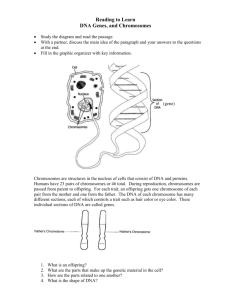
1 Principles of genetics BIOL 311 Practical 5 st level - 3st Year 1435 – 1436 2 LAB-1: Human Genetic Traits EarlobeAttachment Some scientists have reported that this trait is due to a single gene for which unattached earlobes is dominant and an attached earlobe is recessive. Other scientists have reported that this trait is probably due to several genes. Strait and hitchhiker’s thumb This trait is reportedly due to a single gene; strait thumb is dominant and hitchhiker's thumb is recessive. 3 Tongue rolling ability may be due to a single gene with the ability to roll the tongue a dominant trait and the lack of tongue rolling ability a recessive trait.However, many twins do not share the trait, so it may not be inherited. Handedness Some scientists have reported that handedness is due to a single gene with right handedness dominant and left handedness recessive. However, other scientists have reported that the interaction of two genes is responsible for this trait. 4 Freckles This trait is reportedly due to a single gene’ the presence of freckles is dominant, the absence of freckles is recessive. Curly hair Early geneticists reported that curly hair was dominant and strait hair was recessive. More recent scientists believe that more than one gene may be involved Allergies While allergic reactions are induced by things a person comes in contact with, such as dust, particular foods, and pollen, the tendency to have allergies is inherited. If a parent has allergies, there is a one in four (25%) 5 chance that their child will also have allergy problems. The risk increases if both parents have allergies. Some scientists report that there may be a genetic component to this trait while others have found no evidence to support this. Colorblindness Colorblindness is due to a recessive allele located on the X chromosome. Women have two X chromosomes, one of which usually carries the allele for normal color vision. Therefore, few women are colorblind. Men only have one X chromosome, so if they carry the allele for colorblindness, they will exhibit this trait. Thus, colorblindness is seen more frequently in men than in women. 6 LAB-2: Mitosis Mitosis:division of somatic (body) cells Interphase • Interesting things happen! 1. Cell preparing to divide 2. Genetic material doubles • Chromosome pair up! Prophase 1. Chromosomes thicken and shorten -become visible -2 chromatids joined by a centromere 2. Centrioles move to the opposite sides of the nucleus 3. Nucleolus disappears 4. Nuclear membrane disintegrate Metaphase • Chromosomes meet in the middle! 1. Chromosomes arrange at equator of cell 2. Become attached to spindle fibres by centromeres 3. Homologous chromosomes do not associate Anaphase 4. Chromosomes get pulled apart 5. Spindle fibres contract pulling chromatids to the opposite poles of the cell Telophase • Now there are two! 1. Chromosomes uncoil 2. Spindle fibres disintegrate 3. Centrioles replicate 7 4. Nuclear membrane forms 5. Cell divides Mitotic Cell Division Mitotic Cell Division 8 Root tip mitosis in Allium cepa using Feulgen squash method: Bulbs of Allium cepa are placed on the top of jars filled with tap water for root initiation. When the roots are about 1-2 cm in length, they are detached and fixed in a freshly prepared solution of 3:1 ethyle alcohol- acetic acid for 24 hours, then stored in 70% ethyle alcohol in the refrigerator until use. Steps of the experiment 1) Wash the roots in two changes of distilled water and transfer to 1N HCL, in tube, for 6-8 minutes at 60◦ C for hydrolysis. 2) Stain in leuco-basic fuchsin (Feulgen’s reagent) for 1-2 hours and prepare slides following the squash method. 3) Examine under the high power of the microscope and observe the different stage of mitosis. 4) Draw the following phases as you see them under the microscope: interphase- early prophase- prophase- metaphase- anaphasetelophase. 9 LAB-3: Estimation of mitotic index and frequency of mitotic phase The rate of mitotic and frequency of mitotic phase (MI and Ph I). The rate of mitotic division is expressed as mitotic index (MI). Which is determined by calculating the percentage of cells undergoing mitosis in a root tip as follows. Number of dividing cells MI= ------------------------------------------------- * 100 = Number of (dividing + non-dividing) cells The frequency of the mitotic phases is expressed, for each phase as mitotic phase index (Ph I). Which is the percentage of each phase out from the total dividing cells. Number of cells in stage Ph I = --------------------------------------------------*100 Number of dividing cells For the determination of MI and the frequency of mitotic phases root tips of Allium cepa are obtained from bulbs placed on tops of jars filled with water. The root tips are then fixed in 3:1 ethanol: acetic acid for 24 hours and stored in 70% ethanol. 1- Prepare, at least, four slides using the Feulgen squash technique and examine about 10 microscopical fields in each slide. 10 2- Count the number of cells at each stage of mitosis and the number of interphase cells. 