Tuning - BLI-Research-in-Synthetic-Biology-and
advertisement

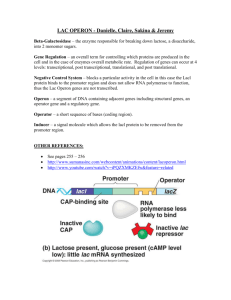
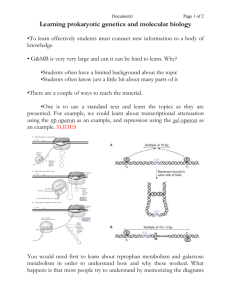
Another engineering principle: Characterization. A stupid engineering joke: • A physicist, a mathematician and an engineer were each asked to establish the volume of a red rubber ball. • The physicist immersed the ball in a beaker full of water and measured the volume of the displaced fluid. The mathematician measured the diameter and calculated a triple integral. The engineer looked it up in his Red Rubber Ball Volume Table. • Basically, engineers want to know the characteristics of their parts and devices. • They list these characteristics in tables, books and files. • Want a beam that can hold 1000 lbs? Look it up. But first • More on bacterial transcription and promoters and such Genome Network Project, Nature Genetics, 2009 Transcriptional Control DNA RNA Environmental change Turn gene(s) on/off protein Proteins to deal with new environment Very important to: 1. express genes when needed 2. repress genes when not needed 3. Conserve energy resources; avoid expressing unnecessary/detrimental genes RNA Structures Vary RNA more like proteins than DNA: structured domains connected by more flexible domains, leading to different functions e.g. ribozymes – catalytic RNA Initiation RNA polymerase α α β β’σ Transcription factors Promoter DNA RNAP binding sites Operator – repressor binding Other TF binding sites Start site of txn is +1 Initiation RNA polymerase 4 core subunits Sigma factor (σ)– determines promoter specificity Core + σ = holoenzyme Binds promoter sequence Catalyzes “open complex” and transcription of DNA to RNA RNAP binds specific promoter sequences Sigma factors recognize consensus -10 and -35 sequences RNA polymerase promoters TTGACA TATAAT Deviation from consensus -10 , -35 sequence leads to weaker gene expression Bacterial sigma factors Sigma factors are “transcription factors” Different sigma factors bind RNAP and recognize specific -10 ,-35 sequences Helps melt DNA to expose transcriptional start site Most bacteria have major and alternate sigma factors Promote broad changes in gene expression E. coli 7 sigma factors B. subtilis 18 sigma factors Generally, bacteria that live in more varied environments have more sigma factors Sigma factors Sigma subunit Type of gene controlled s70 RpoD Growth/housekeeping s54 RpoN N2; stress response ~15 sS RpoS Stationary phase, virulence ~100 sS RpoH Heat shock ~40 sF RpoF Flagella-chemotaxis ~40 Extreme ?heat shock, unfolded proteins ~5 Ferric citrate transport ~5 s32 RpoE FecI # of genes controlled ~1000 E. coli can choose between 7 sigma factors and about 350 transcription factors to fine tune its transcriptional output An Rev Micro Vol. 57: 441-466 T. M. Gruber Lac operon control • Repressor binding prevents RNAP binding promoter • An activating transcription factor found to be required for full lac operon expression: CAP (or Crp) Cofactor binding alters conformation Crp binds cAMP, induces allosteric glucose changes glucose cAMP Crp cAMP Crp lac operon no mRNA mRNA Cooperative binding of Crp and RNAP Binds more stably than either protein alone Interaction of CAP-cAMP, RNA Pol and DNA of lac control region lac operon – activator and repressor CAP = catabolite activator protein CRP = cAMP receptor protein Cis-acting sequence is activator (or CAP) binding site. cAMP signals low glucose activator binding-site lac operon off low lac operon very weakly on lac operon fully induced The ara Operon •another example of operon that has both positive and negative regulation •araB, A, and D encode the 3 arabinose metabolizing enzymes •araC encodes the control protein AraC which is both a positive regulator (in the presence of arabinose) and a negative regulator (in the absence of arabinose). •cAMP-CAP complex also acts as a positive regulator Organization of the ara operon Control of the ara Operon I - Negative araPBAD •When arabinose is absent, the AraC protein acts as a negative regulator. •AraC acts as a dimer, and causes the DNA to loop. Looping brings the I1 and O2 sites in proximity to one another. •One AraC monomer binds to I1 and a second monomer binds to O2. •Binding of AraC prevents RNA Pol from binding to the PBAD promoter Control of the ara Operon II - Positive araPBAD •When arabinose is present, it binds to AraC and changes AraC conformation •An arabinose-AraC dimer complex binds preferentially to I1 and I2, and NOT to O2 which causes ‘opening’ of the loop. This allows RNA Pol to bind to PBAD. •If glucose levels are low, cAMP-CAP complex binds to Pc. •Active transcription occurs. Control can also happen at the Ribosome binding site What about the terminator? • Termination sequence has 2 features: Series of U residues GC-rich self-complimenting region • GC-rich sequences bind forming stem-loop • Stem-loop causes RNAP to pause • U residues unstable, permit release of RNA chain One type of characterization is Tuning • Some promoters bind RNAPs better so they are stronger • Some RBSs make mRNA that bind better to the ribosome so they are stronger • And some are weaker… Tuning • By mixing and matching promoters and RBS parts we can have genetic devices that work at various levels • • • • Weak Promoter + weak RBS = weak device Strong Promoter + strong RBS = strong device Weak Promoter + strong RBS = Medium Promoter + medium RBS = • The synthetic biologists got together and decided on a reference promoter against which others would be measured. • Much like the standard meter. Why would synthetic biologists want to be able to tune a system/device?