Mitochondrial DNA products among RAPD profiles
advertisement

Molecular Ecology (1995) 4, 441-447
Mitochondrial DNA products among RAPD profiles
are frequent and strongly differentiated between races
of Douglas-fir
]. E. AAGAARD, S. S. VOLLMER, F. C. SORENSEN* and S. H. STRAUSS Department of Forest Science, Oregon State University, Corvallis, OR 97331 and *Forestry Sciences Lab, USDA Forest Service, Corvallis OR 97331, USA
Abstract
Racial differentiation and genetic variability were studied between and within the
coastal, north interior, and south interior races of Douglas-fir using RAPD and allozyme
markers. Nearly half of all RAPD bands scored (13: 45%) were found to be amplified from
mitochondrial DNA. They exhibited maternal inheritance among hybrids and back­
crosses between the races, and were much more highly differen~ated (G 5T =0.62 for hap­
lotype frequencies) than were allozymes (Gs-r = 0.26). No evidence of hybridization or
introgression was detected where the coastal and interior races come into proximity in
central Oregon.
Keywords: allozymes, Douglas-fir, genetic differentiation, mitochondrial DNA, RAPD, repetitive
DNA
Received 5 October 1994; second revision received 3 May 1995; accepted 11 May 1995
Introduction
Studies of population genetic structure and genetic varia­
tion using random amplified polymorphic DNA (RAPDs)
are becoming increasingly common. For many uses, the
limitations of RAPDs as a consequence of dominance and
low reproducibility are clearly outweighed by the essen­
tially limitless number of genome markers (reviewed by
Newbury & Ford-Lloyd 1993). As empirical and theoreti­
cal approaches to the treatment of RAPDs in population
genetic studies develop (Lynch & Milligan 1994), concur­
rent studies are needed that compare RAPD diversity to
the well-established and far more prevalent studies using
allozymes (for reviews see Hamrick & Godt 1990).
Population stUdies which have considered both RAPDs
and allozymes show broadly consistent patterns; similar
phylogenetic relationships are elucidated by both molecu­
lar techniques, though RAPDs tend to emphasize greater
differentiation among populations than allozymes (e.g.
Mosseler et al. 1992; Puterka et al. 1993; Heun et al. 1994;
Moreau et al. 1994; Peakall et al. 1995). Possible explana­
tions for the increased differentiation resolved by RAPDs
include selective constraints on coding DNA (reviewed by
Altukhov 1991) and the limitations of conventional starch
Correspondence: Steve Strauss. Fax: (503) 737 1393. E-mail:
strauss@fsl.orst.edu
gel electrophoresis to detect amino acid substitutions
(Neel 1984). In addition, because RAPDs by design ampli­
fy sections of the genome containing repetitive DNA,
RAPD frequencies may reflect processes of genomic
turnover such as nonreciprocal exchange (reviewed by
Dover & Tautz 1986) which accelerate the spread of muta­
tions through populations (Stephan 1989). Finally, RAPDs
can amplify organelle polymorphisms (Lorenz et al. 1994),
which may show distinct patterns of differentiation from
nuclear genes (e.g. Strauss et al. 1993; Hong et al. 1993).
Allozyme variation in Douglas-fir (Pseudotsuga men­
ziesii (Mirb.) Franco) has been well characterized, includ­
ing a range-wide study of populations throughout west­
em North America (Li & Adams 1989). Patterns of
allozyme variation are similar to those of other conifers,
exhibiting limited and primarily quantitative differentia­
tion in allele frequencies between populatio~ (Hamrick &
Godt 1990). In contrast, a preliminary screen with several
RAPD primers suggested large and qualitative differences
in RAPD markers might exist between the coastal variety
(var. menziesii) and interior variety (var. glauca) of Douglas­
fir. These varieties have been recognized based on cone
and needle morphology, growth characteristics (Rehfeldt
1977), and terpene chemistry (von Rudloff 1972).
Allozyme studies have further substantiated varietal dif­
ferences, identifying the coastal and interior variety, as
well as two distinct racial groups within the interior vari­
442
J. E. A A G A A R D
et al.
ety (Li & Adams 1989). Despite a long history of differen­
tiation that may extend back as far as the Miocene (13 mil­
lion years ago: reviewed by Critchfield 1984), the varieties
readily cross, both under controlled pollination and in the
wild (Rehfeldt 1977).