3- Calculate the MI and the frequency of each mitotic phase in the following table: Phase Field number 12345678910- Interphase prophase total Slide II: Slide III: Slide VI: MI:-----------------------------------------------PI: ------------------------------------------------MI:-----------------------------------------------AI:-------------------------------------------------TI:------------------------------------------------- metaphase anaphase telophase 11 LAB-4: The Effect of colchicine on mitosis Colchicine is an alkaloid first obtained from the plant Colchicum autumnale, Colchicine is used at certain law concentrations to completely inhibit the formation of spindle fibers and cause the spiralization of chromosomes at metaphase to proceed further than normal. The inhibition of spindle fibers prevents the chromosomes to arrange themselves at the equatorial plane, and thus arresting mitosis at metaphase. The increased spiralization of chromosomes results in shorter chromosomes which are easily separable and appear scattered in the cytoplasm and the count the chromosome number. On prolonged treatments, however, the centromeres of C-metaphase chromosomes divide and C-anaphase figures are produced. Steps of the experiment 1- Root tips of Allium cepa are also used in this schedule to illustrate the effect of colchicine on mitosis. 2- Before fixation, root tips are treated with 0.05% colchicine for 4 hours at room temperature. 3- Hydrolyze in 1N HCL at 60◦ C for 8 minutes and stain with the feulgen reagent. 4- Prepare a number of slides using the squash technique. 5- Examine your preparation under the high power of the microscope and observe the effects of colchicine on the frequency of mitotic phases and draw 12 the chromosome complement of Allium cepa illustrating the morphological features of chromosomes. 13 LAB-5: Meiosis Meiosis: Meiosis: division of gametic (sex) cells • • • • • • • • • • • • • • • 4 daughter cells produced Each daughter cell has half the chromosomes of the parent 2 sets of cell division involved Meiosis 1 First division of meiosis Prophase 1: Each chromosome duplicates and remains closely associated. These are called sister chromatids. Crossing-over can occur during the latter part of this stage. Metaphase 1: Homologous chromosomes align at the equatorial plate. Anaphase 1: Homologous pairs separate with sister chromatids remaining together. Independent Assortment occurs. Telophase 1: Two daughter cells are formed with each daughter containing only one chromosome of the homologous pair. Meiosis II Second division of meiosis: Gamete formation Prophase 2: DNA does not replicate. Metaphase 2: Chromosomes align at the equatorial plate. Anaphase 2: Centromeres divide and sister chromatids migrate separately to each pole. Telophase 2: Cell division is complete. Four haploid daughter cells are obtained. Meiosis – key differences from mitosis Meiosis reduces the number of chromosomes by half. Daughter cellsdiffer from parent, and each other. Meiosis involves two divisions, Mitosis only one. Meiosis I involves: • Synapsis – homologous chromosomes pair up. Chiasmata form (crossing over of non-sister chromatids). • In Metaphase I, homologous pairs line up at metaphase plate. • In Anaphase I, sister chromatids do NOT separate. • Overall, separation of homologous pairs of chromosomes, rather than sister chromatids of individual chromosome. 14 15 Independent assortment: one of Mendel’s Laws of heredity Steps of the experiment 1- Dissct the flower and remove two anthers on an alcohol-cleaned slide. 2- Add two drops of the stain and squash using the end of a glass rod. 3- Leave for five minutes for the stain to pentratethe tissue and place the cover in position. 4- Apply gentle presure between two layers of filter paper. 5- Examine under the high power of the microscope and observe the different stages of meiosis. For the details of the method see the techniques described before. 16 Exercises Exercise 1: Identify the different stages of Meiosis in the following pictures ……………………………………………….. ……………………………………………….. ……………………………………………….. Excercise 2: Choose the right answer: ) half, homologous chromosomes, sister, diploid, haploid , four, homologous) A cell with the normal number of chromosomes (2N) is called ________________, but in meiosis the number of chromosomes is reduced by ________________ and the resulting cells (1N) is called ___________________. The 2 chromosomes of a pair are called ___________________ chromosomes and have _________________ chromatids. 17 Exercise 3: State TRUEE or FALSE: 1.-Cytokinesis is the division of: A cell ( ) A nucleus ( ) Cytoplasm. ( ) A chromosome ( ) 2.-The first stage of mitosis is: Metaphase ( ) Telophase ( ) Anaphase ( ) Prophase ( ) 3.-Meiosis results in ____ 2 haploid daughter cells ( ) 4 haploid daughter cells ( ) 2 diploid daughter cells ( ) 4 diploid daughter cells ( ) 18 LAB 6: Application of Mendel’s first law HOW TO SOLVE GENETICS PROBLEMS The FOUR STEPS TO SOLVING GENETIC PROBLEMS are shown below. Use these steps everytime to solve genetic problems. NOTES: 1) Complete Dominance Problems: Dominant traits mask recessive traits. Use the same letters, but capital for dominant & lower case for recessive. 