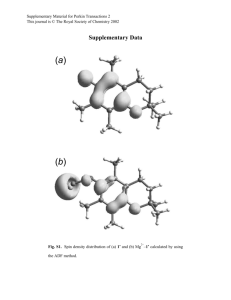
Supplementary Information, “Characterization of putative
advertisement

Supplementary Information, “Characterization of putative glycosylphosphatidylinositolanchoring motifs for surface display in the methylotrophic yeast Hansenula polymorpha” Seon Ah Cheon • Jinhee Jung • Jin Ho Choo • Doo-Byoung Oh • Hyun Ah Kang S. A. Cheon • J. H. Choo • H. A. Kang Department of Life Science, College of Natural Science, Chung-Ang University, Seoul, 156756, Korea J. Jung • D.-B. Oh Biochemicals and Synthetic Biology Research Center, Korea Research Institute of Bioscience and Biotechnology, 125 Gwahak-ro, Yuseong-gu, Daejeon, 305-806, Korea Contents Supplementary Methods: Plasmids constructions Supplementary Table 1 H. polymorpha strains and vectors used in this study. Supplementary Table 2 Primers used in this study. Supplementary Fig. 1 In silico identification of putative GPI-proteins of H. polymorpha. Supplementary Fig. 2 Flowchart of the cell wall fractionation procedure to analyze the locations of GPI-proteins in H. polymorpha. Supplementary Methods Plasmids constructions All vectors constructed in this study are listed in Supplementary Table 1. The 2.2 kb DNA fragment encoding the HpOch1(1-53aa)-msdS-FLAG fragment was obtained by digestion with SpeI/ClaI from pDTMOX-HH1MSF (Cheon et al. 2009) and inserted into the corresponding sites of pDUM2-msdS, which is a modified version of pDUMOX-msdS(HA-HDEL) (Kim et al. 2006) lacking an unique SalI site, resulting in the vector pDUM2-HHMSF. The 0.6 kb DNA fragment containing the partial MOX promoter, α-amylase signal sequence from A. niger, and a c-Myc tag was amplified from pDLMOX-GOD(H) (Kim et al. 2004) using two sequential rounds of PCR with primer sets MOXaa_9F and MOXaa_8B and MOXaa_9F and MOXaa_10B, and cloned between the KpnI and XbaI sites of pDUM2-HHMSF, generating the pDUM2-aaF vector. Then the 1.4 kb msdS fragment generated by digestion with XbaI from pDUM2-HHMSF was reintroduced into the pDUM2-aaF vector, resulting in the pDUM2aaMSF vector containing the msdS gene fused with the α-amylase signal sequence of A. niger and a c-Myc tag. To construct a positive control vector for msdS surface display, the DNA fragment encoding the C-terminal fragment of Tip1p (40 amino acids) (Kim et al. 2002) was amplified from genomic DNA of the DL1-L strain using PCR primers TIP40_11F_SalI/TIP40_12B_SalI (Supplementary Table 2), and cloned into a SalI site of pDUM2-aaMSF, resulting in pDUM2-aaMSF-Tip40. The DNA fragments encoding the Cterminal fragments (40 amino acids) of ten putative GPI-anchored proteins were amplified from the genomic DNA of the A16 (leu2) strain derivative of CBS4732 (ATCC34438) using the PCR primers listed in Supplementary Table 2, and cloned into SalI or SalI/ClaI sites of pDUM-aaMSF, generating the pDUM2-CCW40, -CRH40, -73/40, -CTS40, -EXG40, -133/40, -SPS40, -135/40, and -518/40 vectors, respectively. Supplementary Table 1 H. polymorpha strains and vectors used in this study Strain name Genotype Reference DL1-LdU leu2 ura3Δ::lacZ (Kang et al. 2002) DL1-g11 leu2 ura3Δ::lacZ och1Δ::lacZ (Kim et al. 2006) CBS3742 (A16) leu2 (Lahtchev 