Mitochondrial Disorders & Linkage Analysis
advertisement

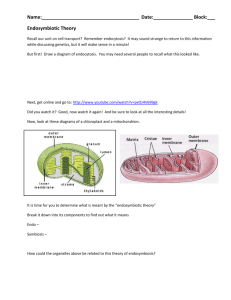
Mitochondrial Disorders Mitochondria – responsible for cellular respiration (the O2-requiring stage of energy production), which takes place via oxidative phosphorylation. Two membranes – outer (MOM) and inner (MIM). o Between membranes = intermembrane space. o Inside MIM = matrix. Oxidative degradation of foodstuffs results in production of simple energy carriers, like NADH and FADH2, which are further metabolized to produce the energy for normal cellular functions. ATP is synthesized via oxidative phosphorylation of ADP in the inner membrane of the mitochondrion. o OxPhos components (5 multi-protein complexes) are embedded in the inner membrane (MIM). o NADH and FADH2 are oxidized by Complexes I and II, then CoQ10 transports electrons (e-) to Complex III. o Complex III transports e- to cytochrome c, which then transfers them to Complex IV where they react with O2. o Proton gradient forms (negative potential inside MIM due to movement of protons into IMS). o Complex V uses energy stored in gradient (proton-motive force) to generate ATP from ADP. Mitochondrial Genome – circular, double-stranded chromosome (mtDNA) that is replicated, transcribed and translated within the mitochondrion. Unique Characteristics: o Uses its own genetic code. o Synthesizes 12S and 16S rRNA. o Exhibits continuous transcription of multiple genes (some polycistronic mRNAs). o Complete lack of introns. o Reminiscent of prokaryotic genomes. Mitochondrial genome encodes for: o 13 polypeptides (in 4 oxphos complexes). o 2 ribosomal RNAs. o 22 transfer RNAs. o **The remainder of proteins that exist in mitochondria (~99% in humans) are transcribed in the nucleus, translated in the cytoplasm and transported into mitochondria. Ploidy of mitochondrial genome: o 2-10 copies of mtDNA per mitochondrion. o Therefore, each cell contains hundreds to thousands of copies of the mitochondrial genome (useful in forensic and anthropologic investigations). Characteristics of mtDNA Affecting Phenotypic Expression 1. mtDNA has a very high mutation rate (7-10 fold higher than nuclear DNA). a. Thus, the mitochondrial genotype can change within a single generation through accumulation of somatic mutations. b. Consequences of mitochondrial mutation rate/accumulation: i. A study was done in which the proof-reading enzyme for mtDNA (mtDNA pol ɣ) was mutated in mice via recombination. ii. Mice homozygous for the mutation in mtDNA pol ɣ exhibited progeria-like symptoms (rapid aging, impaired survival). iii. Phenotype was rescued slightly in mutant mice given endurance training (treadmill running). 1. Prevented mitochondrial depletion/mutations and restored mitochondrial morphology, blunting pathological apoptosis in multiple tissues. 2. Prevented premature mortality and reduced pathology in multiple organs. 2. Both normal (WT) and mutant mtDNA can co-exist within the same cell: a. Homoplasmy – one type of mtDNA per cell. b. Heteroplasmy – multiple types of mtDNA within a cell. i. In normal individuals, ~99.9% of mtDNA molecules are identical. 3. Different tissues can contain different proportions of mutant and normal mtDNA. a. mtDNA replicates autonomously from nuclear DNA. b. Replicative Segregation – mitochondria segregate into daughter cells independently of nuclear DNA. i. Leads to different proportions of mutant and normal mtDNA in different tissues. 4. Tissues differ in dependence upon oxidative phosphorylation: a. Heart, skeletal muscle, brain, and CNS are heavily dependent upon oxphos. b. Other tissues that depend heavily on oxphos include the liver, kidney and insulin-forming islets of the pancreas. c. **Mitochondrial disorders show their clinical manifestations mainly in tissues that are heavily dependent on oxphos. i. Often characterized as myopathies and encephalopathies. 5. Threshold effects in expression of mutant mtDNA --85-90% of mitochondria in a cell may have to be abnormal to discern an aberrant phenotype. Summary: o Clinical phenotype in mitochondrial disease is not simply (or directly) related to mtDNA genotype, rather, it depends on: Inherent oxphos capacity, determined by BOTH nuclear and mitochondrial genes. Accumulation of somatic mtDNA mutations. Degree of heteroplasmy. Tissue-specific requirements for oxphos. Age – progressive mtDNA alterations can lead to worsening conditions with age. Mitochondrial Diseases Maternal Inheritance of mitochondria – non-Mendelian inheritance pattern exclusively through maternal line. o Sperm contain few mitochondria, and none survive within fertilized egg, thus all mitochondria in embryo are maternal in origin (from oocyte cytoplasm). o Transmission solely by the mother to BOTH sons and daughters (sex bias in transmission). o Equal numbers of males and females are affected (no sex bias in affected progeny). o A form of uniparental transmission. Diseases caused by: o Deletions and other structural rearrangements. Mitochondrial Myopathy (MM) and Kearns-Sayre (KS) Neither size nor position of deletions are well correlated with either enzyme deficiency or severity of disease. KS is a multi-system disorder characterized by brain and muscle dysfunction, progressive external ophthalmoplegia (muscle paralysis), and pigmentary retinopathy. o Patients often short of stature, have hearing loss, mental retardation or dementia, and endocrinopathy. Diabetes and deafness sometimes associated with mtDNA deletions or duplications in pedigrees showing maternal inheritance. o Point mutations in protein-encoding genes and tRNA-encoding genes. Leber’s Hereditary Optic Neuropathy (LHON) is a rare neurodegenerative disease of young adults that results in blindness (optic nerve degradation). Caused by a point mutation in NADH dehydrogenase subunit 4. Myoclonic Epilepsy with Ragged Red Fiber (MERRF) is caused by a mutation in tRNALys. Mitochondrial Encephalopathy, Lactic Acidosis, and Stroke-like symptoms (MELAS) is caused by a mutation in tRNALeu. Maternal Myopathy and Cardiomyopathy (MMC) is also caused by a mutation in tRNALeu. Linkage Analysis Linkage – two loci on the same chromosome. Given two loci that are on the same chromosome in mice (with two alleles for each locus): o Determinant of ear color: E – dominant (normal), e- recessive (pale) o Determinant of eye color: R –dominant (normal), r- recessive (ruby) In doubly heterozygous animals (EeRr), in which both dominant alleles are present on one chromosome and both recessive alleles are on the other, one might expect only ER and er gametes, but this is not the case. Due to crossing over of non-sister chromatids of homologous chromosomes, all gametes are possible – (ER, Er, eR, er). o Gametes are classified as either: Parental – chromosome with the same genotype as either of the parental chromosomes (i.e. ER or er). Recombinant – chromosome having a different genotype from either of the parental chromosomes (i.e. eR or Er). **There will not always be a crossover event between any two loci on homologous chromosomes. The physical distance between two loci on the same chromosome is roughly proportional to the probability of a crossover event occurring between two loci. o Thus, the proportion of crossover events (and therefore the proportion of recombinant gametes) is a function of the distance between these two loci on a chromosome. o The frequency of observed recombinants is used as a measure of this distance. o Distances described in “genetic map units.” Map units (m.u.) cannot be directly converted into a physical distance, but they are roughly equivalent to 1-2 x 106 bp. One m.u. is the distance between gene pairs for which 1 product out of every 100 is recombinant. A recombinant frequency (RF or �) of 0.01 (i.e. 1%) is defined as 1 m.u. A map unit is also commonly referred to as a centiMorgan (cM). Example Backcross –Double heterozygote mouse with a double homozygote Recombinant genotypes of offspring arise from crossing over during meiosis in the EeRr parent. Crossing over occurs in eerr parent, but it produces no change in gametes. Linkage Analysis and Human Disease Linkage of a gene and a particular trait begins with studying the segregation of the disease in large families using polymorphic markers. For some diseases, presence of the aberrant phenotype is very closely associated with the inheritance of particular alleles for a locus or set of loci (haplotype). o These loci are said to be in linkage disequilibrium. o Such associations can be used in populations (instead of families) to identify candidate genes (genome-wide association studies). In this pedigree, we also see that individual II-3 has a recombinant chromosome. This also indicates that the disease gene is NOT associated with the A Locus. The haplotype 2,3,3 for loci B,C,D is associated with the disease. **The polymorphic alleles result from the presence or absence of a BamHI site. 2.0 or 2.7 kb bands are observed after hypridization with a specific DNA probe. This figure illustrates how crossing over affects the presence of different alleles in subsequent generations. Haplotypes and Linkage Haplotype – two or more linked loci, considered as a set. Recombination fraction (�) = ratio of recombinant to total haplotypes o � = Recombinant / (Recombinant + Parental) o Range between 0 (complete linkage, thus no recombination), to 0.5 (unlinked – either on separate chromosomes or far apart on same chromosome). o If � = 0.01 = 1 recombinant / 100 gametes = 1 m.u. = 1 cM Lod Scores Linkage analysis is conducted by examination of likelihood (odds) ratios. o Ratios are studied for � ranging from 0-0.5. o Geneticists prefer working with smaller numbers, so odds ratios are transformed by taking their logarithms (base 10). o Log10 of an odds ratio is called a Lod Score (“log of odds”). Lod Scores are calculated as: o Log10 (likelihood of observed data at specified � / likelihood of observed data at � =0.5) o In other words, log10 (likelihood of linkage / likelihood of no linkage) Use of Lod scores: o Lod scores for a particular trait from different families can be combined, which is useful in situations in which finding large families with affected individuals is problematic. Maximum likelihood method – the highest Lod score is considered the best estimate of �. o Lod > +3.0 = linkage o Lod < -2.0 = no linkage It is important to remember that high Lod scores within families indicate linkage, but not all families will exhibit similar linkages. The graph shows completely different interactions between loci in different families. Heterogeneity between families is due to random recombination events that vary from family to family. When calculating the Lod score for this pedigree, the specified � was 0.05. In each individual of generation III, the association of a disease phenotype with inheritance of allele B and a regular phenotype with inheritance of A would exhibit a likelihood of 1-0.05 = 0.95. At the given � , the Lod score is inconclusive for linkage (<+3.0). In this example, a larger family exhibiting complete concordance (as in first example), results in a higher Lod score simply due to a greater amount of data. Lod Score Example Showing Incomplete Concordance (Presence of Recombinant Progeny) In this example, the presence of a recombinant son decreased the Lod score to an insignificant score. Genetic Mapping and Quantitative Trait Loci (QTLs) A QTL is a chromosomal region containing a gene (or genes) that affect the phenotypic expression level of a quantitative trait (i.e. blood pressure, height, intelligence). Using genetic mapping, QTLs can be associated (correlated) with specific phenotypes when they reach a significant Lod threshold: Genome-Wide Association Studies (GWAS) Used to associate loci with disease trait in populations (instead of families). Take advantage of linkage disequilibrium. Can be used in case-control (i.e. patients with or without disease), or cohort studies. The intention is to identify candidate genes/loci, but to date, rarely identify causative variant. Loci identified usually have modest effects.