2) Incomplete Dominance Problems: Neither trait is completely dominant, nor are there recessive traits. Use different letters for different genes, and always use capitals. STEP 1: Choose letters to represent the genes in the cross. STEP 2: Write the genotypes of the parents being crossed. STEP 3: Make a Punnett square. STEP 4: Write the percent probability for each listed genotype and phenotype appearing. 19 Monohybrid cross Exercise 1: For each genotype below, indicate whether it is heterozygous (He) or homozygous (Ho) AA _____ Ee ____ Ii _____ Mm _____ Bb _____ ff ____ Cc _____ Gg ____ kk _____ oo _____ DD _____ HH ____ LL _____ Pp _____ Jj _____ nn _____ Exercise 2: For each of the genotypes below determine what phenotypes would be possible. Purple flowers are dominant to white Brown eyes are dominant to blue eyes flowers. BB_________________ PP___________________ Bb________________ Pp___________________ bb________________ pp___________________ Bobtails in cats are recessive. Round seeds are dominant to wrinkled TT _________________ seeds Tt _________________ RR__________________ tt________________ Rr__________________ rr __________________ Exercise 3: For each phenotype below, list the genotypes (remember to use the letter of the dominant trait) Straight hair is dominant to curly Pointed heads are dominant to round heads. ___ straight _____ pointed 20 ____ curly _____ round Exercise 4: Set up the Punnet squares for each of the crosses listed below. Round seeds are dominant to wrinkled seeds. RR x rr P1 P2 What percentage of theoffspring will be round?_________________ Rr x rr P1 P2 What percent of the offspring will be round? _________________ 21 Practice with Crosses. Show all work! Exercise 5: A TT (tall) plant is crossed with a tt (short plant). What percentage of the offspring will be tall? ___________ Exercise 6: A Tt plant is crossed with a Tt plant.What percentage of the offspring will be short? ______ Exercise 7: A heterozygous round seeded plant (Rr) is crossed with a homozygous round seeded plant (RR). What percentage of the offspring will be homozygous (RR)? __________ Exercise 8: A homozygous round seeded plant is crossed with a homozygous wrinkled seeded plant.What are the genotypes of the parents? __________ x __________ What percentage of the offspring will also be homozygous? ___________ Exercise 9: a. In a pea plant that breeds true for tall, what possible gametes can be produced? Use the symbol D for tall, d for dwarf. b. In a pea plant that breeds true for dwarf, what possible gametes will be produced? c. What will be the genotype of F1 offspring from a cross between these two types? 22 d. Assuming that the allele for tall is dominant, what will be the phenotype of F1 offspring from a cross between these two types? e. What will be the probable distribution of traits in the F2 generation? (Illustrate with a Punnett square). Exercise 10: Considering that the allele for tall plants is dominant to the allele for short plants, and using T for tall and t for short, draw a Punnett square crossing a heterozygous tall plant and a short plant. (do this on paper to figure out your answers) • What percentage of the offspring are homozygous dominant? • What percentage of the offspring are heterozygous? • What percentage of the offspring are homozygous recessive? • What percentage of the offspring are tall? Why? • What percentage of the offspring are short? Why? Exercise 11: In four o'clock, red color exhibits incomplete dominance over white; when both exist together, the flowers are pink. a. In a cross between a red flower and a white one, what is the genotype of the offspring? b. What is the genotypic ratio of the F2 generation if two of the F1 from (a) are crossed? c. List the genotypes of offspring produced by a cross between the F1 generation and red parent. 23 LAB 7: Application of Mendel’s second law Exercise 1: complete the following Example : A tall green pea plant (TTGG) is crossed with a short white pea plant (ttgg). TT or Tt = tall tg tg tg tg tt = short GG or Gg = green TG TG TG TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg TtGg gg = white TG 16 Tall/Green : 0 Tall/White : 0 Short/Green : 0 Short/ White 1) A tall green pea plant (TTGg) is crossed with a tall green pea plant (TtGg) ___________ X ___________ P1 P2 24 ____ Tall/Green : ____ Tall/White : ____ Short/Green : ____ Short/ White 2) A tall green pea plant (TtGg) is crossed with a Short yellow pea plant (ttgg). ___________ X ___________ P1 P2 ____ ____ Short/Green : ____ Short/ White Tall/Green : ____ Tall/White : 3) A Heterozygous tall red flowered plant is crossed with a Homozygous short white flowered plant. ___________ X ___________ P1 P2 ____ ____ Short/Red : ____ Short/White 4) Two Heterozygous Tall, Green pea plants are crossed. ___________ X ___________ P1 P2 Tall/Red : ____Tall/White : 25 ____ Tall/Green : ____ Tall/White : ____ Short/Green : ____ Short/ White Exercise 2: Solve the followingproblems 1. Set up a punnett square using the following information: · Dominate allele for tall plants = D· Recessive allele for dwarf plants = d · Dominate allele for purple flowers = W· Recessive allele for white flowers = w Cross a homozygous dominate parent (DDWW) with a homozygous recessive parent (ddww) P1 P2 2. Using the punnett square in question #1: a. What is the probability of producing tall plants with purple flowers? Possible genotype(s)? b. What is the probability of producing dwarf plants with white flowers? Possible genotype(s)? c. What is the probability of producing tall plants with white flowers? Possible genotype(s)? d. What is the probability of producing dwarf plants with purple flowers? Possible genotype(s)? 26 3. Set up a punnett square using the following information: · Dominate allele for black fur in guinea pigs = B · Recessive allele for white fur in guinea pigs =b · Dominate allele for rough fur in guinea pigs = R · Recessive allele for smooth fur in guinea pigs = r Cross a heterozygous parent (BbRr) with a heterozygous parent (BbRr) P1 P2 4. Using the punnett square in question #3: a. What is the probability of producing guinea pigs with black, rough fur? Possible genotype(s)? b. What is the probability of producing guinea pigs with black, smooth fur? Possible genotype(s)? c. What is the probability of producing guinea pigs with white, rough fur? Possible genotype(s)? d. What is the probability of producing guinea pigs with white, smooth fur? Possible genotype(s)? 5. Set up a punnett square using the following information: · Dominate allele for purple corn kernels = R · Recessive allele for yellow corn kernels = r · Dominate allele for starchy kernels = T· Recessive allele for sweet kernals = t Cross a homozygous dominate parent with a homozygous recessive parent 27 P1 P2 6. Using the punnett square in question #5: a. What is the probability of producing purple, starchy corn kernels? Possible genotype(s)? b. What is the probability of producing yellow, starchy corn kernels? Possible genotype(s)? c. What is the probability of producing purple, sweet corn kernels? Possible genotype(s)? d. What is the probability of producing yellow, sweet corn kernels? Possible genotype(s)? 7. Set up a punnett square using the following information: · Dominate allele for normal coat color in wolves = N · Recessive allele for black coat color in wolves = n · Dominant allele for brown eyes = B· Recessive allele for blue eyes = b · Cross a heterozygous parent with a heterozygous parent P1 P2 8. Using the punnett square in question #7: 28 a. What is the probability of producing a wolf with a normal coat color with brown eyes? Possible genotype(s)? b. What is the probability of producing a wolf with a normal coat color with blue eyes? Possible genotype(s)? c. What is the probability of producing a wolf with a black coat with brown eyes? Possible genotype(s)? d. What is the probability of producing a wolf with a black coat with blue eyes? Possible genotype(s)? 29 LAB 8: What is a Model Organism? • Many aspects of biology are similar in most or all organisms • It is much easier to study particular aspects in particular organisms - for instance, genetics is easier in small organisms that breed quickly, and very difficult in humans! • The most popular model organisms have strong advantages for experimental research • They become even more useful when other scientists have already worked on them, discovering techniques, genes and other useful information. How many are there? • Many (about 80) • Mouse, rat, zebra fish, viruses, chicken, dog, hamster, maize,, etc. • Many scientists have worked on all these over the years, and shared information extensively Which are the main ones? 1) E. coli (bacterium) 2)Saccharomyces cerevisiae (yeast) 3) Arabidopsis thaliana(weed) 4) Drosophila melanogaster (fruit fly) 5) Mus musculus (mouse) 6) Homo sapiens (Man) 30 Arabidopsis thaliana (mustard plant) This is now the main model plant system for genetics. Its small genome, and the recent application of classical genetics has put it far ahead of other models of agricultural importance (tomato, tobacco, corn etc.) Its genome has been fully sequenced. Drosophila sp. ‘Fruit Fly’ - Usually the species Drosophila melanogaster – - Easily raised in lab, rapid generations, mutations easily induced and many observable mutations. - Many clues to development and genetics Caenorhabditis elegans, a nematode: usually called just C. elegans - An excellent model for understanding the genetic control of development and physiology. - C. elegans was the first multicellular organism whose genome was completely sequenced - First to show fixed cell count in body - Gave important clues on programmed cell death Saccharomyces cerevisiae, Baker’s yeast or budding yeast (used in brewing and baking) 31 Humans - Regardless of how thoroughly we may understand other animal systems, sometimes there is no alternative but to study humans directly - i.e. breast cancers - The human is the most medically analyzed and documented of any species. - We have now completely sequenced our own genome too - Over the next decade or so we will understand more about our biology than ever before! 