File
advertisement

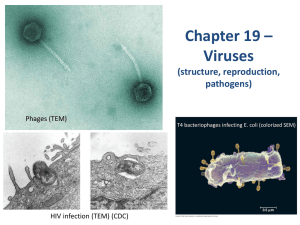
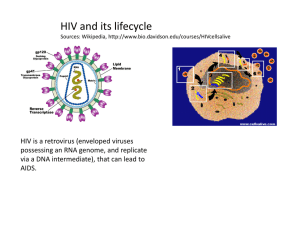
Chapter 19: Viruses Overview: A Borrowed Life virus- infectious particle consisted of genes packaged in a protein coat In the 1800s, researchers thought viruses were the simplest living form because they’re capable of causing a wide variety of diseases & can be spread between organisms Viruses aren’t living because they can’t reproduce or carry out metabolic activities outside a host cell. Molecular biology was born in the lab of biologists studying viruses that infect bacteria. Viruses were valuable experimental systems. Experiments with viruses provided important evidence that genes are made of nucleic acid, working out the molecular mechanisms of the processes of DNA replication, transcription, & translation. The study of viruses has led to the development of techniques that allow scientists to manipulate genes & transfer them from one organism to another, which are important in applications. 19.1 A virus consists of a nucleic acid surrounded by a protein coat The discovery of viruses: Scientific Inquiry Tobacco mosaic disease stunts the growth of tobacco plants & gives their leaves a mottled (mosaic) coloration In 1883, Adolf Mayer discovered that he could transmit the disease from plant to plant by rubbing sap extracted from diseased leaves onto healthy plants. He suggested that the disease was caused by super small bacteria invisible under microscopic view. Dimitri Ivanowsky tested the hypothesis by passing sap from infected tobacco leaves through a filter that removed bacteria. After filtration, the sap still produced mosaic disease. But he still hypothesized that bacteria caused the tobacco mosaic disease. He thought the bacteria were small enough to pass through the filter, or that it made a toxin that could do so. Martinus Beijerinck carried out an experiment that showed that the infectious agent in the filtered sap could replicate. The pathogen replicated only within the host it infected. It couldn’t replicate on its own, in test tubes. Wendell Stanley crystallized the infectious particle, tobacco mosaic virus (TMV). Structure of Viruses Viral Genomes Viruses’ genomes may consist of double-stranded DNA, single-stranded DNA, doublestranded RNA, or single-stranded RNA, depending on the type of virus The genome’s usually organized as a single linear or circular molecule of nuleic acid Viruses can contain 4 – 1000 genes Capsids & Envelopes Capsid- protein shell enclosing the viral genome; shape (rod-shaped, polyhedral, more complex) varies with type; built from large number of protein subunits called capsomeres, but the number of different kinds of proteins in a capsid is small TMV has a rigid, rod-shaped capsid made from 1000(+) molecules of a single type of protein arranged in a helix; a helical virus (rod-shaped) Adenovirus- infect respiratory tracts of animals; have 252 identical protein molecules arranged in a polyhedral capsid with 20 triangular facets (icosahedron); an icosahedral virus Some viruses have accessory structures that help them infect their hosts Viral envelope- membranous envelope that surrounds the capsids of viruses found in animals (ex: influenza); derived from membranes of host cell, contain host cell phospholipids & membrane proteins; contain proteins & glycoproteins (proteins with carbohydrates covalently attached) of viral origin. Bacteriophage (phage)- viruses that infect bacteria The first phages studied included 7 that infected E. coli, named T1 – T7, in the order they were found. o The T-even phages (T2, T4, T6) were very similar in structure. Their capsids have elongated icosahedral heads enclosing their DNA. Attached to the head is a protein tail piece with fibers by which the phages attach to a bacterium. 19.2Viruses replicate only in host cells Viruses lack metabolic enzymes needed for making proteins They are obligate intracellular parasites – they can only replicate within a host cell Each particular virus has a host range- the limited number of cells of host species that a virus can infect Viruses identify host cells by a lock & key fit between viral surface proteins & specific receptor molecules on the outside of cells. Different viruses have different host ranges. o West Nile & equine encephalitis viruses (distinctly different) have a broad range and can infect mosquitoes, birds, horses, & humans o Measles virus has a narrow range and can only affect humans. Viral infection of multicellular eukaryotes is