2002) Plasmid name Description Reference pDUM2-msdS Removed an unique SalI site from pDUMOXmsdS(HA-HDEL), HARS36, HpURA3 This study pDUM2-aaF pMOX- ss*-c-Myc-FLAG This study pDUM2-aaMS pMOX-ss-c-Myc-msdS-FLAG This study pDUM2-aaMSF-TIP1 pMOX-ss-c-Myc-msdS-FLAG-TIP1(C40)** This study pDUM2-aaMSF-CCW14 pMOX-ss-c-Myc-msdS-FLAG-CCW14(C40) This study pDUM2-aaMSF-CRH1 pMOX-ss-c-Myc-msdS-FLAG-CRH1(C40) This study pDUM2-aaMSF-73 pMOX-ss-c-Myc-msdS-FLAG-ORF73(C40) This study pDUM2-aaMSF-CTS2 pMOX-ss-c-Myc-msdS-FLAG-CTS2(C40) This study pDUM2-aaMSF-EXG1 pMOX-ss-c-Myc-msdS-FLAG-EXG1(C40) This study pDUM2-aaMSF-133 pMOX-ss-c-Myc-msdS-FLAG-ORF133(C40) This study pDUM2-aaMSF-SPS2 pMOX-ss-c-Myc-msdS-FLAG-SPS2(C40) This study pDUM2-aaMSF-518 pMOX-ss-c-Myc-msdS-FLAG-ORF518(C40) This study pDUM2-aaMSF-135 pMOX-ss-c-Myc-msdS-FLAG-ORF135(C40) This study pDLMOX-GOD(H) pMOX-ss-GOD-6xHis, HpLEU2 (Kim et al. 2004) * ss, A. niger α-amylase signal sequence ** (C40): C-terminal 40 amino acids Supplementary Table 2 Primers used in this study Primer Name Sequences (5’ to 3’) MOXaa_9F_KpnI acggggtaccttgcatcct MOXaa_8B ttctgagatgagtttttgttcggccaaagcaggtgccgc MOXaaCM_10B tcttctagacagatcctcttctgagatgagtttttgttc TIP40_11F_SalI tagtgggtcgacgctggatctagctccgct TIP40_12B_SalI catgctgtcgacttacataagcagagctgcaag CCW1_C40_1F_SalI tagtgggtcgacgtttcctcttcttctgcagc CCW1_C40_2B_SalI catgctgtcgacctaaagaagaccgatcaaga CRH1_C40_1F_SalI tagtgggtcgacggcaactcctcgtcgcagtct CRH1_C40_2B_SalI catgctgtcgacttagatcaaggcgagtccaaac ORF73_1F_SalI agtgggtcgacacggcaagcacggccagc ORF73_2B_ClaI tccatcgattctatagtacacaaatcagtcc CTS2_3F_SalI agtgggtcgactacccagatgagatggatg CTS2_4B_ClaI tccatcgatttacgagatgaataccagaat EXG1_5F_SalI agtgggtcgacaaatatgcctctgttctgtct EXG1_6B_ClaI tccatcgatttacagtaattctagtcctag ORF333_7F_SalI agtgggtcgactcgtcaacggtcagaaacg ORF333_8B_ClaI tccatcgattcactcaaacataaagcagtg SPS2_11F_SalI agtgggtcgacgacggcgaccacgcaaaac SPS2_12B_ClaI tccatcgatttagagctgcatagcaagc ORF135_13F_SalI agtgggtcgacccacgtgaagacattccg ORF135_14B_ClaI tccatcgattctattgaggagacatgaccatc ORF518_15F_SalI agtgggtcgactacgatgacgaaggaacttc ORF518_16B_ClaI tccatcgattctagatcagggcagccaa *Restriction enzyme sites were underlined Supplementary Fig. 1 In silico identification of putative GPI-proteins of H. polymorpha. For screening GPI-anchored proteins, the 5,483 annotated ORFs of the H. polymorpha CBS4732 strain (ATCC34438) were analyzed systemically using bioinformatic analysis programs, including GPI-SOM (Fankhauser and Maser), PredGPI (Pierleoni et al. 2008), big-PI (Eisenhaber et al. 2004), and fragAnchor (Poisson et al. 2007). Further analysis of putative GPI anchored proteins was performed using SignalP 4.1 server for signal peptides (Petersen et al. 2011), Psort II (Nakai and Horton 1999) and WoLF PSORT (Horton et al. 2007) for protein localization, TMHMM (Sonnhammer et al. 1998) for transmembrane domain, NetNGlyc 1.0 server (http://www.cbs.dtu.dk/services/NetNGlyc/) for N-linked glycosylation sites, NetOGlyc 3.1 and 4.0 servers (Julenius et al. 2005; Steentoft et al. 2013) for O-linked glycosylation sites, and CLC Main benchwork (CLC bio) for analyses of BLASTp, protein alignment, protein properties, and amino acid