slides - Department of Mathematics
advertisement

Feature Identification for Colon
Tumor Classification
UCI Interdisciplinary Computational and Applied Mathematics Program Representative:
Anthony Hou
Joint Work with Melody Lim, Janine Chua, Natalie Congdon
Faculty Advisors: Dr. Fred Park, Dr. Ernie Esser, and Anna Konstorum
Problem Statement
Tumor
spheroids
Control
Chemical Added
Biological Background
Hepatocyte Growth Factor (HGF) has been shown to be
increased in colon tumor microenvironment (in vivo)
Increased HGF is correlated with increased growth &
dispersiveness
Tumor
spheroids
Control
+HGF
Experimental Approach
Data obtained from the Laboratory of Dr. Marian
Waterman, in the Department of Microbiology at UC Irvine
Cell line used: primary, ‘colon cancer initiating cells’
(CCICs)
Cultured CCICs trypsinized and spun down
Experimental Approach (cont.)
Single cells plated in 96 well ultra-low attachment plates
with DMEM, supplement, and with or without HGF at
various concentrations
CCICs imaged at 10x resolution once a day for 12 days
Spheroid grown in media +
50ng/ml HGF, day 8
Our Motivational Goal
Having a set of data, biologists can see the qualitative
effect when the concentration of HGF is high and when
the concentration of HGF is low.
We want to find the feature(s) that can discriminate
between a tumor spheroid that has high and low
concentrations of HGF.
We hope this discovery can indicate which features are
useful in helping biologists measure the amount of HGF in
a certain colon tumor spheroid
Image Processing/Computer
Vision Background
Classification
We humans have an innate ability to learn to identify
one object from another
Now, how can we automate this process with
respect to biological images?
Control
+HGF
Classification Approach
Image Processing
Mathematical features
Shape features: Area, Perimeter/Area, Circularity Ratio,
Texture features: Total Variation/Area, Average
Intensity, Eccentricity
Why these 6 features?
Given feature: Day
Fisher’s Linear Discriminant (FLD) Classification
Processing Data
Raw +HGF tumor
Binary image with
boundary applied
Boundary
of +HGF
tumor
Segmented +HGF
tumor
Thresholded
binary
image
Shape Information
HGF Binary
Features from
Given Shape
•
•
•
•
Area
Perimeter/Area
Circularity Ratio
Eccentricity
Image Information
HGF Segmented
Features from
Given Image
• Total
Variation
• Average
Intensity
Classification
<V1,V2, …Vn>
Tumor gets mapped to feature vectors, which get mapped to points in high
dimensional space. Now how do we separate the 2 groups?
Fisher’s Linear Discriminant
Describe mapping
Fisher’s Linear Discriminant: maximize ratio of inter-class
variance to intra-class variance
Project Overview
Develop classification scheme for colon tumor spheroids
grown in media with and without HGF
Broader goal is to obtain quantitative understanding of
HGF action on tumor spheroids.
Feature vectors can be utilized to quantify HGF action on
tissue growth in vitro.
Results
Ran FLD code on 6 features: Area, Circularity Ratio,
Average Intensity, Eccentricity, Perimeter/Area, TV/Area
Train on half the data
Repeated Random Sub-sampling Cross Validation was used
on all tests
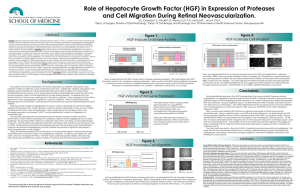
Results
Ran FLD code on 6 features: Area, Circularity Ratio,
Average Intensity, Eccentricity, Perimeter/Area, TV/Area
Percent Correct for Control: 91.50%
Percent Correct for +HGF: 90.99%
Results: Adding Day
Good results, but our goal is to maximize percentage
correct, so included time (day)
Features used: Area, Perimeter/Area, TV/Area, Eccentricity,
Average Intensity, Circularity Ratio, Day
Observed some tumors similar in shape and size, so we
needed a descriptor to separate those. Caused by larger
control tumor from later phase having similar area &
perimeter to earlier-stage HGF tumor.
Results: Adding Day
Good results, but our goal is to maximize percentage
correct, so included time (day)
Features used: Area, Perimeter/Area, TV/Area, Eccentricity,
Average Intensity, Circularity Ratio, Day
Observed some tumors similar in shape and size, so we
needed a descriptor to separate those. Caused by larger
control tumor from later phase having similar area &
perimeter to earlier-stage HGF tumor.
Percent Correct for Control: 98.88%
Percent Correct for +HGF: 100%
Next Approach
Excellent results, but curious to see if same results can be
obtained using less features
Plot all separately to get an idea of their individual
classifying potential
Area
Control=blue
HGF=red
Due to area differences between tumors from control and +HGF
Circularity Ratio Description
C1 = (Area of a shape)/(Area of circle)
where circle has the same perimeter as shape
Circularity Ratio
Control=blue
HGF=red
Given data are relatively circular from both groups (control and +HGF)
Average Intensity Description
Average Intensity: sum of the image intensities over
the shape divided by area
Inversely related to density.