limited to particular tissues o Human cold viruses infect only the cells lining the upper respiratory tract o AIDS virus binds to receptors present only on certain types of white blood cells General Features of Viral Replicative Cycles A viral infection beings when a virus binds to a host cell & the viral genome makes its way inside The way a genome enters depends on the type of virus & type of host cell o T-even phages use their tail apparatus to inject DNA into a bacterium o Others are taken up by endocytosis o Enveloped viruses are taken up by fusion of the viral envelope with the plasma membrane Once the viral genome’s inside, the proteins it encodes can hijack the host, reprogramming the cell to copy the viral nucleic acid & manufacture viral proteins. The host provides the nucleotides for making viral nucleic acids, enzymes, ribosomes, tRNAs, amino acids, ATP. Many DNA viruses use the DNA polymerases of their host cell to synthesize new genomes along the templates provided by the viral DNA RNA viruses use virally encoded RNA polymerases that can use RNA as a template to replicate their genome After the viral nucleic acid molecules & capsomeres are produced, they self-assemble into new viruses. The simplest type of viral relicative cycle ends with the exit of hundreds of viruses from the infected host cell, which damages or destroys the cell, which causes many symptoms of viral infections. The viral progeny that exit a cell can infect additional cells Replicative Cycles of phages Some double-stranded DNA viruses can replicate by 2 alternative mechanisms: the lytic cycle & the lysogenic cycle The Lytic Cycle Lytic cycle- ends in death of host cell; bacteria lyses (breaks open) & releases the phages that were produced within the cell; each of the phages can then infect healthy cells Virulent phage- replicates only by a lytic cycle Successive lytic cycles can destroy an entire bacterial population in just a few hours, but bacteria aren’t extinct because they have defense mechanisms. 1. Natural selection favors bacterial mutants with receptors that are no longer recognized by a particular type of phage Natural selection also favors phage mutants that can bind to altered receptors or are resistant to certain restriction enzymes 2. When phage DNA successfully enters a bacterium, the DNA often is identified as foreign & cut up by cellular enzymes called restriction enzymes, which restricts the ability of the phage to infect the bacterium Bacterial cell’s own DNA is methylated in a way that prevents attack by its own restriction enzymes 3. Instead of lysing their host cells, many phages coexist with them in lysogeny The lytic cycle of phage T4, a virulent phage T4’s genes are transcribed & translated using the host cell’s machinery. One of the 1st translated genes after the viral DNA enters codes for an enzyme that degrades the host cell’s DNA. The phage DNA’s protected from breakdown because it contains a modified form of cytosine that’s not recognized by the enzyme. 1. Attachment: T4 phage uses its tail fibers to bind to specific receptor sites on the outer surface of an E. coli cell 2. Entry of phage & degradation of host DNA: cover of tail bonds, injecting the phage DNA into the cell & leaving an empty capsid outside. The cell’s DNA is hydrolyzed 3. Synthesis of viral genomes & proteins: The phage DNA directs production of phage proteins & copies of the phage genome by host & viral enzymes, using components within the cell wall 4. Assembly: 3 separate sets of proteins self-assemble to form phage heads, tails, & tail fibers. The phage genome is packaged inside the capsid as the head forms. 5. Release: the phage directs production of an enzyme that damages the bacterial cell wall, allowing the fluid to enter. The cell swells & finally bursts, releasing 100s of phage particles. The lysogenic cycle Lysogenic cycle- allows replication of the phage genome without destroying the host Temperate phages- phages capable of using both modes of replicating within a bacterium Phage (lambda) is a temperate phage widely used in biological research that resembles T4, but its tail has only one short tail fiber. Infection of E. coli cell by phage 1. Phage binds to surface of cell & injects its linear DNA genome 2. Within the host, the DNA molecule forms a circle. 