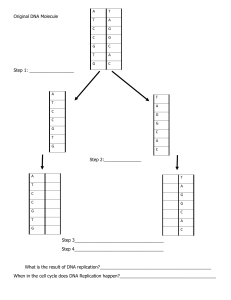
DNA: REPLICATION, RECOMBINATION AND REPAIR Objectives • The mode of DNA synthesis The Meselson-Stahl experiment The semiconservative replication Replication forks and origins • Synthesis of DNA DNA polymerases I,II,III Fidelity of synthesis (self study) • DNA synthesis; A model Unwinding of the helix Initiation of synthesis Continuous and discontinuous synthesis • DNA repair Objectives • Prokaryotic and Eukaryotic DNA synthesis • DNA recombination DNA replication • In a cell, DNA replication begins at specific locations in the genome, called "origins". • Unwinding of DNA at the origin, and synthesis of new strands, forms a replication fork. • In addition to DNA polymerase, the enzyme that synthesizes the new DNA by adding nucleotides matched to the template strand, a number of other proteins are associated with the fork and assist in the initiation and continuation of DNA synthesis. • Cellular proofreading that ensure near perfect fidelity for DNA replication. The Central Dogma • The Flow of Information: DNA → RNA → Protein • A gene is expressed in two steps: DNA is transcribed to RNA. Then RNA is translated into protein DNA replication • DNA replication, the basis for biological inheritance, is a fundamental process occurring in all living organisms to copy their DNA. • In the process of "replication" each strand of the original double-stranded DNA molecule serves as template for the reproduction of the complementary strand. • Two identical DNA molecules have been produced from a single double-stranded DNA molecule. DNA Replication • DNA replication is semiconservative (Meselson and Stahl (1958)) – One strand goes to the next generation – The other is new • Each strand is a template for the other – If one strand is 5’ AGCT 3’ – The other is 3’ TCGA 5’ DNA REPLICATION • Write the strand complementary to – 3’ ACTAGCCTAAGTCG 5’ – Answer • DNA occurs in many forms-linear,circular,single stranded or double stranded. • DNA initiates at specific origins – Unique origin in bacteria – Multiple origins in eukaryotic chromosomes • Is catalysed by DNA polymerases – E coli – DNA polymerase I and DNA polymerase III – Eukaryotes- α, ε and δ DNA REPLICATION • Challenges faced by the cell during replication – How to separate two DNA strands- the cell has to protect the unwound DNA from the action of nucleases that actually attack single strands of DNA. – Synthesis of DNA from the 5’ to the 3’ end- two strands have to be synthesized in the same direction on anti-parallel templates (meansthe template has 5’-3’ strand and one 3’- 5’ strand as does the new synthesized DNA. – How to guard against errors in replication (How do we ensure that the correct base is added to the growing polynucleotide chain. Three possible models were proposed for DNA replication: a. Conservative model proposed both strands of one copy would be entirely old DNA, while the other copy would have both strands of new DNA. b. Dispersive model was that dsDNA might fragment, replicate dsDNA, and then reassemble, creating a mosaic of old and new dsDNA regions in each new chromosome. c. Semiconservative model is that DNA strands separate, and a complementary strand is synthesized for each, so that sibling chromatids have one old and one new strand. This model was the winner in the Meselson and Stahl experiment. Three models for the replication of DNA Peter J. Russl, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. DNA Replication • Is semiconservative – Meselson and Stahl (1958) The Meselson-Stahl Experiment The Meselson-Stahl Experiment 1. Meselson and Stahl (1958) grew E. coli in a heavy (not radioactive) isotope of nitrogen, 15N in the form of 15NH Cl. Because it is heavier, DNA containing 15N is more 4 dense than DNA with normal 14N, and so can be separated by CsCl density gradient centrifugation. 2. Once the E. coli were labeled with heavy 15N, the researchers shifted the cells to medium containing normal 14N, and took samples at time points. DNA was extracted from each sample and analyzed in CsCl density gradients. The Meselson-Stahl experiment, which showed that DNA replicates semiconservatively Peter J. Russell, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. Equilibrium centrifugation of DNA of different densities in a cesium chloride density gradient Peter J. Russell, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. 