32 LAB 9: the genetics of the fruit Fly Drosophila melanogaster • Drosophila melanogaster (Common Fruit Fly) [the name means “blackbodied fruit-lover”] is • a small, common fly found near unripe and rotted fruit. • It has been in use for over a century to study genetics and lends itself well to behavioral studies. • Thomas Hunt Morgan was the preeminent biologist studying Drosophila early in the 1900's. • Morgan was the first to discover sex-linkage and genetic recombination, which placed the small fly in the forefront of genetic research. • Due to its small size, ease of culture and short generation time, geneticists have been using Drosophila ever since. • It is one of the few organisms whose entire genome is known and many genes have been identified. Why use Drosophila? • All female flies used in controlled genetic crosses must be “virgins”. Female flies are 33 • capable of mating as early as 8 hours after emerging from the pupae stage and are polyandrous, • That is, capable of mating with several males. Once mated, females can retain viable sperm for • Several days and this will confuse the results of a subsequent controlled mating. To prevent this, • all adult flies are removed from the culture bottle about 7 hours prior to lab time, so that all • Newly hatched flies will remain virgin. VIRGIN FEMALES • All female flies used in controlled genetic crosses must be “virgins”. Female flies are • capable of mating as early as 8 hours after emerging from the pupae stage and are polyandrous, • That is, capable of mating with several males. Once mated, females can retain viable sperm for • Several days and this will confuse the results of a subsequent controlled mating. To prevent this, • all adult flies are removed from the culture bottle about 7 hours prior to lab time, so that all • Newly hatched flies will remain virgin. Life cycle of D. melanogaster • D. melanogaster exhibits complete metamorphism, meaning the life cycle includes an egg, larval (worm-like) form, pupa and finally emergence (eclosure) as a flying adult. This is the same as the well-knownmetamorphosis 34 of butterflies and many other insects. The larval stage has three instars, or molts. Life cycle by day Day 0: Female lays eggs Day 1: Eggs hatch Day 2: First instar (one day in length) Day 3: Second instar (one day in length) Day 5: Third and final instar (two days in length) Day 7: Larvae begin roaming stage. Pupariation (pupal formation) occurs 120 hours after egg laying Day 11-12: Eclosion (adults emerge from the pupa case). Females become sexually mature 8-10 hours after eclosion 35 • The time from egg to adult is temperature- dependent. The above cycle is for a temperature range of 21-23 degrees C. The higher the temperature, the faster the generation time, whereas a lower (to 18 degrees C) temperature causes a longer generation time. ·Females can lay up to 100 eggs/day. ·Virgin females are able to lay eggs; however they will be sterile and few in number. • After the eggs hatch, small larvae should be visible in the growing medium. If your media is white, look for the black area at the head of the larvae. Some dried premixed media is blue to help identify larvae however this is not a necessity and with a little patience and practice, larvae are easily seen. In addition, as the larvae feed they disrupt the smooth surface of the media and so by looking only at the surface one can tell if larvae are present. However, it is always a good idea to double check using a stereomicroscope. After the third instar, larvae will begin to migrate up the culture vial in order to pupate. 36 Sexing flies: Male and female fruit flies can be distinguished from each other in three ways: 1) Only males have a sex comb, a fringe of black bristles on the forelegs. 2) The tip of the abdomen is elongate and somewhat pointed in females and more rounded in males. 3) The abdomen of the female has seven segments with alternating dark and light segments, whereas that of the male has only five segments which are fused and black. The Drosophila Genome 3 sets of autosomes • 2 and 3 - large metacentric chromosome 37 4 - very small telocentric chromosome X/Y sex Chromosomes X is a large telocentric chromosome Sex determination Males X/Y, 2A Females X/X, 2A Y chromosome is not male determining • • • • X/0, 2A is a sterile males X/X/Y, 2A is a fertile female ratio of X to autosomes determines sex Y chromosome is needed for male fertility Genetic notation • In fruit fly genetics, the normal fly is called a "wild type" and any fly exhibiting a phenotypic mutation is called a "mutant". • Mutant flies are given names that generally denote the type of mutation the fly exhibits. For example, the mutant "ebony" has a much darker body than the wild type fly. • The above description is for a gene located on an autosome (a non-sex chromosome). Of course, fruit flies also have sex chromosomes and they contain a subset of genes as well. If the gene is located on a sex chromosome, we use a slightly different notation. Under normal diploid conditions a female fruit fly has two X chromosomes, a male has an X and a Y chromosome. Sex-linked genes are located on one of the sex chromosomes (usually the X chromosome). 