3. This step depends on the replicative mode: lytic cycle or lysogenic cycle Lytic cycle- viral genes immediately turn the host cell into a -producing factory & cell soon lyses & releases viral products Lysogenic- DNA molecule is incorporated into a specific site on the E. coli chromosome by viral proteins that break the circular DNA molecules & join them to each other. When integrated into the bacterial chromosome in this way, the viral DNA’s a prophage. 1 prophage gene codes for a protein that prevents transcription of most of the other prophage genes. The phage genome is mostly silent within the bacterium. Every time the E. coli cell prepares to divide, it replicates the phage DNA along with its own & passes the copies on to daughter cells. A single infected cell can give rise to a large population of bacteria carrying the virus in prophage form, which enables viruses to propagate without killing the host cells on which they depend Lysogenic implies that prophages are capable of generating active phages that lyse their cells. This occurs when the genome is induced to exit the bacterial chromosome & initiate a lytic cycle. An environmental signal (ex: certain chemical or high-energy radiation) triggers the switchover from lysogenic to lytic Some prophage genes may be expressed during lysogeny. Expression of these genes may alter the host’s phenotype, which has important medical significance o 3 sepcies of bacteria that cause diphtheria, botulism, & scarlet fever wouldn’t be so harmful to humans without certain prophage genes that cause the host bacteria to make toxins Replicative Cycles of Animal Viruses The nature of the genome (number of strands, type of nucleic acid) is the basis for the classification of viruses. Single-stranded RNA viruses are classified into 3 classes (IV – VI) according to how the RNA genome functions in a host cell Many animal viruses have both an envelope and RNA genome. Nearly all animal viruses with RNA genomes have an envelope. Viral Envelopes An animal virus with an envelope (outer membrane) uses it to enter the host cell. Protruding from the outer surface of the envelope are viral glycoproteins that bind to specific receptor molecules on the surface of the host cell. Ribosomes bound to the ER of the host cell make the protein parts of the envelope glycoproteins; cellular enzymes in the ER & Golgi apparatus then add the sugars. The resulting viral glycoproteins, embedded in the host cell membrane, are transported to the cell surface. Like exocytosis, new viral capsids are wrapped in a membrane as they bud from the cell. The viral envelope is derived from the host cell’s plasma membrane. The enveloped viruses are now free to infect other cells. This replicative cycle doesn’t necessarily kill the host cell. Some viruses have envelopes that aren’t derived from plasma membrane 1. Herpeviruses are temporarily cloaked in membrane derived from the nuclear envelope of the host. They then shed this membrane in the cytoplasm & acquire a new envelope made from the membrane of the Golgi apparatus. They have a double-stranded DNA genome & replicate within the host cell nucleus, using a combo of viral & cellular enzymes to replicate & transcribe their DNA. Copies of the viral DNA can remain behind as mini-chromosomes in the nuclei of certain nerve cells, where they remain latent until some physical or emotional stress triggers a new round of active virus production. The replicative cycle of an enveloped RNA virus A virus with asingle-stranded RNA genome functions as a template for synthesis of mRNA. The formation of new envelopes for progeny viruses occurs like this: 1. Glycoproteins on the viral envelope bind to specific receptor molecules on the host cell, promoting viral entry into the cell 2. The capsid & viral genome enter the cell. Digestion of the capsid by cellular enzymes releases the viral genome 3. The viral genome functions as a template for synthesis of complementary RNA strands by a viral RNA polymerase 4. New copies of viral genome RNA are made using complementary RNA strands as templates 5. Complementary RNA strands also function as mRNA, which is translated into both capsid protein (in the cytosol) & glycoproteins for the viral envelope (in ER & Golgi apparatus) 6. Vesicles transport envelope to plasma membrane 7. Capsid assembles around each viral genome molecule 8. Each new virus buds from the cell, its envelope studded with viral glycoproteins embedded in membrane derived from host cell New virus RNA as viral genetic material Genome of class IV single-stranded RNA genomes in animal viruses can directly serve as mRNA & can be translated into viral protein immediately after infection Class V- RNA genome serves as a template for