ele12423-sup-0002-Suppmat
advertisement

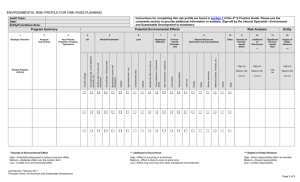
Supporting Information Appendix SA. Details on trees including fossil and extant species and supertree analyses. Input trees for Supertree analyses were based on 30 morphological phylogenetic studies that examined fossil and extant taxa. These are: Taverne (1997), Schultze and Cumbaa (2001), Friedman and Blom (2006), Gardiner et al. (2005), Long et al. (2008), Swartz (2009), Choo (2011), Xu and Gao (2011), Xu and Wu (2012) (early-branching actinopterygian and neopterygian lineages); Grande and Hilton (2006) and Hilton and Forey (2009) (Chondrostei); Grande and Bemis (1998) and Forey and Patterson (2006) (Amiiformes); Arratia (1999) and Arratia (2008) (early teleosts); Taverne (1998), Li and Wilson (1999), Hilton (2003), Zhang (2006), Wilson and Murray (2008) (Osteoglossomorpha); Forey (2004) (clupeomorphs; based on previous studies cited therein); Poyato-Ariza et al. (2010) and Davis et al. (2013) (Gonorynchiformes); (Borden et al. 2013) (paracanthopterygians); Tyler and Santini (2005) (Zeiformes); Otero (2004) (Latinae); Friedman (2008) (Pleuronectiformes); Tyler et al. (1989) (Luvaridae); Tyler and Bannikov (1997) (Siganidae); Santini and Tyler (2003) (Tetraodontiformes). A total of 240 fossil taxa were placed on the molecular phylogeny using a backbone supertree approach; i.e., incongruent clades of extant taxa in the input trees were constrained to conform to the molecular topology. To the best of our knowledge there are no algorithms formally implementing backbone supertrees. This issue was circumvented by up-weighting (“pseudoreplicating”) the molecular topology in the input trees. In many cases, outgroup taxa in the input morphological trees were either added or removed to avoid spurious placements. In some cases, non-disputed monophyletic taxa were not resolved as such in preliminary supertree 1 analyses due to non-overlapping sampling across datasets. Their monophyly was thus enforced on the basis of previous studies. For merging purposes, extant taxa examined by the morphological studies that belong in the same genera as those in the molecular tree were treated as the same taxonomic unit. Other extant taxa present in the morphological trees but not in the molecular phylogeny were removed. Fossil and extant osteoglossomorph relationships hypothesized by several previous studies (i.e., Taverne 1998; Li & Wilson 1999; Hilton 2003; Zhang 2006; Wilson & Murray 2008) are highly incongruent and preliminary supertree analyzes yielded spurious results (i.e., resulted in novel clades not found in any of the input trees). All previous hypotheses were thus reconciled into a single osteoglossomorph supertree prior to the final supertree analysis. Several adjustments were made to both the individual input trees and the osteoglossomorph supertree. (1) Many unstable taxa were excluded from the corresponding input trees, including Jinanichthys, Singida, Kuntulunia, Niierkunia, Huashia, and Joffrichthys. (2) The hypotheses by Li and Wilson (1999) and Wilson and Murray (2008) were combined into a single input tree using the latter topology as a backbone. (3) The relationships of arapaimids, osteoglossids, and pantondontids in the molecular tree were incongruent with most morphological trees. The molecular phylogeny as well as Wilson and Murray’s (2008) tree place arapaimids sister to osteoglossids, whereas the remaining hypotheses indicate as sister-group relationship between pantodontids and osteoglossids. Thus, the three lineages (including stem fossil taxa) were collapsed into a polytomy in all trees that included a Pantodontidae + Osteoglossidae clade. (4) Many marine osteoglossomorphs (i.e., Furichthys fieldsoei, Brychaetoides greenwoodi, Xosteoglossid rebeccae, osteoglossiform indet., Brychaetus sp., and Heterosteoglossum foreyi) were added onto Taverne’s (1998) tree, following the assessment presented by Bonde (2008). 2 While Bonde chose not to conduct a formal phylogenetic analysis, placement of these taxa in clades is supported by synapomorphies. (5) The position of Kipalaichthys, Chanopsis, Laeliichthys, and Paradercites in Taverne’s (1998) cladogram was questioned by Bonde (2008) and thus they were excluded from the tree. (6) Foreyichthys was moved from the stem Osteoglossidae/Arapaimidae + Pantodontidae clade in Taverne’s tree to the crown group, based on the Bonde’s reassessment of the anatomical evidence. (7) The osteoglossomorph supertree estimated after these modifications also resulted in several novel clades that were collapsed into polytomies. The dataset of amiid taxa compiled by Grande and Bemis (1998) and modified Forey and Patterson (2006) (available from http://paleodb.org/bridge.pl?a=getNexusFile&nexusfile_no=159) was reanalyzed in TNT. The strict consensus of 8282 trees with 174 steps is reported. Finally, in order to obtain an adequate placement of flatfish fossils following Friedman (2008), the monophyly of extant Pleuronectiformes