3. After one replication cycle in normal 14N medium, all DNA had density intermediate between heavy and normal. After two replication cycles, there were two bands in the density gradient, one at the intermediate position, and one at the position for DNA containing entirely 14N. 4. Results compared with the three proposed models: a. Does not fit conservative model, because after one generation there is a single intermediate band, rather than one with entirely 15N DNA and another with entirely 14N DNA. b. The dispersive model predicted that a single band of DNA of intermediate density would be present in each generation, gradually becoming less dense as increasing amounts of 14N were incorporated with each round of replication. Instead, Meselson and Stahl observed two bands of DNA, with the intermediate form decreasing over time. c. The semiconservative model fits the data very well. How is DNA replicated? Replication Origins and forks (Bidirectional replication) • The DNA double helix unwinds at a specific point known as the origin of replication (OriC). • DNA replication proceeds toward the direction of the replication fork (leading strand) on one strand and away from the fork (lagging strand) on the other strand or in one direction only. • In eukaryotes, when 2 replication forks are too near, a replication bubble forms • The OriC consists of 245 base pairs-characterised by the presence of repeated sequences of 9 and 13 bases(9mers and 13 mers). Replication origins and forks • Replication begins when proteins bind at a specific site on the DNA known as the origin of replication (ori). • What is a replicon? (self study) – Eukaryotic replication is similar to prokaryotic replication but more complex? (why?) • The closed circular DNA of prokaryotes usually only has one origin of replication (ori) – Linear eukaryotic DNA has multiple ori’s Replication Origins and forks Eukaryotes Prokaryotes What are the functions of DNA polymerases? DNA Polymerases, the DNA Replicating Enzymes 1. First isolation of an enzyme involved in DNA replication was in 1957. Arthur Kornberg won the 1959 Nobel Prize for this work. (DNA polymerase I) 2. Additional DNA polymerases have been isolated, including DNA polymerase II (1970), DNA polymerase III (1971), DNA polymerase IV, and DNA polymerase V. Properties of three bacterial DNA polymerases ( in vivo) • POL I,II III- all possess a 3’ to 5’ exonuclease activity- they can polymerize in one direction, and then reverse directions and excise nucleotides just added. The activity provides a capacity to proofread and remove incorrect nucleotides. Only POL I can initiate DNA synthesis on a template • Pol I/(III)- possess the 5’ to 3’ exonuclease activity. The activity allows the enzyme to excise nucleotides from the end where initial synthesis occurred and in the same direction. They have a potential ability to remove the primer, an important aspect of synthesis. POL II and III can elongate an existing strand (primer- RNA) Roles of DNA Polymerases in vivo • POL I- responsible for removing the primer as well as the synthesis that fills in the gaps. Its exonuclease activity allows for proofreading. It requires a primer and a DNA template to assist in DNA synthesis. • POL II – suspected to be involved in repair synthesis of DNA that has been damaged by external forces like ultraviolet light. • POL III- It is responsible for the polymerization essential to replication. Its 3’ to 5’ exonuclease activity allows it to proof read,excise and then correct base pairs created in error during polymerization. Its active form is known as a holoenzyme. What constitutes a holoenzyme (POL III)? Roles of DNA Polymerases 1. All DNA polymerases link dNTPs into DNA chains . Main features of the reaction: a. An incoming nucleotide is attached by its 5’-phosphate group to the 3’OH of the growing DNA chain. Energy comes from the dNTP releasing two phosphates. The DNA chain acts as a primer for the reaction. b. The incoming nucleotide is selected by its ability to hydrogen bond with the complementary base in the template strand. The process is fast and accurate. c. DNA polymerases synthesize only from 5’ to 3’. 2. The enzyme Kornberg isolated was believed to be the only DNA polymerase in E. coli. The were two requirements for the in vitro DNA synthesis All four deoxyribonucleoside triphosphates (dATP,dCTP,dGTP, dTTP = dNTP) Template DNA DNA chain elongation catalyzed by DNA polymerase Peter J. Russell, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. Fidelity of synthesis (self study) • • • • Kornberg (isolated DNA polymerase I). Objective of his study What was his approach? Briefly describe the method used to test fidelity of synthesis (nearest-neighbour frequency test). How is DNA replicated in prokaryotic cells? objectives - Construct a model of DNA to stimulate DNA replication. - Understand why are RNA primers used in DNA replication - Understand how the DNA