Microarray data preprocessing Affy package (version 3.0, http://www
advertisement

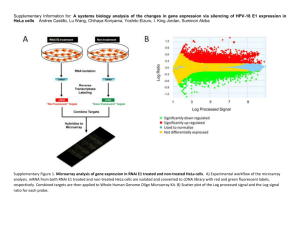
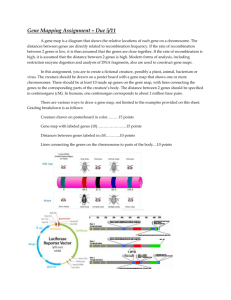
Microarray data preprocessing Affy package (version 3.0, http://www.bioconductor.org/packages/release/bioc/html/affy.html) [1] is written in statistical scripting language R and implements algorithms for processing raw microarray data into expression measures. Firstly, the CEL file data were read into Cel and AffyBatch objects using functions read.cefile and read.affybatch, including the MIAME experiment description data, information about experimental design and supplemental covariate information. A few of steps are involved in turning probe intensity data into gene expression measures, including background correction to eliminate background noise, normalization to detect and correct systematic differences between chips, perfect match correction, and computation of expression values to obtain expression matrix. Screening of differentially expressed genes Limma package (version http://www.bioconductor.org/packages/release/bioc/html/limma.html) 3.0, [2] in R language provides differential expression analysis for microarray data. Limma package is able to fit gene-wise linear models to gene expression data so as to assess differentially expressed genes (DEGs) using empirical Bayes method. Usually, the genes with P-value < 0.05 and |log2 fold change (FC)| > 0.5 are taken as differentially expressed. In the empirical Bayes moderated method, the corresponding P-values can be adjusted to control the false discovery rate using Benjamini and Hochberg method. Protein-protein interaction (PPI) analysis Proteins encoded by genes with similar functions are closely interacted with each other in complex biological metabolisms. As interactions between proteins possess such a crucial part in modern biology [3], STRING (Search Tool for the Retrieval of Interacting Genes, http://string-db.org/) emerges as an online database to provide uniquely comprehensive information for assembling, evaluating and disseminating PPIs in a user-friendly way [4]. Commonly, the value of combined score > 0.4 is regarded as criterion to screen out PPIs. Cytoscape software (version 3.2.1, http://cytoscape.org/) [5], a popular bioinformatics package for biological network visualization and data integration, can be used for global charting of PPI network. The PPI network with topological features provides a global picture used to understand molecular mechanisms and biological processes of disease [6, 7]. In the PPI network, nodes represent proteins, while edges represent interactions between proteins. The hub nodes are known to be the most important nodes in PPI network, which may corresponds to the important proteins in metabolic networks. Functional and pathway enrichment analysis Researches may be confused about biological interpretation of large gene lists derived from microarray analysis. DAVID (the Database for Annotation, Visualization and Integration Discovery) is developed as a publicly available high-throughput functional annotation tool for high-throughput data [8]. Using DAVID tools, KEGG (Kyoto Encyclopedia of Genes and Genomes, http://www.genome.jp/kegg/pathway.html) pathway enrichment analysis [9] can be performed to identify involved biological pathways. Also, GO (Gene Ontology, http://geneontology.org/) term analysis [10] can be implemented to identify biological processes, molecular mechanisms and cellular components enriched by interested genes. Usually, P-value < 0.05 and number of enriched genes > 2 are used as criteria to identify significant GO terms and KEGG pathways. References 1. Gautier L, Cope L, Bolstad BM, Irizarry RA. affy--analysis of Affymetrix GeneChip data at the probe level. Bioinformatics (Oxford, England). 2004;20:307-15. 2. Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015. 3. Jia P, Kao C-F, Kuo P-H, Zhao Z. A comprehensive network and pathway analysis of candidate genes in major depressive disorder. BMC systems biology. 2011;5:S12. 4. Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C. STRING v9. 1: protein-protein interaction networks, with increased coverage and integration. Nucleic acids research. 2013;41:D808-D15. 5. Cline MS, Smoot M, Cerami E, Kuchinsky A, Landys N, Workman C, Christmas R, Avila-Campilo I, Creech M, Gross B, Hanspers K, Isserlin R, Kelley R, Killcoyne S, Lotia S, Maere S, Morris J, Ono K, Pavlovic V, Pico AR, Vailaya A, Wang PL, Adler A, Conklin BR, Hood L, Kuiper M, Sander C, Schmulevich I, Schwikowski B, Warner GJ, Ideker T, Bader GD. Integration of biological networks and gene expression data using Cytoscape. Nat Protoc. 2007;2:2366-82. 6. Kar G, Gursoy A, Keskin O. Human cancer protein-protein interaction network: a structural perspective. PLoS computational biology. 2009;5:e1000601. 7. Song J, Singh M. How and when should interactome-derived clusters be used to predict functional modules and protein function? Bioinformatics. 2009;25:3143-50. 8. Da Wei Huang BTS, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols. 2008;4:44-57. 9. Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic acids research. 2000;28:27-30. 10. Gene Ontology C. The Gene Ontology (GO) database and informatics resource. Nucleic acids research. 2004;32:D258-D61.