HHMI at AVI BioPharma PMOs as Antibiotics: Gene- E. coli
advertisement

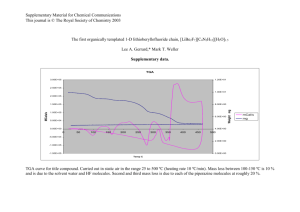
HHMI at AVI BioPharma PMOs as Antibiotics: Genespecific Inhibition of E. coli Mentor: Dr. Bruce Geller Susan Puckett Background Antibiotics today— a biological arms race Some bacteria have developed resistance and immunity to available antibiotics MRSA, TB, Pseudomonas, VRE Overview AVI BioPharma, OSU-LARC Using man-made DNA mimics called phosphorodiamidate morpholino oligomers (PMOs) to block production of essential protein in E. coli PMOs work as antibiotics to inhibit bacterial growth Antisense mechanism: forming complementary base pairs with nucleic acid to change gene expression Escherichia coli bacterium, prokaryote E. coli is a model organism A lot of information available Easy to obtain, work with and grow Safe (few dangerous strains, many harmless unless in large quantities) We are trying to find ways to inhibit the growth of E. coli using DNA mimics (PMOs) Eventually this technology can be used with other disease causing organisms Phosphorodiamidate Morpholino Oligomers (PMOs) DNA mimics Structure is different enough so that nucleases don’t break it down Used to anneal to mRNA in order to block ribosomes from attaching, disrupting protein synthesis PMOs can be manufactured to match RNA to form complementary base pairs in the same manner nucleic acids bind to each other PMO: a picture RNA strand: 5’……….AUG AGC ACU AUC GAA GAA CGC GUU………..3’ PMO: G TGA TAG CTT C 1) A U G A G RNA PMO C A C U A U C G A A G G T G A T A G C T T C C A C U A U C G A A G G T G A T A G C T T C A 2) A U G A G A Protein Synthesis Three Stages Initiation of Translation Elongation Termination In the initiation of translation stage, the ribosome finds the ribosome binding site (RBS) on the mRNA and attaches. PMOs are thought to prevent the two parts of the ribosome from coming together and attaching to the mRNA. PMOs and RNA What size of PMO works best? What gene in E. coli to target? What locus on the gene to target? 11 base pair PMO most effective for inhibiting translation AcpP gene necessary for production of essential protein Targeting just downstream of the ribosome binding site PMOs and Peptides Problem: getting PMO into cell Peptides attached to PMOs increase the amount of compound that enters the cell Question: gene specific or general toxicity? Experiment: showing specificity Wildtype (original) strain of E. coli (W3110) and mutant strain of E. coli in the AcpP gene (LT 1.7) mixed with AcpP PMO, mutant PMO, scrambled PMO, or water. Incubated, recorded optical density (OD600) using microplate reader every hour to measure growth. Optical Densities vs. Time (Growth Curves) 5’….AUG AGC ACU AUC GAA GAA CGC GUU…..3’ G TGA TAG CTT C AcpP mRNA AcpP PMO 5’….AUG AGU ACC AUU GAG GAA CGC GU…..3’ A TGG TAA CTC C AcpP mutant mRNA AcpP PMOmut4 Conclusion of experiment Order of bases matters: sequence specific inhibition PMOs do inhibit bacterial growth Experiment: Minimum Inhibitory Concentration (MIC) How much PMO is needed to prevent growth of E. coli? Experiment tests the ability of different concentrations of peptidePMOs to inhibit strains of E. coli. PMO and bacteria are combined in a 96-well plate, incubated overnight, and examined. Results Various PMOs tested as well as some traditional antibiotics such as tetracycline, rifampin Example: MIC determined to be 2.5 uM (micromolar) for the (RFF)3R-AcpP PMO optical density readings 40 uM 20 uM 10 uM 5 uM 2.5uM 1.25 uM 0.004 0.000 0.001 -0.001 0.575 0.451 0.003 0.001 0.001 0.571 0.000 0.119 0.003 0.015 0.000 0.000 0.000 0.410 Watch for contamination, mutants Isolation of Mutants Growth at the MIC for (RFF)3R-AcpP PMO was used for a second MIC Significantly higher MIC Streak plated Colonies were picked and named W3110R1 through 10 (W3110 was the parent strain) Gram stain tests Sequencing of AcpP target region Frequency of mutation: 1.4 x 10-6 on plates with 20uM PMO Evidence of peptide related mutation No change in sequence of AcpP target Targeting a different gene with the same peptide still has resistance Therefore we think the mutation must be affecting the peptide uptake into the cell This is useful because very little is known about how the peptide gets into the cell. FtsZ Graphs 0 uM FtsZ PMO 08-01-06 FtsZ: another gene in E. coli essential for protein production 1.00E+11 1.00E+10 1.00E+09 CFU/mL 1.00E+08 1.00E+07 1.00E+06 1.00E+05 1.00E+04 1.00E+03 1.00E+02 1.00E+01 160 uM FtsZ PMO 08-01-06 1.00E+00 W3110R1 W3110 parent E. coli 1.00E+11 1.00E+10 1.00E+09 1.00E+08 CFU/mL Colony-forming unit (CFU) per mL: found from plate counting 1.00E+07 1.00E+06 1.00E+05 1.00E+04 1.00E+03 1.00E+02 1.00E+01 1.00E+00 W3110R1 W3110 parent E. coli Minimum Inhibitory Concentrations 45 40 Concentration (µM) 35 30 25 W3110 W3110R 20 15 10 5 0 (RFF)3RAcpP11 D-(RFF)3RAcpP11 (KFF)3KAcpP11 (RXX)3-AcpP PMO (RTR)-AcpP11 (RXR)4AcpP11 (RX)6BAcpP11 Conclusion (RFF)3R-AcpP sequence specific inhibition MIC of (RFF)3R-AcpP is 2.5 uM Mutants can be isolated and mutants appear to be peptide related Future experimentation Identifying mutant gene in resistant strain Other bacteria Testing in vitro and in vivo Acknowledgements Dr. Bruce Geller Luke Tilley Brett Mellbye AVI BioPharma Dr. Kevin Ahern Howard Hughes Medical Institute Publications: Lucas D. Tilley, Brett L. Mellbye, Susan E. Puckett, Patrick. L. Iversen, Bruce L. Geller. 2006. Antisense Peptide-Phosphorodiamidate Morpholino Oligomer Conjugate: Dose-Response in Mice Infected with Escherichia coli. J. Antimicrob. Chemother. In press.