Materials and methods
Seeds from the three major divisions (races) of Douglas-fir
identified by the allozyme study of Li & Adams (1989)
were sampled from throughout the Western United States
(Fig. 1). To sample the coastal variety and north interior
race of the interior variety, three seeds (one seed per
maternal tree) were selected from each of 20 sources span­
ning an east-west transect which crossed the putative
transition zone between the two varieties in central
Oregon (mean latitude 44°45'; see Sorensen 1979 for
details). Seeds from the south interior race of the interior
variety representing nine distinct geographical locations
throughout Arizona, New Mexico, and Utah (2-3 seeds
per source) were obtained from bulked seed collections
stored at the Lucky Peak Nursery (Boise National Forest,
Idaho).
Seeds were soaked overnight in water, embryos
removed, and embryo DNA extracted according to a mod­
ification of the CfAB protocol of Wagner et al. (1987) in
which the DNA was further purified by four phenol/ chlo­
roform/isoamyl alcohol (25:24:1) extractions and a final
ethanol precipitation. Reagents used in the RAPD amplifi­
cations (final volume 25 14L) were 100-,uM dNTPs, 0.2-14M
primer, 1.8-mM Mg2•, 1 unit of Taq polymerase, and 2 ng
of template DNA. Increasing template DNA up to 25 ng
had little effect on RAPD profiles of several primers test­
ed. The thermocycle protocol consisted of denaturation for
one minute at 93 °C, annealing at 37 oc for 1 min, and
extension at 72 oc for 2 min repeated over 45 complete
cycles using an MJ Research thermocycler (model PTC­
1()()....60 ).
Pre-screening of 105 random decamer primers (UBC,
Vancouver, British Columbia) revealed 10 primers produc­
ing 41 bands that were chosen for further study (UBC
primers 266: CCACfCACCG, 268: AGGCCGCfTA, 275:
CCGGGCAAGC, 300: GGCfAGGGCG, 409: TAGGC­
GGCGG, 414: AAGGCACCAG, 419: TACGTGCCCG, 428:
GGCTGCGGTA, 438: AGACGGCCGG, and 446:
GCCAGCGTTC). These primers were chosen on the basis
of 100% reproducibility of selected bands between repli­
cates and ability to amplify DNA from all seed samples
(seeds were in freezer storage for 5-20 years). Negative
controls included in primer screening showed no amplifi­
cation artefacts and established that all bands resulted
from template DNA.
Initial scoring of RAPD phenotypes assumed bands of
like phenotype (mobility, intensity) result from dominant
alleles at independent nuclear loci, and are in
Hardy-Weinberg equilibrium within populations.
Dominance of most RAPD bands is widely known
(Waugh & Powell 1992; Heun & Helentjaris 1993). With
few exceptions conifers, including Douglas-fir (e.g. Moran
& Adams 1989), show only minor and transient depar­
tures from Hardy-Weinberg proportions. However,
because RAPD bands amplified from total genomic DNA
of plants may also be of cytoplasmic origin (Lorenz et al.
1994), we examined inheritance of several bands in inter­
racial crosses of Douglas-fir made previously (unpub­
lished data). Parents studied were all F1 hybrids between
the north interior and coastal races, and their progeny
were the result of crossing to another hybrid or 'back­
crossing' to each race. In addition, we analysed the mater­
nally derived, haploid conifer megagametophytes of the
seed from these crosses. Diploid embryos and their associ­
ated haploid megagametophytes were studied in 44 prog­
eny (four from each of 11 crosses; four F1 hybrids and
seven back-crosses). Because chloroplast DNA is paternal­
ly inherited (Neale et al. 1986) in Douglas-fir and shows
extremely low genomic diversity within or among races
(Ali et al. 1991), and mitochondrial DNA is maternally
inherited (Marshall & Neale 1992), the genomic origin of
uniparentally inherited bands could be unambiguously
defined.