was enforced (see also Betancur-R. et al. 2013b; Betancur-R. & Orti 2014). Details on time-calibration of the supertree cladogram are given in the main text. The complete time tree with fossil and extant species is shown in Fig. S1. Appendix SB. Fossils and branch-length scaling of fossil-based trees. Table SB1. List of 240 fossils placed onto the global phylogeny using a supertree approach. Individual trees were obtained from the aforementioned studies (Appendix SA). Habitat and minimum age information was obtained from the corresponding references, the Paleobiology 3 Database (www.paleodb.org), and other sources. In many cases, the absolute minimum age is based on the corresponding stratigraphic horizon (http://www.geosociety.org/science/timescale/). Species with unknown habitat information are coded as either marine or freshwater to assess sensitivity of analyses to coding ambiguity (see main text). # 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 Fossil Dialipina Cheirolepis trailli Cheirolepis schultzei Cheirolepis canadensis Osorioichthys marginis Tegeolepis clarki Howqualepis rostridens Donnrosenia schaefferi Gogosardina coatesi Mimipiscis bartrami Mimipiscis toombsi Moythomasia n. sp. Moythomasia nitida Moythomasia durgaringa Stegotrachelus finlayi Krasnoyarichthys jesseni Novagonatodus kasantsevae Limnomis delaneyi Cuneognathus gardineri Mansfieldiscus sweeti Melanecta anneae Woodicthys bearsdeni Wendyichthys dicksoni Kentuckia hlavini Mesopoma Pteronisculus Birgeria Saurichthys Chondrosteus acipenseroides Peipiaosteus Protopsephurus liui Psammorhynchus longipinnis Boreosomus Min. Age (MA) 398 390 372 372 368 361 383 383 372 372 372 372 372 372 383 359 347 359 359 347 318 318 318 359 338 247 251 241 191 125 125 71 251 4 Habitat F F M FM M M F F M M M M M M F M F F F F M M M M M M M M M F F F M 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 Australosomus Beishanichthys Fukangichthys Scanilepis Evenkia Perleidus Felberia Luganoia lepidosteoides Macrosemius Kyphosichthys Semionotus Watsonulus eugnathoides Ionoscopus cyprinoides Oshunia brevis Ophiopsis procera Macrepistius arenatus Liodesmus gracilis Liodesmus sprattiformis Caturus furcatus Amblysemius pachyurus Sinamia zdanskyi Ikechaoamia orientalis Ikechaoamia meridionalis Amiopsis woodwardi Amiopsis dolloi Amiopsis prisca Amiopsis lepidota Amiopsis damoni Solnhofenamia elongata Nipponamia satoi Calamopleurus cylindricus Calamopleurus mawsoni Melvius thomasi Melvius chauliodous Vidalamia catalunica Pachyamia latimaxillaris Pachyamia mexicana Maliamia gigas Tomognathus mordax Amia hesperia Amia pattersoni Amia scutata Cyclurus ignotus 251 247 202 196 251 239 237 235 151 242 199 246 146 109 146 100 146 146 146 146 112 134 134 137 124 92 146 137 146 126 110 128 66 74 136 98 100 48 90 48 50 36 37 5 M F F FM F M M M M M FM M M M M FM M M M M F F F FM F FM M FM M F FM F FM FM FM M M FM M F F F FM 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 Cyclurus fragosus Cyclurus oligocenicus Cyclurus gurleyi Cyclurus kehreri Cyclurus macrocephalus Cyclurus valenciennesi Cyclurus efremovi Pseudamiatus heintzi Dapedium Hypsocormus Pachycormus Mesturus Vinctifer Aspidorhynchus Belonostomus Pholidophorus macrocephalus Pholidophorus bechei Leptolepis coryphaenoides Tharsis Domeykos Varasichthys Luisichthys Protoclupea Chongichthys Notelops Apsopelix Crossognathus Goulmimichthys Rhacolepis Ascalabos Pachythrissops Allothrissops Thrissops Anaethalion zapporum Anaethalion angustus Anaethalion knorri Tongxinichthys microdus Jiuquanichthys liui Kuyangichthys microdus Lycoptera davidi Tanolepis ningjiagouensis Paralycoptera wui Xixiaichthys 66 33 50 48 38 66 59 62 191 146 176 161 100 140 65.5 146 191 176 146 157 157 157 157 156 100 80 106 89 113 146 146 151 146 151 151 146 126 100 100 130 113 110 126 6 F F F F F F F FM M M M M FM M M M M M M M M M M M FM M M M FM M M M M M M M M F F F F F F 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 Plesiolycoptera daqingensis Yanbiania wangqingica Eohiodon woodruffi Jiaohichthys Eohiodon rosei Hiodon consteniorum Thaumaturus Ostariostoma Furichthys fieldsoei Palaeonotopterus Brychaetoides greenwoodi Musperia radiata Phaerodusichthys tavernei Monopterus gigas Heterosteoglossum foreyi Chauliopareion Xosteoglossid rebeccae Opsithrissops osseus osteoglossiform indet Foreyichthys bolcensis Ridewoodichthys caheni Thrissopterus catullii Sinoglossus lushanensis Cretophareodus Brychaetus sp Brychaetus muelleri Phareodus testis Phareodus queenslandicus Phareodus encaustus Paraclupea Ellimmichthys Armigatus Diplomystus Triplomystus Sorbinichthys Spratticeps Santanaclupea Tischlingerichthys Notogoneus Charitopsis Charitomosus Judeichthys Hakeliosomus 66 130 49 130 49 34 34 66 55 94 55 34 62 49 55 40 55 55 55 49 56 49 40 71 55 49 50 35 50 145 145 94 50 94 94 100 113 146 83 100 83 100 100 7 F F F F F F F F M F M F F M M F M M M M M M F F M M F F F F FM M F M M M FM M F M M M M 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 