38 • The genotypic notation for a mutant gene for white eye color on the X chromosome would look like: • Xw Xw = white-eyed female • Xw+Xw = wild type heterozygote female • Xw Y = white-eyed male • Xw+ Y = wild type male • Each mutation is also given a letter code. Thus, in the case of ebony, the code is a lower case e. The wild type fly is denoted by a superscript + over the mutant letter code. For example, e+ denotes a wild type fly for the ebony body trait - meaning it has normal body color (not ebony). 39 • To get use to the idea of phenotypic mutations, you will be given several strains of mutant flies. In your lab notebook, describe their morphology paying particular attention to eye color, body color, and wing shape. You should begin with a wild type fly so that you will have some basis for comparison. You may want to draw a picture. Determining the Type of Inheritance in a Cross So now you know what a wild type fly and several mutant flies look like phenotypically, but that really doesn’t tell us what their genotypes are (Why not?). In order to determine the genotypes, we must look at the ratios of phenotypes of the offspring. You will be given several bottles of fruit flies. Each bottle represents the F1 (first filial) generation of a cross set up two weeks ago. Recall that there must be a P (parental) generation before you can have a F1 generation. The idea here is to work back to the parental genotypes from the ratios of the F1 phenotypes you are given today. In the first cross you are given there is a single mutation. Anesthetize the flies using ether (your instructor will demonstrate). Observe them carefully under the dissecting microscope and record your observations in your lab notebook. The first step is to identify the sex and type of fly — mutant or wild type. Once you have determined the type of mutation, you should write out all possible crosses that could have produced these offspring. Remember, a single trait can be inherited on an autosome or on a sex chromosome. Is your mutation on an autosome or sex chromosome? How would you tell? 40 1. Observe the wild type flies carefully and complete the following table. 1-From your observations, draw each of these stages in Drosophila melanogaster development. Larva Pupa Egg Characteristics of Fruit flies Description of: Antenna Eye color Eye shape Body color Wing shape Wing morphology Adult Wildtype (normal) Present Red Normal Normal Normal Normal mutant Absent Bright, purple, sepia, white Bar, eyeless, lobe, star Black, ebony, sable, tan, yellow Apterous, miniature, vestigial Curly, curved, dumpy, scalloped Your fly 41 Fill the table below (Tab1) after observing under dissecting microscopes: Table 1. External characteristics observed in male and female fruit flies. Characteristic observed Male Female Overall size Color of abdomen Shape of abdomen tip Are sex combs present? What are the possible crosses that could have produced this pattern of offspring? If the mutation was inherited as a simple autosomal recessive, then we might suspect the parental generation was either: • w w X w+w+ • w+w X w+w • w+w X w w • w+w X w+w+ Remember a w represents the mutant trait and a w+ represents the wild type. 42 Which of these 4 possibilities would give you the same approximate pattern of offspring you actually observed? Which of these 4 possibilities can be ruled out? Take parental cross 1 above for example. In a cross between a homozygous wild type and a homozygous mutant type you would expect to get all heterozygous wild type offspring. Since heterozygous offspring all appear phenotypically to be wild type flies, you could now rule this cross out because you actually observed 30 mutant flies in you F1 population. • Similarly, if this mutation was inherited as a sex-linked trait, you might predict one of the following parental crosses: • XwXw x XwY • Xw+Xw+ x Xw+Y • Xw+Xw x XwY • Xw+Xw x Xw+Y • Again you can do each cross and quickly rule out the ones that do not fit with your observed pattern of offspring. Assume you have ruled out all crosses but a parental cross between two heterozygotes for a trait located on an autosome (i.e. a parental cross of w+w x w+w). In this cross you would expect to see a phenotypic ratio of 3 wild type for every 1 mutant type regardless of the sex of the fly. Does this expected outcome fit with your observed data? It sure looks like it, but how can you be sure? • To test your hypothesis that the observed ratio of 70:30 is the same as the expected ratio of 3:1, we can use a statistic called the Chi square statistic Chi square statistic 43 • • • Calculate the chi square statistic x2 by completing