RAPD data were analysed for genetic differentiation
and diversity. Calculations were made for maternally
inherited markers based on haplotype frequencies at a
single locus as done previously (Strauss et al. 1993), and
for the remaining RAPD markers assuming nuclear origin
and dominance. For most calculations, individuals were
pooled into one of three racial groups disregarding taxo­
nomic hierarchy (coastal, north interior, or south interior
races: Li & Adams 1989). Only those loci showing poly­
morphism among one or more races (0.99 criterion) were
used in the analysis. Null allele frequencies were correct­
ed for sampling (Lynch & Milligan 1994). Parameters cal­
culated included Nei's (1973) gene diversity statistics cor­
rected for sample size and population number, and Nei's
(1978) unbiased genetic distances (D) between racial
groups and between seed sources (locations) of the coastal
and north interior races. The latter was calculated. solely to
illustrate the sharpness and location of the racial bound­
ary, and all RAPD bands were included regardless of their
inheritance pattern (i.e. all bands were treated as nuclear
loci). Measurements of diversity included gene diversity
(Hs) and both mean (n) and effective mean (n.) number of
alleles per locus. The effective mean number of alleles per
locus weights an allele according to its frequency to pro­
vide a measure of allelic evenness (Hedrick 1983 ). Genetic
differentiation between races using Nei's (1973) Gsr is
nearly equivalent to the familiar Fsr of Wright (Wright
1965) and the two may be treated interchangeably in most
RAPD DIFFERENTIATION BETWEEN RACES OF DOUGLAS-FIR
443
cases (Nei 1977). The computer program GeneStat-PC 3.3
(Lewis 1994) was used for all calculations.
To compare genetic differentiation and diversity
between RAPDs and allozymes, we re-analysed data from
Li &: Adams' (1989) range-wide allozyme study of
Douglas-fir (provided by P. Li and W. T. Adams, personal
communication). The 29 seed sources of our RAPD study
were geographically matched with 31 populations from Li
&: Adams' study (roughly 15 maternal trees per popula­
tion; see Li &: Adams 1989 for details) and for quantitative
analysis pooled into one of three racial groups based on Li
and Adams' cluster diagram (Fig. 1; coastal: 16-23; north
interior: 25-28, 65-69, 73, 75, 78; and south interior 84-87,
97-102).
Results
RAPDs
Fig. 1 Locations of Douglas-fir seed sources and populations
used for the RAPD and allozyme analyses. Seed sources used in
the RAPD study come from 29 locations representing the coastal
(C), north interior (N), and south interior (S) races. The approxi­
mate locations of the 31 populations taken from Li &: Adams'
(1988) allozyme study are shown as numerals. All populations
and seed sources are located within the states of Arizona (AZ),
Idaho (ID), New Mexico (NM), Oregon (OR), and Utah (UT).
I
2
G)
(,)
c
<U
en oe
i5
(,)
;:
G)
06
c
G)
"a;
04
_en
z
CO I I I I I
--. - . ­
- - --- -.
North
lnlrrlor
- ·-·--·
- - -- ---____
- ·- ---___ ...,. ...---·__ _­­
--... - -... ···--­ ------------
02 Transect Position - West to East
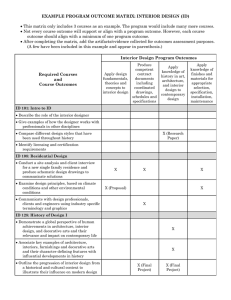
Fig. 2 Nei's (1978) genetic distances between the eight coastal
and 12 north interior sample locations (seed sources). Genetic dis­
tances were calculated in a pairwise manner between the source
furthest east and all other sources (A) and between the source
furthest west and all other sources (B). Also shown are RAPD
phenotypes from primer UBC 275 exh.tbiting strong race-speci­
fidty for three coastal and four north interior seed embryos (three
replicates per embryo).
Of the 41 fragments analysed in the RAPD study, 29
showed variability between or within the coastal, north
interior, and south interior races of Douglas-fir. Of these,
10 exhibited race specificity. All of the fragments agreed as
to the position of the racial boundary between the coastal
and north interior races (coastal and interior variety) in
central Oregon; the break occurred between the lowest
elevation Santiam population (915 m) and the Grizzly
population (1555 m), which are separated by 55 km
(described in Sorensen 1979). In no cases were unique
combinations of fragments found that might indicate
introgression or interracial hybridization (Fig. 2).
Based on strict maternal inheritance, 13 RAPD markers
(45%) generated from seven primers (UBC primers 266,
268, 275, 409, 419, 428 and 438) were of mitochondrial ori­
gin (data not shown). Of the remaining bands, five
showed clear biparental inheritance and 11 were ambigu­
ous in the crosses available. Using the 13 maternally inher­
ited markers we identified nine distinct haplotypes
among the coastal, north interior, and south interior races.
Only a single haplotype was found in the coastal race
whereas six haplotypes were found in the south interior
race. Gene diversity ranged from a high of 0.76 in the
south interior race to zero in the coastal race (Table 1).