Ramallichthys Mahengichthys Gordichthys Rubiesichthys Aethalionopsis Parachanos Dastilbe Tharrias Humbertia Erichalcis Leptolepides haerteisi Leptolepides sprattiformis Orthogonikleithrus hoelli Orthogonikleithrus leichi Sphenocephalus Mcconichthys Trichophanes Libotonius Amphiplaga Erismatopterus Lateopisciculus Massamoricthys Archaeozeus skamolensis Protozeus kuehnei Cretazeus rinaldii Palaeogadus Eolates gracilis Eolates aquensis Heteronectes Amphistium Joleaudichthys Numidopleura Eobothus Kushlukia permira Kushlukia sp. Avitoluvarus dianae Avitoluvarus mariannae Luvarus necopinatus Ruffoichthys Eosiganus Siganopygaeus Protosiganus Cretatriacanthus guidottii 100 45 130 139 113 83 112 112 100 105 146 146 146 146 72 56 34 34 50 50 56 56 58 58 72 28 49 28 49 49 41 41 49 56 48 56 56 56 49 38 56 28 70 8 M F F F F FM FM FM M M M M M M M F F F F F F F M M M M M F M M M M M M M M M M M M M M M 206 Plectocretacicus clarae 95 M 207 Protriacanthus gortanii 90 M 208 Prohollardia avita 23 M 209 Carpathospinosus propheticus 23 M 210 Bolcabalistes varii 49 M 211 Eospinus daniltshenkoi 49 M 212 Spinacanthus cuneiformis 49 M 213 Protobalistum imperialis 49 M 214 Proaracana dubia 49 M 215 Eolactoria sorbinii 49 M 216 Oligolactoria bubiki 28 M 217 Prodiodon tenuispinus 49 M 218 Prodiodon erinaceus 49 M 219 Zignodon fornasieroae 49 M 220 Pshekhadiodon parini 41 M 221 Heptadiodon echinus 49 M 222 Archaeotetraodon jamestyleri 5 M 223 Sphoeroides hyperostosus 4 M 224 Archaeotetraodon winterbottomi 23 M 225 Protacanthodes ombonii 49 M 226 Protacanthodes nimesensis 49 M 227 Acanthopleurus trispinosus 23 M 228 Cryptobalistes brevis 28 M 229 Acanthopleurus collettei 28 M 230 Acanthopleurus serratus 28 M 231 Eoplectus bloti 49 M 232 Eotetraodon pygmaeus 49 M 233 Zignoichthys oblongus 49 M 234 Triodon antiquus 34 M 235 Eomola bimaxillaria 41 M 236 Moclaybalistes danekrus 59 M 237 Oligobalistes robustus 28 M 238 Balistomorphus ovalis 28 M 239 Balistomorphus orbiculatus 28 M 240 Balistomorphus spinosus 28 M M: Marine. F: Freshwater FM: Unknown habitat or found in both marine and freshwater habitats. Table SB2. Comparisons of the root age for extant ray-finned fishes (MRCA of Polypterus and Danio) obtained with the fossil-based tree under different values of the “vartime” variable in 9 Paleotree (timePaleoPhy routine). Compare these ages with the divergence time estimated using the extant only molecular tree, based and node-age calibrations (383 Ma). Values of vartime above 1.0 impact the age of the root and were not included in downstream analyses. Vartime 0.1 0.5 1.0 2.0 5.0 10.0 Root age (Ma) 395 395 395 403 438 740 Appendix SC. Ancestral state reconstructions. Assessment of sensitivity of ancestral state (habitat) reconstructions: (i) default analyses (BiSSE using trees with extant taxa; stochastic mapping [SIMMAP-MK2] using trees with fossil and extant taxa) reported in the main text but depicting complete trees (Fig. SC1); (ii) alternative analyses using ML-MK2 reconstructions (Fig. SC2); (iii) linear plots and regressions comparing ML-MK1 vs. ML-MK2 reconstructions (Fig. SC3); (iv) linear plots comparing SIMMAP-MK2 reconstructions for alternative trees with fossil and extant taxa, scaled under different values of the Vartime parameter (0.1, 0.5, 1.0) in Paleotree (Fig. SC4); and (v) SIMMAP-MK2 analyses using alternative tree topologies (six additional trees obtained with RAxML search replicates; Fig. SC5). The rationale for choosing MK2 (instead of the simpler MK1) as the default model is explained with detail in Appendix SG. 10 Figure SC1. Ancestral state (habitat) reconstructions (ASR) under different models (BiSSE and SIMMAP), coding scenarios (BEAM - brackish and euryhaline as marine; BEAF - brackish and euryhaline as freshwater), and using different trees (including or excluding fossils). Ovals highlight major differences in ancestral habitat reconstructions among early-branching linages. 11 12 Figure SC2. Ancestral state (habitat) reconstructions (ASR) under ML (Ape’s function Ace) under different models (MK1 and MK2) and coding scenarios (BEAM - brackish and euryhaline as marine; BEAF - brackish and euryhaline as freshwater). Ovals highlight major differences in ancestral habitat reconstructions among early-branching linages. Figure SC3. Linear plots of state probability (freshwater probability) at each node for ML-MK1 vs. ML-MK2 using Ape’s function Ace (see also Fig. SC2). BEAM: brackish and euryhaline taxa coded as marine; BEAF: brackish and euryhaline taxa coded as freshwater. 13 Figure SC4. Linear plots of state probability (freshwater probability) at each node obtained with SIMMAP for alternative trees with fossil and extant taxa. The alternative trees used different values of the Vartime parameter for scaling of fossil branches in Paleotree. State probabilities among alternative reconstructions are highly similar. BEAM: brackish and euryhaline taxa coded as marine; BEAF: brackish and euryhaline taxa coded as freshwater. 