the following steps: For each observed number in the table subtract the corresponding expected number (O — E). Square the difference [ (O —E)2 ]. Divide the squares obtained for each cell in the table by the expected number for that cell [ (O - E)2 / E ]. Sum all the values for (O - E)2 / E. This is the chi square statistic. • Having now obtained our chi square statistic x2 = 1.31, we look up in a table of the Chi • • Square X2 distribution the probability attached to it. Before we can do this, however, we need to know the degrees of freedom. When a comparison is made between one sample and another, a simple rule is that the degrees of freedom equal (number of columns minus one) x (number of rows minus one) not counting the totals for rows or columns. For our data this gives (2-1) x (2-1) = 1. Entering the Chi square distribution table with 1 degree of freedom and reading along the row we find our value of x2 (1.31) lies between 0.455 and 2.706. The corresponding probability is 0.5<P<0.1. 44 • This is well below the conventionally accepted significance level of 0.05 or 5%, so the null hypothesis that the two distributions are the same is verified. In other words, when the computed x2 statistic exceeds the critical value in the table for a 0.05 probability level, then we can reject the null hypothesis of equal distributions. Since our x 2 statistic (1.31) did not exceed the critical value for 0.05 probability level (3.841) we can accept the null hypothesis that a ratio of 70:30 is the same as a 75:25 ratio (within 5% error). 45 LAB 10: Extraction of DNA from Fish Fins Objectives: To isolate DNA from fish fins. 1. To obtain the purified form of DNA which can be further used for molecular analysis. 2. To understand the principle and process of DNA extraction. Theory: Deoxyribonucleic acid (DNA) is a complex nucleic acid containing the genetic code with the instructions for the development and functioning of all known living organisms, with the exception of some viruses. The DNA segments that carry this genetic information are called genes. DNA is transcribed into RNA which is then used as a template in the synthesis of proteins. DNA isolation refers to the process of extracting DNA from a cell in a pure form. The extraction of DNA is an important preliminary step in which purified DNA is obtained from other cellular components such as proteins, RNA and lipids. DNA can be isolated from any nucleated cell from diverse sources, both living and dead, such as whole blood, hair, sperm, bones, nails, tissues, faeces, shed feathers, egg shells, saliva, epithelial cells, urine, bacteria, animal tissues or plants. Isolation of DNA is required for a variety of applications in science, medicine and forensics. For example, scientists introduce DNA into the cells of animals or plants for SNP analysis, DNA methylation analysis, copy number variation (CNV) or comparative genomic hybridization (CGH) analysis, Southern Blotting, PCR etc. In medicine, DNA isolation is helpful for diagnostic 46 purposes. Furthermore, DNA extraction is an important tool in forensic science for the identification of individuals (for example rapists, thieves, accident, or war victims), paternity determination, body identification etc. There are a number of different procedures for the preparation of genomic DNA. They all start with some form of cell lyses, followed by de-proteination and recovery of DNA. The main differences between various approaches lie in the extent of de-proteination and in molecular weight of the DNA produced. Factors affecting the methods of DNA isolation are the age, source and size of the sample. The presence of proteins, lipids, polysaccharides etc. during DNA preparation can interfere with DNA analysis methods by reducing the quality of DNA. The extraction methods to efficiently purify DNA from various sources have to be adapted depending on factors such as sample size, the freshness of the sample, and the biochemical content of the cells from which DNA is being extracted. The isolation method must vary depending on the size of sample. The freshness of the sample also affects the extraction technique. Extraction methods are also variable according to the biochemical content of the source cells. For example, in the case of bacteria, the main biochemicals present in a cell extract are protein, DNA and RNA. Therefore, phenol extraction or protease treatment, followed by removal of RNA with ribonuclease, leaves a pure DNA sample. These treatments may not be sufficient if the cells also contain significant quantities of other biochemicals. Plant tissues are particularly difficult in this respect as they often contain large amounts of carbohydrates that are not removed by phenol extraction. 