frequencies Supplementary Fig. 2 Flowchart of the cell wall fractionation procedure to analyze the locations of GPI-proteins in H. polymorpha. The cell wall proteins of H. polymorpha were isolated as described in Materials and Methods References Cheon SA, Choo J, Ubiyvovk VM, Park JN, Kim MW, Oh DB, Kwon O, Sibirny AA, Kim JY, Kang HA (2009) New selectable host-marker systems for multiple genetic manipulations based on TRP1, MET2 and ADE2 in the methylotrophic yeast Hansenula polymorpha. Yeast 26:507-521 Eisenhaber B, Schneider G, Wildpaner M, Eisenhaber F (2004) A sensitive predictor for potential GPI lipid modification sites in fungal protein sequences and its application to genome-wide studies for Aspergillus nidulans, Candida albicans, Neurospora crassa, Saccharomyces cerevisiae and Schizosaccharomyces pombe. J Mol Biol 337:243-253 Fankhauser N, Maser P (2005) Identification of GPI anchor attachment signals by a Kohonen self-organizing map. Bioinformatics 21:1846-1852 Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:W585-587 Julenius K, Molgaard A, Gupta R, Brunak S (2005) Prediction, conservation analysis, and structural characterization of mammalian mucin-type O-glycosylation sites. Glycobiology 15:153-164 Kang HA, Sohn JH, Agaphonov MO, Choi ES, M.D. T-A, Rhee SK (2002) Development of expression systems for the production of recombinant proteins in Hansenula polymorpha DL-1. In: Gellissen G, editor. Hansenula polymorpha-Biology and Applications. Weinheim: Wiley-VCH. p 124-146 Kim MW, Kim EJ, Kim JY, Park JS, Oh DB, Shimma Y, Chiba Y, Jigami Y, Rhee SK, Kang HA (2006) Functional characterization of the Hansenula polymorpha HOC1, OCH1, and OCR1 genes as members of the yeast OCH1 mannosyltransferase family involved in protein glycosylation. J Biol Chem 281:6261-6272 Kim MW, Rhee SK, Kim JY, Shimma Y, Chiba Y, Jigami Y, Kang HA (2004) Characterization of N-linked oligosaccharides assembled on secretory recombinant glucose oxidase and cell wall mannoproteins from the methylotrophic yeast Hansenula polymorpha. Glycobiology 14:243-251 Kim SY, Sohn JH, Pyun YR, Choi ES (2002) A cell surface display system using novel GPIanchored proteins in Hansenula polymorpha. Yeast 19:1153-1163 Lahtchev K (2002) Basic genetics of Hanseula polymorpha. In: Gellissen G, editor. Hansenula polymorpha : biology and applications: Wiley-VCH Nakai K, Horton P (1999) PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization. Trends Biochem Sci 24:34-36 Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785-786 Pierleoni A, Martelli PL, Casadio R (2008) PredGPI: a GPI-anchor predictor. BMC Bioinformatics 9:392 Poisson G, Chauve C, Chen X, Bergeron A (2007) FragAnchor: a large-scale predictor of glycosylphosphatidylinositol anchors in eukaryote protein sequences by qualitative scoring. Genomics Proteomics Bioinformatics 5:121-130 Sonnhammer EL, von Heijne G, Krogh A (1998) A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6:175182 Steentoft C, Vakhrushev SY, Joshi HJ, Kong Y, Vester-Christensen MB, Schjoldager KT, Lavrsen K, Dabelsteen S, Pedersen NB, Marcos-Silva Let al (2013) Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J 32:1478-1488