Greater than 60% of the observed haplotype diversity
resulted from differentiation between populations (Gsr
- 0.62; Table 1).
Differentiation between races of Douglas-fir was also
extensive when the remaining 16 polymorphic RAPD
bands were considered. Differentiation between races
accounted for more than 70% of the total diversity
observed (Gsr • 0.73; Table 1). Paired comparisons of the
races showed that differentiation between the coastal and
either the north interior or south interior race was similar
to the total differentiation between all races (Gsr - 0.76 and
444
J.
E. AAGAARD et al.
0.82, respectively). Differentiation between the interior
races was less (Gsr ... 0.52). Similarly, genetic distance was
substantial between the coastal and either interior race,
but modest between the interior races; mean genetic dis­
tance ranged from a high of 1.01 (coastal x south interior)
to a low of 0.24 (north interior x south interior).
ranging from a high of 0.10 (coastal x south interior) to a
low of 0.04 (north interior x south interior).
Surprisingly, an inverse pattern of genic and allelic
diversity between races was found for allozymes relative
to RAPDs. Allozyme gene diversity was highest in the
coastal race (0.21) and lowest in south interior race (0.09),
whereas the reverse was true for RAPDs (Table 1). Mean
and effective mean number of alleles per locus paralleled
these trends with the greatest values in the coastal race
(2.79 and 1.36, respectively) and the lowest in the south
interior race (2.53 and 1.15, respectively).
Allozymes
The degree and pattern of racial differentiation and diver­
sity based on allozymes differed dramatically from those
of RAPDs (Table 1). Of 20 loci analysed by Li & Adams
(1988), 19loci exhibited polymorphism between or within
races and five loci showed race-specific alleles. Of the total
genetic variability found for allozyme data, less than 30%
(Gsr,. 0.26) was attributed to racial differentiation. Also in
contrast to results from RAPDs, the coastal and north inte­
rior races had a similar amount of differentiation as that
found between north interior and south interior races (G5T
- 0.21 and 0.19, respectively), whereas coastal and south
interior races were nearly twice as differentiated (Gsr
'"'0.36). Genetic distances concurred with these patterns,
Discussion
We found several striking contrasts between estimates of
racial differentiation and diversity with RAPD and
allozyme markers. Most notable was the much greater dif­
ferentiation reflected in Gsr values found for RAPDs
(Table 1). This may result from several distinct causes.
First, because conventional starch gel electrophoresis fails
to detect many allozymes distinct in their amino acid
sequence, some allozyme differentiation often remains
Table 1 Racial differentiation, genetic distances, and genic and allelic diversity based on RAPDs and allozymes between or within races
of the coastal and interior varieties of Douglas-fir. Standard errors (SE) are calculated for Gsr from variance among locus-specific estimates.
Calculations of differentiation and diversity for mitochondrial RAPDs are based on nine mitochondrial haplotypes identified from 13
maternally inherited RAPD markers
Differentiation (Gsr) between races
coastal x north interior x south interior
coastal x north interior
coastal x south interior
north interior x south interior
Nei's genetic distance (D) between races
coastal x north interior
coastal x south interior
north interior x south interior
Gene diversity (Hs) calculated within races
coastal
north interior
south interior
Mean gene diversity (H5) averaged over races
Total gene diversity (HT)
Observed (n) and effective (nJ number of
alleles per locus within races•
coastal
north interior
south interior
Average over races•
RAPDs
Allozymes
Mitochondrial
RAPDs
0.73 ±0.09
0.76 ±0.12
0.82±0.09
0.52 ±0.11
0.26±0.03
0.21 ±0.03
0.36±0.04
0.19 ±0.03
0.62
0.89
0.45
0.39
0.49
1.01
0.24
0.06
0.10
0.04
0.07±0.04
0.15±0.05
0.18 ±0.05
0.13 ±0.03
0.49 ±0.05
0.21 ±0.04
0.17±0.05
0.09±0.04
0.16 ±0.03
0.21 ±0.05
0.00
0.11
0.76
0.29
0.76
n
n,
n
n,
n
n,
1.25 ±0.11
1.44±0.13
1.44 ±0.13
1.38 ±0.06
1.12 ±0.07
1.25 ±0.09
1.29 ±0.09
1.22 ±0.05
2.79 ±0.22
2.95 ±0.19
2.53 ±0.21
2.75 ±0.12
1.36 ± 0.10
1.31 ±0.10
1.15 ±0.08
1.27 ±0.06
1
2
6
3
1.13
3.86
2.00
•for mitochondrial RAPDs, the number of alleles refers to the number of mitochondrial haplotypes.