helix unwinds, initiates synthesis (primer introduction), continuous and discontinuous. - Link the process of DNA replication to the behavior of chromosomes in mitosis and meiosis (cell cycle) - Repair mechanisms (self study) Molecular Model of DNA Replication 1.Table:1 shows key genes and DNA sequences involved in replication. DNA Synthesis • DNA synthesis: A model – A mechanism must exist by which the helix undergoes localised unwinding and is stabilised in the open configuration so that the synthesis may proceed along both strands. – As unwinding proceeds, increased coiling creates tension further down the helix which must be reduced. – A primer must be synthesized so that polymerization can commence under the direction of polymeraseIII – When polymerase III commences synthesis of the compliment of both strands of the parent molecule, the two strands are anti-parallel to another. The other becomes continuous and the other becomes discontinuous in the opposite direction. – The primers must be removed before the completion of replication. Gaps that are created must be filled with DNA complementary to the template at each location. – The newly synthesized DNA strand that fills each gap must be ligated to the adjacent strand of DNA UNWINDING OF DNA Unwinding of DNA -The OriC consists of 245 base pairs-characterised by the presence of repeated sequences of 9 and 13 bases(9mers and 13 mers). – - DNA helicase: unwinds the double helix by breaking the Hbonds at the replication fork(region where enzymes replicating DNA bind to an untwisted, s.s. DNA strand). Helicases require energy supplied by the hydrolysis of ATP in order to break hydrogen bonds and denature the double helix. dnaA is responsible for the initial step in unwinding the helix. A number of its subunits bind to each of the several 9mers. It is essential for subsequent binding of dnaB and dnaC proteins that further open and destabilise the helix. • SSBPs- The single stranded binding proteins (SSBs) bind to the exposed bases to prevent them from annealing . Stabilises the conformation. • DNA gyrase: relieves tension from the unwinding of the DNA strands during bacterial replication. It cuts nicks in both strands of DNA, allowing them to swivel around one another and then resealing the cut strands Initiation, continuous and discontinuous DNA synthesis • In prokaryotes, there are 3 enzymes known to function in replication & repair – DNA polymerase I, II & III • In eukaryotes, there are 7 enzymes known to function in replication & repair Building Complimentary Strands in prokaryotes • RNA primers are synthesized by primase and are temporary • The leading strand (uses 3’-5’ template) is synthesized continuously • The lagging strand (uses 5’-3’ template) is synthesized discontinuously in short fragments Building Complimentary Strands in prokaryotes • DNA polymerase III builds the complimentary strand of DNA • DNA polymerase III adds complimentary nucleotides (deoxyribonucleoside triphosphates) in the 5’ to 3’ direction, using RNA primers as starting points – The segments are called Okazaki fragments Building Complimentary Strands in prokaryotes • DNA polymerase I removes the RNA primers from the leading strand and fragments from the lagging strand and replaces them with the appropriate deoxyribonucleotides. Building Complimentary Strands in prokaryotes • DNA ligase joins the Okazaki fragments into one strand on the lagging strand of DNA through the formation of a phosphodiester bond. Building Complimentary Strands in prokaryotes • As the 2 new strands of DNA are synthesized, 2 d.s. DNA molecules are produced that automatically twist into a helix. Initiation, continous and discontinous synthesis Summary of DNA synthesis • Synthesis is initiated at a specific origin within each replicon. In E.coli is called OriC • Unwinding proteins (helicases) denature the DNA helix at the origin, while the SSBPs stabilize the denatured DNA • Synthesis is bidirectinal, creating two replication forks which move in opposite directions away from the origin • As the replication fork moves away from the origin, increased coiling of the helix occurs and DNA gyrases diminishes the tension. It cuts and seals DNA strands after uncoiling occurs. • Initiation of DNA synthesis involves a RNA primer synthesized under the direction of a primase. The resultant RNA is complementary to its DNA template Summary of DNA synthesis • DNA polymerase III polymerizes complementary DNA strands by elongating an existing chain in the 5’ to 3’ direction. • As the replication forks move away from the origin, synthesis is continuous on the leading strand, but is discontinuous on the lagging