( t 0.375) 2 Reflex actions • • • • Vaso-constriction - contract veins Venous constriction – increase venous dead space Increased HR Increased ventricular contractility Varying HR as a function of Pao Disease States • Pericardial Tamponade - build up of fluid in pericardium - dead space volume V0,pcd by 10 ml / 10 heart beats • Pulmonary Embolism - Rpul 20% each time • Cardiogenic shock - e.g. left ventricle infarction, blocked coronary artery - not enough oxygen to myocardium - Ees,lvf, P0,lvf • Septic shock - blood poisoning - reduce systemic resistance • Hypovolemic shock – severe drop in total blood volume Results • Healthy human Output Value Volume in left ventricle 111.7/45.7 ml Volume in right ventricle 112.2/46.1 ml Cardiac output 5.3 L/min Max Plv 119.2 mmHg Max Prv 26.2 mmHg Pressure in aorta 116.6/79.1 mmHg Pressure in pulmonary artery 25.7/7.8 mmHg Avg pressure in pulmonary vein 2.0 mmHg Avg pressure in vena cava 2.0 mmHg Results • • Pericardial tamponade • Pulmonary Embolism Ppu – 7.9 mmHg CO – 4.1 L/min MAP – 88.0 mmHg All other disease states capture physiological trends and magnitudes Identifiability • Add 10% noise to outputs • Apply integral-based optimization Integral Method - Concept x ax b sin( t ) c, x (0) 1 a 0.5, b 0.2, c 0.8 • Discretised solution analogous to measured data (simple example with analytical solution ) • Work backwards and find a,b,c • Current method – solve D. E. numerically or analytically 1.85 x (t ) 1.8 (ab cos t ba 2 sin t ca 2 c )) 1.75 1.7 - Find best least squares fit of x(t) to the data 1.65 x - Non-linear, non-convex optimization, computationally intense 1.6 q 1.55 1.5 P1 P2 R • integral method – reformulate in terms of integrals 1.45 1.4 12 1 ( eat ( a c ab ca 2 a 3 ) (a 1)a 2 – linear, convex optimization, minimal computation 13 14 15 16 time 17 18 19 Integral Method - Concept Integrate x ax b sin( t ) c, both sides from t0 to t ( t0 4 ) • t x dt t (ax b sin( t ) c) dt t t 0 0 x(t ) x(t0 ) a t x dt b t sin( t ) dt c t 1 dt • t t t 0 0 0 x(t ) x(t0 ) a t x dt b(cos( t0 ) cos(t )) c(t t0 ) t 0 Choose 10 values of t, between t0 4 and 6 form 10 equations in 3 unknowns a,b,c a tt x dt b(1 cos( ti )) c(ti t0 ) x (ti ) x (t0 ), i 1,,10 0 Integral Method - Concept tt x dt cos( t0 ) cos( t1 ) t1 t0 a x (t1 ) x (t0 ) b t x dt cos( t ) cos( t ) t t c x ( t ) x ( t ) 0 10 10 0 0 10 t 1 0 10 0 • Linear least squares (unique solution) Method Starting point CPU time (seconds) Solution Integral - 0.003 [-0.5002, -0.2000, 0.8003] Non-linear [-1, 1, 1] 4.6 [-0.52, -0.20, 0.83] Non-linear [1, 1, 1] 20.8 [0.75, 0.32, -0.91] • Integral method is at least 1000-10,000 times faster depending on starting point • Thus very suitable for clinical application Identifiability • • Pericardial tamponade (determining V0,pcd) Change True value (ml) Optimized Value Error (%) First 180 176 2.22 Second 160 158 1.25 Third 140 138 1.43 Fourth 120 117 2.50 Fifth 100 100 0 Pulmonary Embolism (determining Rpul) Change True value (mmHg s ml-1) Optimized Value Error (%) First 0.1862 0.1907 2.41 Second 0.2173 0.2050 5.67 Third 0.2483 0.2694 8.50 Fourth 0.2794 0.2721 2.60 Fifth 0.3104 0.2962 4.59 Identifiability • • Cardiogenic shock (determining [Ees,lvf, P0,lvf] (mmHg ml-1, mmHg) Change True values Optimized Value Error (%) First [2.59,0.16] [2.61,0.15] [0.89,5.49] Second [2.30,0.19] [2.30,0.18] [0.34,4.39] Third [2.02,0.23] [2.02,0.21] [0.43,8.03] Fourth [1.73,0.26] [1.70,0.24] [1.48,9.85] Fifth [1.44,0.30] [1.43,0.27] [0.47,9.39] Septic Shock (determining Rsys) Change True value (mmHg s ml-1) Optimized Value Error (%) First 1.0236 1.0278 0.41 Second 0.9582 0.9714 1.37 Third 0.8929 0.8596 3.73 Fourth 0.8276 0.8316 0.49 Fifth 0.7622 0.7993 4.86 Identifiability • Hypovolemic Shock (determining stressed blood volume) Change True value (ml) Optimized Value Error (%) First 1299.9 1206.5 7.18 Second 1177.3 1103.7 6.26 Third 1063.1 953.8 10.28 Fourth 967.8 1018.9 5.28 Fifth 928.5 853.4 8.10 Conclusions • Minimal cardiac model simulate time varying disease states • Accurately captured physiological trends and magnitudes capture wide range of dynamics • Integral based parameter ID method - errors from 0-10%, with 10% noise - identifiable using common measurements • Rapid feedback to medical staff Acknowledgements Engineers and Docs Dr Geoff Chase Dr Geoff Shaw The Danes Steen Andreassen The honorary Danes Dr Bram Smith Questions ??? AIC2, Kate, Carmen and Nick