14 Figure SC5. Ancestral state (habitat) reconstructions (ASR) under SIMMAP using six alternative topologies obtained with the RAxML tree searches and under alternative coding scenarios (BEAM - brackish and euryhaline as marine; BEAF - brackish and euryhaline as freshwater). In all cases, early branching lineages are reconstructed with a high probability of freshwater occupancy, indicating that the ASRs are robust to topological variations. 15 Appendix SD. State-dependent diversification (SSE). Table SD1. Sampling fractions defined across 50 (mostly nested) clades to account for sampling biases in the BiSSE analyses (uneven sampling). We also attempted to define sampling fractions across 57 non-nested clades, but there were two situations that violated the method’s assumptions: (1) one of the sates is absent in some clades, which results in an undefined sampling fraction (a/0); and (2) the two states exist in the clade but all the sampled taxa belong to one state only, which results in sampling fractions of 0 that cannot be handled by the method. As per recommendation of R. Fitzjohn (pers. comm.), we thus defined sampling fractions across 50 clades that included all habitat states (both sampled and unsampled species). Clades are shown in reverse phylogenetic order; clade names follow our current classification scheme for bony fishes (Betancur-R. et al. 2013a; Betancur-R. et al. 2014). Clade Cottioidei Notothenioidei Perciformes Centrarchiformes s.l. Terapontiformes Terapontiformes + Centrarchiformes s.l. Tetraodontiformes Tetraodontiformes + Lophiiformes Sciaenidae Eupercaria Beloniformes Cyprinodontiformes Atheriniformes Atherinomorphae Cichlomorphae Ovalentaria BEAM Freshwater 0.0294 1.0000 0.0698 0.2763 0.0455 0.1917 0.0769 0.0769 0.0385 0.1005 0.0404 0.0067 0.0551 0.0163 0.0110 0.0148 16 BEAM Marine 0.0670 0.1032 0.0660 0.1746 0.1235 0.1458 0.1301 0.1043 0.0755 0.0884 0.1195 0.2593 0.1101 0.1288 0.1429 0.0809 BEAF Freshwater 0.0442 1.0000 0.0748 0.2763 0.0877 0.1955 0.0625 0.0625 0.0588 0.1162 0.0696 0.0115 0.0687 0.0251 0.0110 0.0220 BEAF Marine 0.0659 0.0974 0.0653 0.1746 0.1029 0.1374 0.1332 0.1055 0.0750 0.0863 0.1049 0.1111 0.0843 0.0979 0.2500 0.0731 Pleuronectiformes Carangiaria Carangiaria + Anabantaria Carangiaria + Anabantaria + Ovalentaria Syngnathiformes + Scombriformes Gobiiformes Kurtiformes Gobiaria Percomorpha Polymixiiformes + Euacanthomorphacea Zeiogadaria Paracanthomorphacea Acanthomorphata Neoteleostei Osmeriformes Stomiatii Protacanthopterygii Euteleostei Diplomystoidei + Siluroidei Siluriformes Siluriformes + Characiformes Characiphysae Otophysi Gonorynchiformes Ostariophysi Alepocephaliformes + Ostariophysi Clupeiformes Otomorpha Clupeocephala Osteoglossocephalai Elopomorpha Teleostei Neopterygii Actinopteri ROOT BEAM: brackish and euryhaline as marine BEAF: brackish and euryhaline as freshwater 0.0238 0.0606 0.0640 0.0205 0.0286 0.0058 0.1333 0.0094 0.0284 0.0284 0.2500 0.7000 0.0298 0.0298 0.2857 0.2857 0.0365 0.0309 0.0140 0.0104 0.0115 0.0124 0.0099 0.0938 0.0102 0.0102 0.0200 0.0103 0.0172 0.0177 0.1250 0.0177 0.0181 0.0183 1.00000 0.1027 0.1357 0.1357 0.0983 0.0954 0.0386 0.0569 0.0419 0.0833 0.0851 0.0718 0.0714 0.0849 0.0865 0.2647 0.0846 0.0805 0.0864 0.0187 0.0187 0.0185 0.0185 0.0175 0.6667 0.0500 0.0615 0.0235 0.0412 0.0847 0.0847 0.0164 0.0806 0.0806 0.0806 1.00000 0.0533 0.1290 0.0813 0.0294 0.0357 0.0120 0.1579 0.0156 0.0387 0.0387 1.0000 0.5385 0.0399 0.0399 0.2571 0.2500 0.0538 0.0422 0.0143 0.0106 0.0117 0.0125 0.0099 0.1471 0.0104 0.0104 0.0292 0.0107 0.0222 0.0226 0.0652 0.0227 0.0230 0.0233 1.00000 0.1032 0.1314 0.1314 0.0915 0.0966 0.0409 0.0545 0.0438 0.0813 0.0832 0.0714 0.0718 0.0831 0.0849 0.3333 0.0741 0.0562 0.0843 0.0125 0.0125 0.0125 0.0125 0.0123 0.5000 0.0353 0.0578 0.0176 0.0376 0.0829 0.0829 0.0149 0.0787 0.0787 0.0787 1.00000 Table SD2. Assessments of best-fit models and preliminary rates obtained using a maximum likelihood framework. The ten GeoSSE models tested were the full (λM, λF, 17 λMF, μM, μF, qMF, and qFM), no intermediate speciation (no λMF), equal speciation (λM~λF), equal extinction (μM~μF), equal speciation and equal extinction (λM~λF and μM~μF), equal speciation and dispersal (λM~λF and qMF~qFM), equal extinction and dispersal (μM~μF and qMF~qFM), and all-equal rates (no sMF, λM~λF, μM~μF, and qMF~qFM). The nine BiSSE models tested were similar except that the full model has only six parameters (λM, λF, μM, μF, qMF, and qFM) and thus does not account for intermediate speciation (λMF). Method BiSSE BiSSE GeoSSE GeoSSE BiSSE BiSSE Sampling Even Even Even Even Uneven (50 clades) Uneven (50 clades) Coding scenarios BEAF BEAM BAF BAM BEAF BEAM Best-fit model Full Full Full No λMF λM~λF Full Net. Div. ratio 2.4 2.8 2.6 2.7 1.6 1.8 Turnover ratio 0.91 0.89 0.89 0.88 0.96 0.93 Net diversification rates ratio: net diversification rates marine/net diversification rates freshwater Turnover rates ratio: turnover rates marine/turnover rates freshwater BAM: brackish as marine (GeoSSE) BAF: brackish as freshwater (GeoSSE) BEAM: brackish and euryhaline as marine (BiSSE) BEAF: brackish and euryhaline as freshwater (BiSSE) 18 Figure SD1. MCMC plots of net diversification (speciation minus extinction) and turnover (extinction/speciation) rates across habitats based on the Clupeocepahala subtree. BAM: brackish as marine (GeoSSE); BAF: brackish as freshwater (GeoSSE); BEAM: brackish and euryhaline as marine (BiSSE); BEAF: brackish and euryhaline as freshwater (BiSSE). 