RAPD DIFFERENTIATION BETWEEN RACES OF DOUGLAS-FIR
cryptic (Neel 1984). RAPDs, on the other hand, are theo­
retically sensitive to many kinds of mutations, including
those that affect sequences of priming sites and their rela­
tive positions. Second, allozymes represent variation in
functional gene products (enzymes) whereas the majority
of RAPD variation is likely to result from noncoding DNA.
Thus, natural selection may act more strongly on
allozymes. Balancing selection has been proposed as an
agent for maintaining allozyme polymorphism (Altukhov
1991; Karl & Avise 1992); it may retain allozyme frequen­
cies at similar values between isolated populations or
racial groups while neutral loci differentiate due to genet­
ic drift. Third, because RAPD products represent portions
of the genome that contain repetitive DNA, specifically
regions bounded by complementary inverted repeats,
their evolution may be fundamentally distinct from genes
encoding allozymes. Processes such as unequal exchange
may allow repetitive DNA to differentiate between isolat­
ed populations in a concerted and rapid manner termed
'molecular drive' (Dover & Tautz 1986; Stephan 1989;
Cabot et al. 1993). Complementary inverted repeats are
nearly universal footprints of transposable elements,
which on evolutionary time scales can invade genomes
rapidly (Charlesworth et al. 1994). Thus, it would not be
surprising if some types of RAPD variants could increase
in frequency more rapidly than allozymes.
The mitochondrial genome also contributed strongly to
the RAPD phenotypes we observed. RAPDs may readily
amplify mitochondrial DNA fragments from total genom­
ic DNA because of the common occurrence of comple­
mentary inverted repeats in plant mitochondrial genornes
(e.g. Thormann & Osborn 1992; Lorenz et al. 1994). Th~
mitochondrial genome exhibits rapid changes of structure
(for reviews see Sederoff 1987 & Schuster & Brennicke
1994) and has been found to be much more highly differ­
entiated among populations than allozymes (Strauss et al.
1993). The large number of variable mitochondrial RAPD
bands found, and the diverse primers that amplified them,
suggest that the mitochondrial genome of Douglas-fir is
large, complex, and dynamic.
Surprisingly, allelic and genic diversities between races
were inverted for RAPDs relative to that for allozymes. The
coastal race was more diverse for allozymes than was the
south interior race, but less diverse for RAPDs (Table 1). We
believe that this results, at least in part, from pooling seed
samples of the south interior race over several populations
derived from isolated mountain ranges in three states
(Fig. 1). Inter-population differentiation for allozymes is
much stronger in this region than for the coastal or north
interior regions (Li & Adams 1989). Based on the present
results, this differentiation is likely to be even greater for
RAPDs. Pooling over locations in the south interior race
may therefore enhance diversity for RAPDs greatly; RAPD
diversity within populations of the south interior race
445
might still be less than that for the coastal race, as allozyme
studies predict (Li & Adams 1989).
The sharp boundary found between the coastal and
north interior races in central Oregon based on RAPDs
(Fig. 2) agrees with most results from other genetic analy­
ses. Li & Adams' (1989) allozyme study also demonstrated
strong racial differentiation, but did indicate that a narrow
zone of transition might exist between the races in central
Oregon. Sorensen's (1979) nursery study found a gradual
transitional zone for some growth traits but strong differ­
entiation for others along the same transect that we sam­
pled in our RAPD study. Terpene composition agreed
most strongly with our RAPD data, suggesting a pro­
nounced racial boundary in nearby populations (Zavarin
& Snajberk 1973). Because intervarietal hybrids are known
to readily occur in zones of contact (Rehfeldt 1977) and
gene flow through pollen and seed dispersal is extensive
in conifers (reviewed by Adams 1992), this suggests that
the close proximity of the varieties in central Oregon may
be a geologically recent event.
In sum, it appears that RAPDs provide considerably
greater sensitivity than allozymes for the detection Qf
genetic differentiation, at least for long-isolated gene pools
such as races of Douglas-fir. Mitochondrial DNA has pro­
vided a major portion of this strong differentiation.
Additional research is needed to determine whether
nuclear DNA amplified by RAPDs is also more highly dif­
ferentiated than are allozymes. Although substantial dif­
ferentiation was also observed for the 16 bands that did
not show maternal inheritance, many of these could also
be of mitochondrial origin in the crosses studied.