RNA synthesis. RNA genome’s transcribed into complementary RNA strands, which function both as mRNA & as templates for the synthesis of additional copies of genomic RNA. All viruses that require RNA RNA synthesis to make mRNA use a viral enzyme capable of carrying out this process. The viral enzyme’s packaged with the genome inside the viral capsid. Retroviruses (class VI)- RNA animal viruses with the most complicated replicative cycles; equipped with reverse transcriptase (transcribes RNA template into DNA, providing RNA DNA info flow) o HIV (human immunodeficiency virus) causes AIDS (acquired immunodeficiency syndrome) They’re enveloped viruses that contain 2 identical molecules of single-stranded RNA & 2 molecules of reverse transcriptase After HIV enters a host cell, its reverse transcriptase molecules are released into the cytoplasm, where they catalyze synthesis of viral DNA. The newly made viral DNA then enters the cell’s nucleus & integrates into the DNA of a chromosome Provirus, integrated viral DNA, never leaves the host’s genome, remaining a permanent resident of the cell The host’s RNA polymerase transcribes the proviral DNA into RNA molecules, which can function both as mRNA for the synthesis of viral proteins & as genomes for the new viruses that’ll be assembled & released from the cell The replicative cycle of HIV 1. The envelope of glycoproteins enable the virus to bind to specific receptors on certain WBCs 2. The virus fuses with the cell’s plasma membrane. The capsid proteins are removed, releasing viral proteins & RNA 3. Reverse transcriptase catalyzes the synthesis of a DNA strand complementary to the viral RNA 4. Reverse transcriptase catalyzes the synthesis of a second DNA strand complementary to the 1st 5. The double-stranded DNA’s incorporated as a provirus into the cell’s DNA 6. Proviral genes are transcribed into RNA molecules, which serve as genomes for the next viral generations & as mRNAs for translation into viral protein 7. The viral proteins include capsid proteins & reverse transcriptase (made in cytosol) & envelope glycoproteins (made in ER) 8. Vesicles transport the glycoproteins to the cell’s plasma membrane 9. Capsids are assembled around viral genomes & reverse transcriptase molecules 10. New viruses bud off from the host cell Evolution of viruses Viruses aren’t descendents of precellular forms of life, rather they’ve evolved after the first cells appeared Most favored hypothesis of virus origin: viruses originated from naked bits of cellular nucleic acids that moved from one cell to another via injured cell surfaces. The evolution of genes coding for capsid proteins have facilitated the infection of uninjured cells. Candidates from the original sources of viral genomes (all are mobile genetic elements) 1. Plasmids- small, circular DNA molecules found in bacterial & unicellular eukaryotes called yeasts. They exist apart from the cell’s genome, can replicate independently of the genome, & are occasionally transferred between cells 2. Transposons- DNA segments that can move from one location to another within a cells genome Viral genome can have more in common with the genome of its host than with the genomes of viruses that infect other hosts. Some viral genes are identical to the hosts’ Genetic sequences of some viruses are similar to seemingly distinct viruses. This genetic similarity reflects the persistence of groups of viral genes that were favored by natural selection during the early evolution of viruses & the eukaryotic cells that served as their hosts. Debate about origin has been revived by reports of mimivirus, the largest virus discovered. Virus most likely evolved before the first cells & then developed an exploitative relationship with them. It is a double-stranded DNA virus with an icosahedral capsid 400 nm in diameter. Some of its genes code for proteins involved in translation, DNA repair, protein folding, & polysaccharide synthesis. Virus Diseases in Animals Viruses may damage or kill cells by causing the different release of hydrolytic enzymes from lysosomes Some viruses cause infected cells to produce toxins & lead to disease symptoms Some have molecular components that are toxic The degree of damage partly relies on the ability of the infected tissue to regenerate by cell division. o People recover from cols because the epithelium of the respiratory Many of the temporary symptoms associated with viral infections (ex: fever, aches) result from the body’s efforts at defending itself against infection.