19 Appendix SE. Transition rates. Table SE1. Estimates of transition rates obtained with alternative methods (SIMMAP, SSE) coding scenarios (BEAM, BEAF, BAM, BAF), sampling fraction schemes (even, uneven), and trees (extant or extant + fossils). Method Sampling fractions Tree/coding BiSSE Even Extant/BEAF BiSSE Even Extant/BEAM GeoSSE Even Extant/BAF GeoSSE Even Extant/BAM BiSSE Uneven (50 clades) Extant/BEAF BiSSE Uneven (50 clades) Extant/BEAM SIMMAP-MK2 _ Extant/BEAM SIMMAP-MK2 _ Extant/BEAF SIMMAP-MK2 _ Extant + fossils/BEAM SIMMAP-MK2 _ Extant + fossils/BEAF qFM: freshwater-to-marine transition rate qMF: marine-to-freshwater transition rate BEAM: brackish and euryhaline as marine (BiSSE, SIMMAP) BEAF: brackish and euryhaline as freshwater (BiSSE, SIMMAP) BAM: brackish as marine (GeoSSE) BAF: brackish as freshwater (GeoSSE) Transition rate ratio (qMF/qFM) 11.6 21.5 31.6 36.2 2.0 2.1 4.9 1.6 1.4 1.2 Appendix SF. Clade-based analyses. Table SF1. List of 67 non-nested target clades and their estimates of stem and crown ages, species richness, mean habitat state (freshwater) probability, and discretized habitat state. Clade state probabilities were estimated by averaging the results obtained with different models (BiSSE, SIMMAP, ML) and coding strategies (BEAM and BEAF). 20 Habitat discretization resulted in 20 freshwater (freshwater probability >0.9) and 35 marine clades (freshwater probability < 0.1); 12 remaining clades had ambiguous state probabilities (>0.1 and <0.9) and were thus excluded from consideration (listed as “?”). Taxa denoted with asterisk have a crown age of 80–50 Ma (9 freshwater, 19 marine clades, and 5 ambiguous that were excluded), which were selected for reduced cladebased comparisons (see main text). Clade Stem Age (Ma) Crown Age (Ma) Polypteriformes Acipenseriformes Holostei Elopiformes Albuliformes Notacanthiformes* Anguilliformes* Hiodontiformes Osteoglossiformes Clupeiformes Alepocephaliformes* Gonorynchoidei Chanoidei + Knerioidei Cobitoidea* Cyprinoidea* Gymnotiformes* Characiformes Loricaroidei Diplomystoidei + Siluroidei Lepidogalaxiiformes Argentiniformes* Galaxiiformes* Esociformes* Salmoniformes Retropinnidae Osmeriformes (in part) Stomiatiformes Ateleopodiformes Aulopiformes Neoscopelidae Myctophidae* Lampridiformes 382.6 350.1 322.5 196.2 150.8 101.0 101.0 227.1 227.1 230.2 219.7 175.9 175.9 99.3 99.3 147.8 137.0 115.8 115.8 231.8 159.6 145.6 104.5 55.8 73.7 73.7 129.4 192.3 182.8 73.6 73.6 150.3 29.2 138.9 267.9 133.6 40.5 50.7 79.4 9.5 163.1 188.9 53.3 18.2 147.1 78.8 63.3 69.0 114.8 110.6 106.2 3.5 70.5 50.0 79.4 35.3 22.5 34.3 83.8 8.1 113.9 42.4 51.7 81.7 21 Mean state Clade (freshwater) richness probability 12 0.999 28 0.988 8 0.962 9 0.439 13 0.011 27 0.000 934 0.000 2 1.000 228 0.998 398 0.820 140 0.006 5 0.065 33 0.867 1143 1.000 2916 1.000 204 1.000 2004 1.000 1385 1.000 2183 1.000 2 1.000 89 0.033 50 0.814 13 0.984 216 0.536 6 0.546 35 0.495 427 0.001 13 0.000 262 0.000 7 0.000 252 0.000 24 0.001 Discretized clade habitat Freshwater Freshwater Freshwater ? ? Marine Marine Freshwater Freshwater ? Marine Marine ? Freshwater Freshwater Freshwater Freshwater Freshwater Freshwater Freshwater Marine ? Freshwater ? ? ? Marine Marine Marine Marine Marine Marine Percopsiformes* Zeiformes* Gadariae* Polymixiiformes Beryciformes Holocentriformes* Ophidiiformes* Batrachoidiformes Kurtiformes* Gobiiformes* Syngnathiformes* Scombriformes* Synbranchiformes* Anabantoidei* Channoidei* Carangiaria* Polycentridae Cichllidae* Atheriniformes* Cyprinodontiformes* Beloniformes* Pomacentridae + Mugilidae + Blenniiformes + others Labriformes* Moronidae + Ephippiformes Lobotiformes + Sciaenidae Acanthuriformes + Pomacanthidae + Chaetodontidae + Haemulidae + Lutjanidae + others Spariformes* Lophiiformes + Tetraodontiformes* Uranoscopiformes Pempheriformes Terapontiformes + Centrarchiformes Serranidae* Percidae Notothenioidei* Cottioidei* 135.0 107.1 107.1 154.8 146.5 145.0 132.8 126.8 102.3 102.3 94.6 94.6 80.7 70.7 70.7 96.4 94.2 88.7 77.4 76.4 76.4 63.2 66.5 78.8 13.7 125.3 52.5 75.4 39.8 80.3 72.8 74.3 50.3 73.7 62.8 67.4 69.4 46.1 76.4 70.9 66.9 71.9 9 33 612 10 182 84 531 84 349 1996 662 274 124 69 163 1068 4 1642 345 1227 258 0.980 0.000 0.001 0.000 0.000 0.001 0.000 0.000 0.052 0.202 0.000 0.000 0.999 1.000 1.000 0.000 0.989 0.988 0.500 0.512 0.502 Freshwater Marine Marine Marine Marine Marine Marine Marine Marine ? Marine Marine Freshwater Freshwater Freshwater Marine Freshwater Freshwater ? ? ? 