Practical2_day1
advertisement

Comparing human populations to identify positive selection. Practical.
Simon Myers and Gil McVean
The Concept
As you saw in the first practical, mutations under positive selection can arise and then reach
high frequency in a population (and eventually fix). This has important implications when we
are comparing diversity in two populations, for example in humans. Although most diversity
is shared among human groups, some genetic differences exist between human populations
as a result of historical separation – for example between Yorubans from Nigeria, and
Europeans from Utah, two populations that have been separated at least ~50,000 years, and
for whom extensive variation data has already been gathered, by the HapMap project1.
Two possible scenarios might lead to some positively selected mutations looking more
different between separated populations than “typical” mutations. First, the two groups A
and B may live in different environments, so selection on an existing mutation may only act
in one population, say A. In this case, this mutation might have reached high frequency in
population A, where it is advantageous, but still be rare in population B. Secondly, selective
forces may be the same in both groups, but a new advantageous mutation may arise in
population A after the populations separated. In this case, selection may again allow the
mutation to reach high frequency in population A (more rapidly than a typical mutation
would) and it will be virtually absent in population B.
Either case can result in the same signal – a mutation at very different frequencies in
populations A and B. Looking for such unusually differentiated SNPs can then help identify
selection acting in one or both populations. The key to finding such mutations is to identify
particular SNPs (Single Nucleotide Polymorphisms), based on diversity surveys, that are
unusually diverged between groups based on the distribution of SNPs in the genome.
Knowledge of the function of a particular mutation can add evidence and aid understanding
about the selection operating.
The Data
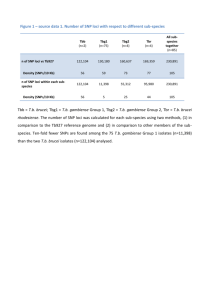
The file “chrom15data.txt” contains the population frequencies of SNPs, based on the
HapMap data1, in a region of the human genome of length 5 million bases and on
chromosome 15. The file contains six columns giving first a unique name for the SNP (called
the rs identifier), its position in bases along the chromosome, and then the frequency of the
SNP observed in samples from each of four populations originating from Europe (CEU),
Africa (YRI), Japan (JPT) and China (CHB), respectively.
Tasks
1. Load in “chrom15data.txt”. To do this and store the result as “data”, you can use
the following commands:
fid = fopen('chrom15data.txt', 'r');
data = textscan(fid, '%s %n %n %n %n %n');
fclose(fid);
For each pair of populations A and B, plot the column giving SNP frequency in population
A versus the column of SNP frequencies in population B. (There will be 6 plots
altogether.) Which pairs of groups show the most similar and most different allele
frequencies? Can you explain this?
2. Find the SNP(s) with the largest difference in absolute frequency between CEU and YRI,
between CEU and JPT and between CEU and CHB. Use the results to identify the single
SNP which most strongly separates CEU and the other three groups.
3. A more formal measure of differentiation between two groups A and B at a particular
SNP is given by FST. For a single SNP, this is calculated via the following formula:
FST 1.0
f A (1 f A )
f B (1 f B )
2 f (1 f )
Here fA and fB are the observed SNP frequencies for populations A and B, and f is the
mean SNP frequency:
f
fA fB
2
FST measures the proportion of variation at the SNP explained by between population
variability, relative to the total variation when populations are combined, and lies
between 0 and 1. Larger FST values indicate more differentiated SNPs.
Calculate and store a vector of single SNP FST values between CEU and JPT for each of the
1889 SNPs in the region. Make a plot of SNP position against the FST value. Which SNP
has the largest FST between these two populations?
4. Look for functional evidence at the SNP identified in 2. and 3. by going to the following
URL, which is the genome browser for the HapMap:
http://hapmap.ncbi.nlm.nih.gov/cgi-perl/gbrowse/hapmap27_B36/
Type in the rs identifier of the SNP you have found into the “Landmark or Region” box
and press enter. What results have genome-wide association studies found for this SNP?
By zooming out, find the nearest gene to the SNP you have found. Click on this gene to
obtain additional information.
5. What trait do these results suggest selection may have acted on, in human populations?
Reference
1.
Frazer, K.A. et al. A second generation human haplotype map of over 3.1 million
SNPs. Nature 449, 851-61 (2007).