strand, producing short strands known as okazaki fragments. • The RNA primers are removed and the resulting gaps are filled with DNA under the direction of DNA polymerase I • Along the lagging strand, the Okazaki fragments are joined by DNA ligase. Summary of DNA synthesis • As this process proceeds along the length of the replicon, proofreading by DNA polymerase I and III occurs and semi conservative replication is achieved DNA Repair DNA Repair (Proofreading) • DNA polymerase III and DNA polymerase I proofread the newly synthesized DNA strands. • When mistakes occur, either enzyme can function as an exonuclease. – The enzyme backtracks and excises the incorrectly paired nucleotide – Then it continues forward adding nucleotides to the complimentary strand DNA Repair • Repairs must be made immediately to avoid errors being copied in subsequent replications. • Errors missed by proofreading can be corrected by one of several repair mechanisms that operate after the completion of DNA replication. Self study • DNA damage repair mechanisms – Excision repair – Mismatch repair – The SOS response – Double strand Break repair Replication of circular DNA and the supercoiling problem 1. Some circular chromosomes (e.g., E. coli) are circular throughout replication, creating a theta-like (θ) shape. As the strands separate on one side of the circle, positive supercoils form elsewhere in the molecule. Replication fork moves about 500 nt/ second, so at 10 bp/turn, replication fork rotates at 3,000 rpm. 2. Topoisomerases relieve the supercoils, allowing the DNA strands to continue separating as the replication forks advance. Rolling Circle Replication 1. Another model for replication is rolling circle, which is used by several bacteriophages, including ΦX174 (after a complement is made for the genomic ssDNA) and λ (after circularization by base pairing between the “sticky” ssDNA cos ends) 2. Rolling circle replication begins with a nick (singlestranded break) at the origin of replication. The 5’ end is displaced from the strand, and the 3’ end acts as a primer for DNA polymerase III, which synthesizes a continuous strand using the intact DNA molecule as a template. 3. The 5’ end continues to be displaced as the circle “rolls”, and is protected by SSBs until discontinuous DNA synthesis makes it a dsDNA again. The replication process of double-stranded circular DNA molecules through the rolling circle mechanism Chapter 3 slide 56 Prokaryotic DNA replication in bacteriophages Bacteriophage M13 • On infecting an E.coli cell, the viral strand directs the synthesis of its complemetary strand to form a circular duplex replicative form (RF), which can either be nicked (RF II) or supercoiled (RF I). • As the M13(+) strand enters the E.coli cell, it becomes coated by SSB except a segment of 57nt that form the hairpin. RNA polymerase commences primer synthesis 6nt before start of the hairpin and extends the RNA 20 to 30 residues to form a segment of RNA-DNA hybrid duplex. The DNA is displaced from the hairpin and becomes coated with SSB so that when RNA polymerases reaches it, primer synthesis stops. Pol III holoenzyme then extends the RNA primer around the circle to form the (-) strand. The primer is removed by Pol I catalyzed nicktranslation, thereby forming RFII, which is converted to RF I by sequential actions of DNA ligase and DNA gyrase. • • • • Prokaryotic DNA replication in bacteriophages (Self study) Bacteriophage φX174 • The reaction sequence begin as that of M13. the (+) strand is coated with SSb except for a 44 unit hairpin. A 70-nt sequence containing hairpin is known as pas (primosome binding site), which is recognised & bound by the PriA, Pri B and Pri C proteins. • DNaB and DNaC proteins are added to the DNA with help of DNaT protein in an ATP requiring process. DNaC is then released yielding a preprimosome, which in turn binds primase yielding the primasome. • At randomly selected sites, the primasome reverses its migration while primase synthesizes an RNA primer.. • Pol III holoenzyme extends the primers to form Okazaki fragments. • Pol I excises the primers and then replaces them by DNA. The fragments are then joined by DNA ligase and supercoiled by DNA gyrase to form φX174 RF I. How is DNA replicated in eukaryotic cells? DNA Replication in Eukaryotes 1.DNA replication is very similar in both prokaryotes and eukaryotes, except that eukaryotes have more than one chromosome. Replicons 1. Eukaryotic chromosomes generally contain much more DNA than those of prokaryotes, and their replication forks move much more slowly. If they were like typical prokaryotes, with only one origin of replication per chromosome, DNA