47 DNA molecule Fish fins are an excellent and reliable source of high quality DNA with a number of advantages. In most cases DNA is obtained from the collection of blood from the fish. In the case of rare or endangered species, this method is not advisable. Therefore, fin tissue is often used for DNA extraction because sampling is relatively fast, logistically simple, and is non-lethal. A small piece of tissue provides plenty of DNA for PCR and restriction digests. Most importantly, collection of fish fin sample does not harm the fish. This is a very important consideration especially relevant regarding rare and the ornamental species. Fish fin samples are used for DNA-based studies on genetic diversity, mating systems and parentage determination of fish populations with minimal disturbance. Principle: The isolation of DNA usually begins with the lysis of cells or tissues inorder to destroy the protein structures and allows the release of nucleic acids from the nucleus. Lysis is carried out in a lysis solution containing important ingredients: sodium chloride which provides an osmotic shock to the cells; Tris HCl, which is a buffer to retain constant pH; EDTA, which sequesters the divalent metal ions that is required for nuclease activity and thereby inhibiting its action; a detergent, usually SDS, which disrupts the cell membrane and nuclear envelope, causing the cells to burst 48 open and release their DNA. The DNA is still rapped very tightly around histone proteins. Proteinase K (a serine protease) is the common enzyme used in DNA extraction that cuts apart the histones to free the DNA and finally results in the breakdown of cells and dissolving of membranes. The nucleic acids are then purified from the protein- nucleic acid complex by phase extraction with a mixture of organic solvents namely Phenol, Chloroform and Isoamyl alcohol in a ratio of 25:24:1. Phenol dissociates proteins from DNA. Chloroform denatures the proteins and lipids and helps to maintain the separation of the organic and aqueous phase. It also makes the DNA less soluble in the phenol, thus reducing losses to the organic phase. Isoamyl alcohol is often added to prevent foaming. At pH 7-8, the DNA partitions to the aqueous phase while the protein is denatured and extracted into water- immiscible organic phase, which is separated from the nucleic acid containing aqueous phase by centrifugation. When large amount of protein is present, it forms a white precipitate between the organic and aqueous phase. The DNA is then precipitated with cold ethanol or isopropanol after adjustment with 3M sodium acetate and then centrifuging. The DNA is insoluble in the alcohol and will come out of solution, and the alcohol serves as a wash to remove the residual salts. The alcohol is then removed, and DNA is stored in a biological buffer, like TE (Tris-EDTA) buffer. Contaminating RNA in the DNA sample can eliminated by digestion with an RNase. 49 However, since shearing forces are generated at every step, the resulting DNA molecules in the final preparation rarely exceed 100-150 kb in length. DNA of this size is adequate for Southern analysis on standard agarose gels, as a template in PCRs, and for the construction of genomic DNA libraries in bacteriophage λ vectors. Materials Required: Fish fins. Scalpel. Forceps. Mortar and pestle. Centrifuge. Vials. Micropipette and Pipette tips. Water bath. Filter paper / Blotting paper. 50 Reagents: Extraction Buffer 200ML 1M Tris Cl (pH 8.0) - 2.5ml. 0.5M EDTA (pH 8.0) - 50ml. Pancreatic RNAase - 5mg. Proteniase K - (Required). TE Buffer: Tris base - 10mM. EDTA - 1mM. Phenol: Chloroform: Isoamylalcohol (50ml)- (25:24:1). Chloroform: Isoamylalcohol (50Ml) - (48:2). 70%, 100% Ethanol (Ice cold). Sodium Acetate 3M. Procedure: 1. Pluck out 20-100 mg of fish fin using a sterile scalpel and a forceps. 2. Homogenize the fins in 600μl of extraction buffer using sterile mortar and pestle. 3. Collect the paste into 1.5ml Micro Centrifuge Tube(MCT). 4. Incubate at 37°C for 1 hr in a water bath. 5. Again incubate at 55°C for 1 hour in the water bath. 6. Centrifuge at 5000 rpm for 10 minutes. 7. Remove the supernatant put it in a new MCT and add equal volume of Phenol: Chloroform: Isoamylalcohol (25:24:1). Mix slowly and thoroughly by repeated inversion of the MCT. 8. Centrifuge at 12000 rpm for 10 minutes and collect the top aqueous layer in a fresh MCT. Do not disturb the intermediate layer. 9. Add Chloroform: Isoamylalcohol in the ratio 24:1. Mix slowly and thoroughly by inversion of the tube. 51 10. Centrifuge the tube at 12000 rpm for 10 minutes. 11. Collect the aqueous layer into a fresh MCT and add 0.1 volume of 3M sodium acetate and equal volume of ice cold ethanol (100%). Mix the solution thoroughly until the DNA pellet is obtained. (Now you can see the DNA clumps in the tube.) 12. Incubate the MCT at -20°C for 1 hour. 13. Take out the tube and centrifuge at 1000 rpm for 10 minutes and decant the supernatant (Ethanol). 14. Wash the tube with 70% ethanol, by centrifuging at 1000 rpm for 10 minutes and decant the ethanol carefully without losing the pellet. (If you find difficulty use pipette for removing.) 15. Air dry the DNA pellets at room temperature by keeping the tube opened and also blot using filter paper / blotting paper. 16. Resuspend the pellet in 50-100μl of TE buffer or sterile triple distilled water.