Smaller values indicate less light passing through,
suggesting a denser object
+HGF 10ng/ml Day 11 (10x)
Control Day 8 (10x)
Average Intensity
Control=blue
HGF=red
• Control Group is similar in Average Intensity, whereas +HGFs are denser
• Not all are very dense, so there are some overlap with controls
Eccentricity Description
Measure of elongation of an object
Eccentricity
Control=blue
HGF=red
Due to most tumors from both groups being circular except for a few
outliers
Perimeter to Area Ratio
Why Normalize Perimeter by Area?
We do so because a small, jagged object may have
the same area as a large, circular object. Thus, we
divide by area, creating a more effective classifier.
Perimeter to Area Ratio
Control=blue
HGF=red
This is to be expected because the +HGF tumor spheroids have more dispersion,
resulting in greater area, in contrast to the control tumor spheroids.
Total Variation to Area Ratio
Description
At every point, estimate its gradient (difference in
intensities in x and y direction). Use discretization of Total
Variation. Also normalized by area.
Texture
Control
Day 11
(10x)
+HGF 10ng/ml
Day 12 (10x)
Total Variation to Area Ratio
Control=blue
HGF=red
Due to similar densities/intensities in tumors from both groups
Intuition Through Trial and
Error
Given the individual results, we combined the two
strongest features, area and perimeter/area, and plot
them both using a scatter plot
Area vs. Perimeter/Area
Control=blue
HGF=red
Results
We obtained reasonably accurate results, having only two
controls on the +HGF side if we draw an imaginary line to
separate the two groups
Ran FLD code on Area and Perimeter/Area
Results
We obtained reasonably accurate results, having only two
controls on the +HGF side if we draw an imaginary line to
separate the two groups
Ran FLD code on Area and Perimeter/Area
Percent Correct for Control: 89.03%
Percent Correct for +HGF: 96.92%
Evaluation
Reasonably decent results, but decided to add the feature
Day
Evaluation
Reasonably decent results, but decided to add the feature
Day
Results: Area, Perimeter/Area, Day
Percent Correct for Control: 100%
Percent Correct for +HGF: 100%
“Bad” Features
Plotting graphs of “good” features and running FLD
showed how strong those features really are.
Our first thoughts: Were the “good” features too strong
that the “bad” features couldn’t exhibit their full potential
as classifiers?
CR, TV/Area, Average Intensity, Eccentricity
Intuition
Decided to run FLD test to see if they perform better
as a group by themselves
Results: CR, TV/Area, Average Intensity, Eccentricity
Intuition
Results: CR, TV/Area, Average Intensity, Eccentricity
Percent Correct for Control: 75.33%
Percent Correct for HGF: 55.27%
Why?
Final Thoughts
Our belief: “bad” features are not necessarily
useless.
Data sets vary; some may include tumors with different
textures, shapes, area, and so on
Our set of features are extremely versatile
After feature identification, features can be used to
further pursue broader goals such as the quantification of
a certain chemical’s effect on their tumors
Conclusion
Effectiveness of area vector is obviously in accordance with
biological hypothesis that HGF increases cellular mitosis
rate, resulting in larger tumors.
Effectiveness of perimeter/area vector quantifies
contiguous cell spread, supporting hypothesis stating HGF
results in a spheroid with greater perimeter/area ratio.
Tried a lot of fancy ways, but turns out the strongest
features were the simplest ones that also agreed with
biologists’ intuition.
Conclusion (cont.)
Including Day Vs. Not Including Day
Day + less features = better results
Less features (without day) = worse results
Use more features (without day) = good results; separation
in high dimensions
Future Goals
Develop methods to quantify cell spread for cells that are
no longer attached to the tumor.
Develop an automated segmentation scheme
Occlusions
Existing strong methods worked, but needed more
preprocessing
+HGF 10ng/ml Day 13 (10x)
Future Experiments
EXPERIMENT IDEA #1:
Run experiment w/ different concentrations of HGF
We want to quantify how HGF acts with respect to increasing
concentration
Utilize developed feature vectors to classify images from
different concentrations of HGF.
Future Experiments
EXPERIMENT IDEA #2:
Stain spheroids for proteins associated with stem and
differentiated cell compartments
Stains can be incorporated into new feature vectors to
identify whether HGF-induced changes in stem /
differentiated cell concentrations are significant enough to
improve image classification.
Acknowledgements
NSF
Professors Jack Xin, Hongkai Zhao, Sarah Eichorn
Advisors: Dr. Fred Park, Dr. Ernie Esser, and Anna
Konstorum
Laboratory of Dr. Marian Waterman
Group: Janine Chua, Melody Lim, Natalie Congdon
MBI
References
[1] Thomas Brabletz, Andreas Jung, Simone Spaderna, Falk Hlubek, and
Thomas Kirchner. Opinion: migrating cancer stem cells - an integrated
concept of malignant tumour progression. Nat Rev Cancer, 5(9):744{749,
Sep 2005.
[2] Caroline Coghlin and Graeme I Murray. Current and emerging concepts
in tumour metastasis. J Pathol, 222(1):1{15, Sep 2010.
[3] A De Luca, M Gallo, D Aldinucci, D Ribatti, L Lamura, A D'Alessio, R De
Filippi, A Pinto, and N Normanno. The role of the egfr ligand/receptor
system in the secretion of angiogenic factors in mesenchymal stem cells. J
Cell Physiol, Dec 2010.