replication would take many days. 2. Instead, eukaryotic chromosomes contain multiple origins, at which DNA denatures and replication then proceeds bidirectionally until an adjacent replication fork is encountered. The DNA replicated from a single origin is called a replicon, or replication unit 3. In eukaryotes, replicon size is smaller than it is in prokaryotes, replication is slower, and each chromosome contains many replicons. Number and size of replicons vary with cell type. 4. Not all origins within a genome initiate DNA synthesis simultaneously. Cell-specific patterns of origin activation are observed, so that chromosomal regions are replicated in a predictable order in each cell cycle Fig. 3.13 Temporal ordering of DNA replication initiation events in replication units of eukaryotic chromosomes Peter J. Russell, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. Eukaryotic Replication Enzymes 1. Enzymes of eukaryotic DNA replication aren’t as well characterized as their prokaryotic counterparts. The replication process is similar in both groups—DNA denatures, replication is semiconservative and semidiscontinuous and primers are required. 2. Fifteen DNA polymerases are known in mammalian cells: a. Three DNA polymerases are used to replicate nuclear DNA. Pol α (alpha) extends the 10-nt RNA primer by about 30nt. Pol δ(delta) and Pol ε(epsilon) extend the RNA/DNA primers, one on the leading strand and the other on the lagging (it is not clear which synthesizes which). b. Other DNA pols replicate mitochondrial or chloroplast DNA, or are used in DNA repair. Eukaryotic DNA Polymerases • –DNA polymerases in eukaryotes • At least 7 polymerases: α, β, γ, δ, ε, ς, η • Polymerases α, δ and ε work together in the replication of nuclear DNA • Polymerases β, ς, η and ε function in nuclear DNA repair • Polymerase γ functions in the replicationof mitochondrial DNA • DNA replication • Properties: – Replication is semiconservative – Replication is directional: 5’ -> 3’ – Replication requires: • A template • A primer Initiation of Replication (SELF STUDY) 1. Eukaryotic origins are generally not well characterized; those of the yeast Saccharomyces cerevisiae are among the best understood. 2. Chromosomal DNA fragments (about 100bp) that are able to replicate autonomously when introduced into yeast as extracellular, circular DNA are known as ARSs (autonomously replicating sequences). 3. ARSs are yeast replicators. The three sequence elements typically found in ARSs are A, B1, and B2. 4. Initiator protein in yeasts is the multiprotein origin recognition complex (ORC), which binds to A and B1. Other replication proteins join, including one that unwinds DNA at B2. The yeast origin of replication is between regions B1 and B2. 5. DNA and histones must be doubled in each cell cycle. G1 prepares the cell for DNA replication, chromosome duplication occurs during S phase, G2 prepares for cell division, and segregation of progeny chromosomes occurs during M phase, allowing the cell to divide. 6. Cell cycle control is complex, and only outlined here. Yeasts, in which chromosomal replication is well studied, serve as a eukaryotic model organism. 7. Initiation of replication has two separate steps, controlled by cyclin-dependent kinases (Cdks) that are present throughout the cell cycle, except during G1. a. In the absence of Cdks during G1, replicator selection occurs. ORC and other proteins assemble on each replicator to form pre-replicative complexes (pre-RC). b. When cell enters S phase, Cdks are present, and activate preRCs to initiate replication. c. Cdk activity inhibits another round of pre-RC formation until the cell again enters G1, when Cdks are absent. How are ends of chromosomes Replicated? Replicating the Ends of Chromosomes 1. When the ends of chromosomes are replicated and the primers are removed from the 5’ ends, there is no adjacent DNA strand to serve as a primer, and so a single-stranded region is left at the 5’ end of the new strand. If the gap is not addressed, chromosomes would become shorter with each round of replication The problem of replicating completely a linear chromosome in eukaryotes Chapter 3 slide 72 Peter J. Russell, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. 2. Most eukaryotic chromosomes have short, species-specific sequences tandemly repeated at their telomeres. Blackburn and Greider have shown that chromosome lengths are maintained by telomerase, which adds telomere repeats without using the cell’s regular replication machinery. 