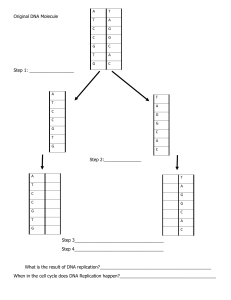
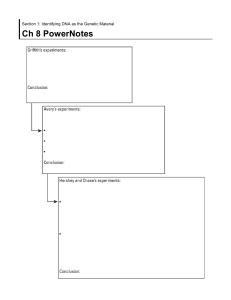
Chapter 7: Nucleic Acids 7.1: DNA Structure and Replication How does the huge DNA fit into a much smaller cell in our organisms? More than 50% of a chromosome is built of protein and the bulk of these proteins has a support and packaging role. The packaging of DNA is important because it makes sure that those excessive DNA are able to fit nicely in a cell that is many times smaller. This packaging is achieved by coiling the DNA double helix and looping it around protein beads called nucleosomes. In eukaryotic organisms, the DNA is packaged with histone proteins to create a compacted structure called a nucleosome ● nucleosomes help to supercoil the DNA, resulting in a greatly compacted structure that allows for more efficient storage ● Supercoiling helps to protect the DNA from damage and also allows chromosomes to be mobile during mitosis and meiosis. Hershey and Chase experiment: In the mid-twentieth century, scientists were still unsure if the DNA (core) or proteins (coat) contain genetic material. This is because 50% of chromosomes consist of proteins. How do we know now for sure that the DNA contains the genetic material and not proteins? Hershey and Chase conducted an experiment which aimed to prove that DNA contains the genetic code. Hershey and Chase grew viruses (T2 bacteriophage) in one of two isotopic mediums in order to radioactively label a specific viral component ○ Viruses grown in radioactive sulfur (35S) had radiolabeled proteins (sulfur is present in proteins but not DNA) ○ Viruses grown in radioactive phosphorus (32P) had radiolabeled DNA (phosphorus is present in DNA but not proteins) The viruses were then allowed to infect a bacterium (E. coli) and then the virus and bacteria were separated via centrifugation. ● The larger bacteria formed a solid people while the smaller viruses remained in the supernatant. ● The bacterial pellet was found to be radioactive when infected b y the 32P-viruses (DNA) but not the 35S-viruses ● This demonstrated that DNA, not protein was the genetic material, because DNA was transferred to the bacteria. Through the X-Ray diffraction, Rosalind Franklin discovered that DNA is a double helix. ■ DNA was purified and then fibres were stretched in a thin glass tube (to make most of the strands parallel) ■ The DNA was targeted by a X-ray beam, which was diffracted with it contacted an atom ■ The scattering pattern of the X-ray was recorded on a film and used to elucidate the details of molecular structure. From the scattering pattern produced by a DNA molecule, certain inferences could be made about its structure. ● Composition: DNA is a double-stranded molecule ● Orientation: Nitrogenous bases are closely packed together on the inside and phosphates from an outer backbone ● Shape: The DNA molecule twists at regular intervals (every 34 Angstrom) to form a helix (two strands= double helix) Now that the scientists know that DNA contains the genetic material and that it is a double helix, they still need to know how these 2 strands attach. Chargaff analyzed the composition of DNA and discovered the principle of base pairing. He did that by discovering that the number of purine bases, which are double-ring bases (adenine and guanine) always equalled the number of pyrimidines, which are single-rings bases (cytosine and thymine) and that the number of adenine equals the number of thymine and the number of cytosine equals the number of guanine. DNA replication: DNA sequencing: Sequencing is a technique by which the nucleotide base order of a DNA sequence is elucidated (typically via Sanger method) ● Dideoxynucleotides (ddNTPs) lack the 3’-hydroxyl group needed to form covalent bonds (they terminate replication) ● Four PCR mixtures are prepared – each with stocks of normal bases and one dideoxynucleotide (ddA, ddT, ddG, ddC) Whenever the dideoxynucleotide is randomly incorporated, the DNA sequence is terminated at that base position ● Because a complete PCR cycle generates millions of sequences, every base position is likely to have been terminated These sequences are separated by gel electrophoresis to determine base sequence (according to ascending sequence length) Automated machines can determine the sequence quickly if fluorescent labeling of the dideoxynucleotides has occurred DNA replication: DNA replication is a semi-conservative process that is carried out by a complex system of enzymes: Helicase: It unwinds and The double-stranded DNA by breaking the hydrogen bonds between base pairs this occurs at specific regions creating a replication fork of two strands running antiparallel directions DNA Gyrase reduces supercoiling by relaxing tension which builds up during DNA unwinding, preventing DNA breakage. SSB single stranded binding proteins: These proteins bind to the DNA atrands after they have been separated and prevent the strand from re-annealing. And prevent the single strandeed DNA from being digested by nucleases. DNA primase: It generates a small RNA primer on each of the template strands And it proves ann initiation point for DNA polymerase III which can extend a nucleotide chain but cannot start one. DNA polymerase III: DNA polymerase III attaches to the 3’ end of the primer and covalently joins the free nucleotides (a attaches to T and C with G) together in 5’ to 3’ direction And as DNA strands are antiparallel, DNA polymerase moves in opposite directions on the two strands. On the leading stand, it moves towards the replication fork and can synthesise continuously. On the lagging strand, DNA polymerase iii is moving away from the replication fork and synthesizes in pieces that are called okazaki fragments. DNA polymerase I removes the RNA primers from the lagging strand and replaces them with nucleotides. DNA ligase joins the okazaki fragments on the lagging strand to form a continuous strand. And it does this by covalently joining the sugar-phosphate backbones together with a phosphodiester bond