97.2 93.6 93.8 92.2 94.6 76.7 88.6 86.4 1920 625 24 298 0.085 0.000 0.007 0.001 Marine Marine Marine Marine 92.2 85.9 85.7 80.3 691 253 0.000 0.000 Marine Marine 80.8 95.3 93.2 80.4 89.0 89.6 787 165 195 0.000 0.000 0.000 Marine Marine Marine 93.2 80.2 66.0 76.0 73.1 84.7 80.1 40.3 63.0 70.8 282 535 233 156 1266 0.060 0.000 0.994 0.004 0.000 Marine Marine Freshwater Marine Marine Table SF2. Estimates of net diversification rates for the 55 target clades (35 marine and 20 freshwater) using both crown and stem equations implemented in the method-of- 22 moments (Magallon & Sanderson 2001). Rates were calculated based on the mean values of ε 0.30) habitats and confidence limits were assessed all cases, net diversification rates are higher for marine (MA) vs. freshwater (FW) clades, although only two comparisons (crown-based) are significant (*P< 0.05). Extinction (ε) 0.0 FW= 0.53; MA= 0.30 0.9 Mean net div. rates FW - Crown 0.0504 Mean net div. rates MA Crown 0.0705 U test P value 0.043* Mean net div. rates FW - Stem 0.0419 Mean net div. rates MA - Stem 0.0503 U test P value 0.326 0.0468 0.0330 0.0689 0.0433 0.019* 0.082 0.0360 0.0249 0.0470 0.0296 0.110 0.362 Table SF3. Estimates of net diversification rates for a subset of the clades in Table SF1 (9 freshwater and 19 marine clades) whose crown age is 80–50 Ma. Both crown and stem equations implemented in the method-of-moments are compared (Magallon & Sanderson 2001). Rates were calculated based on the mean values of ε (extinction) estimated with fractions. In all cases, net diversification rates are higher for marine (MA) vs. freshwater (FW) clades, although only one comparison (crown-based) is marginally significant (*P = 0.068) presumably due to reduced statistical power. Extinction (ε) Mean net div. rates FW - Crown Mean net div. rates MA Crown 23 U test P value Mean net div. rates FW - Stem Mean net div. rates MA - Stem U test P value 0.0 FW= 0.53; MA= 0.30 0.9 0.0266 0.0436 0.06813 0.0476 0.0522 0.7355 0.0596 0.0426 0.0716 0.0490 0.2051 0.3568 0.0481 0.0330 0.0550 0.0357 0.4982 0.7355 Table SF4. Estimates of net diversification rates for a subset of the clades in Table SF1 (9 freshwater and 19 marine clades) whose crown age is 80–50 Ma. Both crown and stem equations implemented in the method-of-moments are compared (Magallon & Sanderson 2001). Rates were calculated based on the mean values of ε (extinction) estimated with BiSSE/GeoSSE for freshwater (= 0.53) and marine (= 0.30) habitats and confidence limits were assessed under arbitrarily high (= 0.90) and low (= 0.0) extinction fractions. Most rate estimates based on crown equations are significantly higher than those based on stem equations for marine (MA) clades, whereas all crown vs. stem comparisons for freshwater (FW) clades are non-significant. Note that these results may suffer from limited statistical power. Extinction (ε) 0.0 FW= 0.53; MA= 0.30 0.9 Mean net div. rates FW - Crown 0.0266 Mean net div. rates FW - Stem 0.0476 U test P value 0.297 Mean net div. rates MA - Crown 0.0436 Mean net div. rates MA - Stem 0.0522 U test P value 0.284 0.0596 0.0426 0.0481 0.0330 0.340 0.340 0.0716 0.0490 0.0550 0.0357 0.099** 0.008** **P< 0.01 24 Figure SF1. Correlations of (log) species richness against crown and stem ages, for 19 marine and 9 freshwater clades. Note that these results may suffer from limited statistical power. Appendix SG. Comparisons of model fitting using Markov models that assume fixed diversification parameters but allow transition rates to be either symmetric (MK1) or asymmetric (MK2). 25 One of the reviewers of the paper (Graham Slater) commented: “What would be useful, straightforward and clearer for this manuscript would be to first compare the fit of different flavors of MK models to the extant only and extant plus extinct datasets using ML. This could be done, for instance, by fitting equal rates, symmetric (meaning transition rates within, but not among pairs of states, are equal), and all rates different models using fitDiscrete in Geiger.” We followed his suggestion and tested the fit of both datasets (extant only and fossil-based) to the MK1 (equal rates) and MK2 (all-rates-different) models using the fitDiscrete function in Geiger as well as the Ace function in Ape. In all cases the tests failed to reject the null MK1 model (ΔAIC <4; Table SG1), although those using the fossil-based dataset were marginally below significance. Table SG1. Results of model fitting using Markov models that assume fixed diversification parameters but allow transition rates to be symmetric (MK1; qMF~qFM) or asymmetric (MK2; qMF≠qFM). ΔAIC fitDiscrete (Geiger) ML Extant/BEAM 2.00 ML Extant/BEAF 1.50 ML Extant + fossils/BEAM 3.56 ML Extant + fossils/BEAF 3.79 BEAM: brackish and euryhaline as marine BEAF: brackish and euryhaline as freshwater Method Tree/coding ΔAIC Ace (Ape) 0.67 1.54 3.55 3.79 These results were surprising (particularly for the extant-only dataset) given that model fitting using SSE strongly indicated