Regardless of origin, however, RAPDs provide a powerful
means for racial identification.
Acknowledgements
We wish to thank Peng Li and Tom Adams for providing Douglas­
fir allozyme data, Brian Ferguson and Dick Thatcher of the US
Forest Service for making available the southwest seed collec­
tions, Amy Brunner and Konstantin Krutovski for help with writ­
ing and organization, and three anonymous reviewers for their
helpful comments. ]EA received financial support in part from
NSF grants DEB 9300083 and BSR 895702.
References
Adams WT (1992) Gene dispersal within forest tree populations.
New Forests, 6, 217-240.
Ali IF, Neale DB, Marshall KA (1991) Chloroplast DNA restriction
fragment length polymorphism in Sequoill sempervirens D. Don
End!., Pseudotsuga menziesii (Mirb.) Franco, Calocedrus ckcur­
mzs (Torr.), and Pinus taeda L. Theoretical and Applied Genetics,
81,83-89.
Altukhov YP (1991) The role of balancing selection and overdom­
inance in maintaining allozyme polymorphism. Genetica, 85,
79-90.
446
J.
E. AAGAARD et al.
Cabot EL, Doshi P, Wu ML. Wu 0 (1993) Population genetics of
tandem repeats in centromeric heterochromatin - unequal
crossing over and chromosomal divergence at the responder
locus of Drosophilll melmwgaster. Gtnetics, 135, 477-487.
Charlesworth 8, Sniegowski P, Stephan W (1994) The evolution­
ary dynamics of repetitive DNA in eukaryotes. Nature, 371,
215-220.
Critchfield WB {1984) Impact of the Pleistocene on the genetic
structure of North American conifers. In: Proceedings of the 8th
North Amerian For~l Biology Workshop (ed. Lanner RM), pp.
70-118. Utah State University, Logan, UT.
Dover GA. Tautz D (1986) Conservation and divergence in multi­
gene families: alternatives to selection and drift. Philosophical
TranSRcticns of the Royal Society, 312, 275-289.
Hamrick JL, Godt M] (1990) Allozyme diversity in plant species.
In: Plant Population Genetics, Breeding, and Genetic Resourc~
(eds BrownAHD, Clegg MT, Kahler AL, Weir B S), pp. 43-63.
Sinauer Associates, Sunderland, MA.
Hedrick PW (1983) Measures of genetic variation. In: Genetics of
Populations {Hedrick PW), pp. 39-81. Science Books
International, Boston.
Heun M, Helentjaris T (1993) Inheritance of RAPDs in F1 hybrids
of com. Theoretical and Applied Genetics, 85, 961-968.
Heun M, Murphy JP, Phillips m (1994) A comparison of RAPD
and isozyme analyses for determining the genetic relation­
ships among Avena sterilis L. accessions. Theoretical and Applied
Gtnetics, 87, 68496.
Hong Y, Hipkins VD, Strauss SH (1993) Chloroplast DNA diversi­
ty among trees, populations and species in the California
Cl064!d-Cone Pines (Pinus radiata, Pinus muricata and Pinus
attenuata). Gtnetics, 135, 1187-1196.
Karl SA, Avise JC (1992) Balancing selection at allozyme loci in
oysters: implications from nuclear RFLPs. Science, 256,
100-102.
Lewis PO (1994) GeneStat-PC 3.3. Department of Statistics, north
Carolina State University, Raleigh, NC.
LiP, Adams WT (1989) Range-wide patterns of allozyme variation
in Douglas-fir (Pseudotsuga menziesii). Canadian Journal of Forest
Restarch, 19, 149--161.
Lorenz M, Weihe A, BOmer T (1994) DNA fragments of organellar
origin in random amplified polymorphic DNA (RAPD) pat­
terns of sugar beet (13m vulgaris L.). Theoretical and Applied
Genetics, 88,715-779.
Lynch M, Milligan BG (1994) Analysis of population genetic struc­
ture with RAPD markers. Moleculllr Ecology, 3, 91-99.
Marshall KA. Neale DB (1992) The inheritance of mitochondrial
DNA in Douglas-fir (Pseudotsuga mmziesi1). Canadian Journal of
Forest Resttlrch, 22, 73-75.