3. In the ciliate Tetrahymena, the telomere repeat sequence is 5’ TTGGGG-3’ a.Telomerase, an enzyme containing both protein and RNA, binds to the terminal telomere repeat when it is single stranded, synthesizing a 3-nt sequence, TTG. b.The 3’ end of the telomerase RNA contains the sequence AAC, which binds the TTG positioning telomerase to complete its synthesis of the TTGGGG telomere repeat. c.Additional rounds of telomerase activity lengthen the chromosome by adding telomere repeats. 4. After telomerase adds telomere sequences, chromosomal replication proceeds in the usual way. Any shortening of the chromosome ends is compensated by the addition of the telomere repeats. 5. If the sequence of the telomerase RNA is mutated, telomeres will correspond to the mutant sequence, rather than the organism’s normal telomere sequence. Using an RNA template to make DNA, telomerase functions as a reverse transcriptase called TERT (telomerase reverse transcriptase). 6. Telomere length may vary, but organisms and cell types have characteristic telomere lengths. Mutants affecting telomere length have been identified, and data indicate that telomere length is genetically controlled. Shortening of telomeres eventually leads to cell death, and this may be a factor in the regulation of normal cell death. Synthesis of telomeric DNA by telomerase Peter J. Russell, iGenetics: Copyright © Pearson Education, Inc., publishing as Benjamin Cummings. How is the Genetic Information rearranged by Genetic recombination? THEORETICAL PROBLEM AND SOLUTION Homologous Recombination • I. Overview of homologous recombination (the breakage and rejoining of DNA into new combinations at sites of significant sequence similarity) • A. Requirement 1: identical or very similar sequences in the crossover region: homologous recombination does not occur between dissimilar sequences • B. Requirement 2: complementary base pairing between double-stranded DNA molecules Homologous Recombination • C. Requirement 3: heteroduplex formation: if similar complementary sequences base pair, mismatches must exist, unless the sequences are identical • D. Requirement 4: recombination enzymes must catalyze the various covalent and noncovalent rearrangement. The Holliday model for genetic recombination • • • • Figure 1-1. The Holliday model for genetic recombination. One strand of each DNA molecule is cut at the same position and then pairs with the other molecule to form a heteroduplex (region of blue paired with black). The strands are then ligated to form the Holliday junction. This DNA structure can isomerize between forms I and II. Cutting and ligating resolves the Holliday junction. Depending on the conformation of the junction, the flanking markers A, B, a, and b will recombine or remain in their original configuration. The product DNA molecules will contain heteroduplex patches. Migration of Holliday junctions • Figure 1-2. Migration of Holliday junctions. • By breaking the hydrogen bonds holding the DNAs together in front of the branch and reforming them behind, the junction migrates and extends the regions of pairing (i.e., the heteroduplexes) between the two DNAs. • The heteroduplex region is crosshatched. In the example, two mismatches, GA and CT, form in the heteroduplex region because one of the DNA molecules has a mutation in this region. Single-strand invasion model (self study) • Figure 1-3. The single-strand invasion model. A single-stranded break in one of the two DNA molecules frees a single-stranded end that invades the other DNA molecule. The gap left on the cut DNA is filled by DNA polymerase (dashed line). The displaced strand on the other DNA molecule is degraded, and the two ends are joined (arrows). Initially, a heteroduplex, represented by crosshatching, will form on only one of the two DNA molecules. Branch migration will cause another heteroduplex to form on the other DNA molecule. Isomerization can recombine the flanking DNA molecules, as in the Holliday model Double-strand break repair model • Figure 1-4. The double-strand break repair model. (1) A double-strand break in one of the two DNAs initiates the recombination event. The arrows indicate the degradation of the 5' ends at the break. (2) One 3' end, or tail, then invades the other DNA, displacing one of the strands. (3) This 3' end serves as a primer for DNA polymerase, which extends the tail until it can eventually be joined to a 5' end (black arrow). Meanwhile, the displaced strand (in black) serves as a template to fill the gap left in the first DNA (dashed lines). Two Holliday junctions form and may produce recombinant flanking DNA, depending upon how they are resolved Self study • How are RNA genomes Replicated? – Reverse transcriptase • Enzymes of General Recombination – RecA – RecBCD – RuvA,B and C