asymmetrical transition rates in most cases 26 (ΔAIC values of up to 74; results of Table 2 summarized on Table SG2 for ease of comparison). Table SG2. Results of model fitting using SSE models (extant-only dataset) where transition rates are fixed as equal (qMF~qFM) while speciation and extinction are free to vary (λM≠λF; μM≠μF). Adapted from Table 2. Method BiSSE BiSSE GeoSSE GeoSSE BiSSE BiSSE Sampling fractions Even Even Even Even Unveven (50 clades) Unveven (50 clades) Tree/coding Extant/BEAF Extant/BEAM Extant/BAF Extant/BAM Extant/BEAF Extant/BEAM qMF~qFM (ΔAIC) 62*** 65*** 74*** 70*** 4* 2 BEAM: brackish and euryhaline as marine (BiSSE, SIMMAP) BEAF: brackish and euryhaline as freshwater (BiSSE, SIMMAP) BAM: brackish as marine (GeoSSE) BAF: brackish as freshwater (GeoSSE) The fitDiscrete and Ace results are not only contrary to those using SSE, but also challenge the biological notion that habitat transitions in fishes are asymmetric (see Introduction). This bizarre result led us to hypothesize that these functions may lack statistical power. To test this idea we first used the tree and habitat data from our previous study with ariid catfishes (Betancur-R et al. 2012) where we show that transition rates are highly asymmetric: there were 12 independent events of freshwater colonization by marine ariid lineages vs. (possibly) a single event of freshwater-to-marine invasion in recent times (Fig. 1). The ΔAIC score from this comparison was, again, not significant (fitDiscrete = 1.91; Ace = 2.72). 27 We then simulated 10 birth-death trees (λ=0.1, μ=0.03) and binary characters for 1500 taxa using the tree.bd and sim.character functions in diversitree. Character states were simulated under MK2 assuming highly asymmetrical rates, with state shifts being more than an order of magnitude higher for one transition type than the other (q01 = 0.0003; q10= 0.006). These transition parameter values were based on our results using BiSSE, BDAM coding, and even sampling – i.e., the BiSSE result with the highest ΔAIC (65) in our model comparisons (see Table SG2). Once again, both fitDiscrete and Ace failed to reject the null hypothesis of equal transition rates in all comparisons, although one ΔAIC score was marginally below 4.0 (Table SG3) Table SG3. Results of MK1 vs. MK2 model fitting using 10 datasets simulated with highly asymmetrical transition rate parameters (q01 = 0.0003; q10= 0.006). Simulation 1 2 3 4 5 6 7 8 9 10 ΔAIC fitDiscrete (Geiger) 0.99 1.98 1.84 1.98 1.98 2.00 1.92 3.65 1.99 1.98 ΔAIC Ace (Ape) 0.99 1.98 1.98 1.98 1.98 1.98 1.98 1.98 1.98 1.98 It is beyond the scope of this study to properly assess the power of fitDiscrete and Ace, as that would require more exhaustive analyses. However, these preliminary results show that these tests may suffer from high rates of type II error. We thus base our 28 conclusions on the asymmetry of habitat transition following the SSE result and conduct all major ancestral state reconstructions using the MK2 model (asymmetric rates) under SIMMAP and ML. It is noteworthy, however, that the ASRs using Ace are robust to model choice (MK1 vs. MK2; Fig. SC2). 29 REFERENCES 1. Arratia, G. (1999). The monophyly of Teleostei and stem-group teleosts. Consensus and disagreements. In: Mesozoic Fishes 2 – Systematics and Fossil Record (eds. Arratia, G & Schultze, HP). Verlag Dr. F. Pfeil München, pp. 265-334. 2. Arratia, G. (2008). The varasichthyid and other crossognathiform fishes, and the Break-up of Pangaea. Geological Society London Special Publications, 295, 7192. 3. Betancur-R, R., Ortí, G., Stein, A.M., Marceniuk, A.P. & Alexander Pyron, R. (2012). Apparent signal of competition limiting diversification after ecological transitions from marine to freshwater habitats. Ecology Letters, 15, 822-830. 4. Betancur-R., R., Broughton, R.E., Wiley, E.O., Carpenter, K., Lopez, J.A., Li, C. et al. (2013a). The tree of life and a new classification of bony fishes. PLoS Currents Tree of Life, 2013 Apr 18. 5. Betancur-R., R., Li, C., Munroe, T.A., Ballesteros, J.A. & Orti, G. (2013b). Addressing gene-tree discordance and non-stationarity to resolve a multi-locus phylogeny of the flatfishes (Teleostei: Pleuronectiformes). Systematic Biology, 62, 763–785. 6. Betancur-R., R. & Orti, G. (2014). Molecular evidence for the monophyly of flatfishes (Carangimorpharia: Pleuronectiformes). Molecular phylogenetics and evolution, 73, 18-22. 7. Betancur-R., R., Wiley, E.O., Miya, M., Lecointre, G. & Orti, G. (2014). Phylogenetic Classification of Bony Fishes. Version 3. Available at: http://www.deepfin.org/Classification_v3.htm Last accessed July 2014. 8. Bonde, N. (2008). Osteoglossomorphs of the marine Lower Eocene of Denmark – with remarks on other Eocene taxa and their importance for palaeobiogeography. In: Fishes and the Break-up of Pangaea (eds. Cavin, L, Longbottom, A & Richter, M). Geological Society, London, Special Publications London, pp. 253–310. 