Moran GF, Adams WT (1989) Microgeographical patterns of
allozyme differentiation in Douglas-fir from Southwest
Oregon. Forest Science, 35 (1), 3-15.
Moreau F, Kleinschmit J, Kremer A (1994) Molecular differentia­
tion between Q. petraea and Q. robur assessed by random
amplified DNA fragments. Forest Gtnetics, 1 (1), 51-M.
Mosseler A. Egger KN, Hughes GA (1992) Low levels of genetic
diversity in red pine confirmed by random amplified poly­
morphic DNA markers. Canadian Journal of For~t Restarch, 22,
1332-1337.
Neale DB, Wheeler NC, Allard RW (1986) Paternal inheritance of
chloroplast DNA in Douglas-fir. Canadian Journal of Forest
~' 16, 1152-1154.
Nee! JV (1984) A revised estimate of the amount of genetic varia­
tion in human proteins: implications for the distribution of
DNA polymorphisms. American journal of Human Genetics, 36,
1135-1148.
Nei M (1973) Analysis of gene diversity in subdivided popula­
tions. Proceedings of the National Academy of Science USA, 70,
3321-3323.
Nei M (1977) F-statistics and analysis of gene diversity in subdi­
vided populations. Annals of Human Genetics, 41, 225-233.
Nei M (1978) Estimation of average heterozygosity and genetic
distance from a small number of individuals. Genetics, 89,
583-590.
Newbury HJ, Ford-Lloyd BV (1993) The use of RAPD for assess­
ing variation in plants. Plant Growth and Regulation, 12, 43-51.
Peakall R. Smouse PE, Huff DR (1995) Evolutionary implications
of allozyme and RAPD variation in diploid populations of
dioecious buffalograss Buchloe dactyloides. Molecular Ecology, 4,
135-147.
Puterka GJ, Black IV WC, Steiner WM, Burton RL (1993) Genetic
variation and phylogenetic relationships among worldwide
collections of the Russian wheat aphid, Diuraphis noxia
(Mordvilko), inferred from allozyme and RAPD-PCR markers.
Heredity, 70, 604-618.
Rehfeldt GE (1977) Growth and cold hardiness of intervarietal
hybrids of Douglas-fir. Theoretical and Applied Genetics, 50,
3-15.
von Rudloff E (1972) Chemosystematic studies in the genus
Pseudotsuga. I. Leaf oil analysis of the coastal and Rocky
Mountain varieties of the Douglas fir. Canadian Journal of
Botany, 50, 1025-1040.
Schuster W, Brennicke A (1994) The plant mitochondrial genome:
physical structure, information content, RNA editing.. and
gene migration to the nucleus. Annual Review of Plant
Physiology and Plant Molecular Biology, 45, 61-78.
Sederoff RR (1987) Molecular mechanisms of mitochondrial
genome evolution in higher plants. American Naturalist, 130,
530-545.
Sorensen FC (1979) Provenance variation in Pseudotsuga menziesii
seedlings from the var. menziesii-var. glauca transition zone in
Oregon. Silvae Gtnetica, 28 (2-3), 96-103.
Stephan W (1989) Tandem-repetitive noncoding DNA: forms and
forces. Molecular Biology and Evolution, 6 (2), 198-212.
Strauss SH. Hong Y-P, Hipkins VD (1993) High levels of popula­
tion differentiation for mitochondrial DNA haplotypes in
Pinus radiata, muricata, and attenuata. Theoretical and Applied
Gtnetics, 86, 605-611.
Thormann CE, Osborn TC (1992) Use of RAPD and RFLP markers
for germplasm evaluation. In: Proceedings of the Symposium:
Applications of RAPD Technology to Plant Breeding, pp. 9-11.
Joint Plant Breeding Symposia Series, Minneapolis. MN.
Wagner DB, Fumier GR Saghai-Madroof MA, Williams SM,
Cancik BP, Allard RW (1987) Chloroplast DNA polymor­
phisms in lodgepole and jack pines and their hybrids.
Proceedings of the National Academy of Sciences of the USA, 84,
2097-2100.
Waugh R. Powell W (1992) Using RAPD markers for crop
improvement. Tmuis in Biotechnology, 10, 186-191.
WrightS (1965) The interpretation of population structure by F­
statistics with special regard to systems of mating. Evolution,
19, 395-420.
Zavarin E, Snajberk K (1973) Geographic variability of monoter­
penes from cortex of Pseudotsuga menziesii. Pure and Applied
Chemistry, 34, 411-434.