9. Borden, W.C., Grande, T. & Smith, W.L. (2013). Comparative osteology and myology of the caudal fin in the Paracanthopterygii (Teleostei: Acanthomorpha). In: Mesozoic Fishes 5 - Global Diversity and Evolution (eds. Arratia, G & Schultze, H-P). Verlag F. Pfeil Muenchen. 10. Choo, B. (2011). Revision of the actinopterygian genus Mimipiscis (=Mimia) from the Upper Devonian Gogo Formation of Western Australia and the 30 interrelationships of the early Actinopterygii. Earth and Environmental Science Transactions of the Royal Society of Edinburgh, 102, 77-104. 11. Davis, A.M., Arratia, G. & Kaiser, T.M. (2013). The first fossil shellear and its implications for the evolution and divergence of the Kneriidae (Teleostei: Gonorynchiformes). In: Mesozoic Fishes 5 - Global Diversity and Evolution (eds. Arratia, G, Schultze, H-P & Wilson, MVH). Verlag F. Pfeil Muenchen, pp. 325-362. 12. Forey, P. (2004). Basal clupeomorphs and ellimmichthyiform phylogeny. In: Mesozoic Fishes 3 – Systematics, Paleoenvironments and Biodiversity (eds. Arratia, G & Tintori, A). Verlag Dr. F. Pfeil München, pp. 391-404. 13. Forey, P.L. & Patterson, C. (2006). Description and systematic relationships of †Tomognathus, an enigmatic fish from the English Chalk. Journal of Systematic Palaeontology, 4, 157-184. 14. Friedman, M. (2008). The evolutionary origin of flatfish asymmetry. Nature, 454, 209-212. 15. Friedman, M. & Blom, H. (2006). A NEW ACTINOPTERYGIAN FROM THE FAMENNIAN OF EAST GREENLAND AND THE INTERRELATIONSHIPS OF DEVONIAN RAY-FINNED FISHES. Journal of Paleontology, 80, 1186-1204. 16. Gardiner, B.G., Schaeffer, B. & Masserie, J.A. (2005). A review of the lower actinopterygian phylogeny. Zoological Journal of the Linnean Society, 144, 511-525. 17. Grande, L. & Bemis, W.E. (1998). A comprehensive phylogenetic study of amiid fishes (Amiidae) based on comparative skeletal anatomy. An empirical search for interconnected patterns of natural history. Journal of Vertebrate Paleontology (Memoir 4, supplement) 18, 690. 18. Grande, L. & Hilton, E.J. (2006). An Exquisitely Preserved Skeleton Representing a Primitive Sturgeon from the Upper Cretaceous Judith River Formation of Montana (Acipenseriformes: Acipenseridae: N. Gen. And Sp). Journal of Paleontology, 80, 1-39. 19. Hilton, E.J. (2003). Comparative osteology and phylogenetic systematics of fossil and living bony-tongue fishes (Actinopterygii, Teleostei, Osteoglossomorpha). Zoological Journal of the Linnean Society, 137, 1–100. 20. Hilton, E.J. & Forey, P.L. (2009). Redescription of †Chondrosteus acipenseroides Egerton, 1858 (Acipenseriformes, †Chondrosteidae) from the lower Lias of Lyme Regis (Dorset, England), with comments on the early evolution of sturgeons and paddlefishes. Journal of Systematic Palaeontology, 7, 427-453. 31 21. Li, G.-Q. & Wilson, M.V.H. (1999). Early divergence of Hiodontiformes sensu stricto in East Asia and phylogeny of some Late Mesozoic teleosts from China. In: Mesozoic Fishes 2 – Systematics and Fossil Record (eds. Arratia, G & Schultze, H-P). Verlag Dr. Friedrich Pfeil Mü nchen pp. 369-384. 22. Long, J.A., Choo, B. & Young, G.C. (2008). A new basal actinopterygian fish from the Middle Devonian Aztec Siltstone of Antarctica. Antarctic Science, 20. 23. Magallon, S. & Sanderson, M.J. (2001). Absolute diversification rates in angiosperm clades. Evolution, 55, 1762-1780. 24. Otero, O. (2004). Anatomy, systematics and phylogeny of both Recent and fossil latid fishes (Teleostei, Perciformes, Latidae). Zoological Journal of the Linnean Society, 141, 81-133. 25. Poyato-Ariza, F.J., Grande, T. & Diogo, R. (2010). Gonorynchiform Interrelationships: Historic Overview, Analysis, and Revised Systematics of the Group. In: Gonorynchiformes and Ostariophysan Relationships (eds. Grande, T, PoyatoAriza, FJ & Diogo, R). Science Publishers, pp. 227-338. 26. Santini, F. & Tyler, J.C. (2003). A phylogeny of the families of fossil and extant tetraodontiform fishes (Acanthomorpha, Tetraodontiformes), Upper Cretaceous to Recent. Zoological Journal of the Linnean Society, 139, 565–617. 27. Schultze, H.-P. & Cumbaa, S.L. (2001). Dialipina and the characters of basal actinopterygians. In: Major events in Early Vertebrate Evolution, Paleontology, Phylogeny, Genetics and Development (ed. Ahlberg, PE). Systematic Association Special Volume - Taylor & Francis London and New York., pp. 315-332. 28. Swartz, B.A. (2009). Devonian actinopterygian phylogeny and evolution based on a redescription ofStegotrachelus finlayi. Zoological Journal of the Linnean Society, 156, 750-784. 29. Taverne, L. (1997). Osorioichthys marginis, ‘Paéonisciforme’ du famennien de Belgique, et la phylogénie de Actinoptérygiens dévonians (Pisces). Bulletin de l’institut Royal des Sciences Naturelles de Belgique, 67, 57–78. 30. Taverne, L. (1998). Les ostéoglossomorphes marins de l’Éocène du Monte Bolca (Italie): Monopteros Volta 1796, Thrissopterus Heckel, 1856 et Foreyichthys Taverne, 1979. Considérations sur la phylogénie des téléostéens ostéoglossomorphes. Studie e Ricerche sui Giacimenti Terziari di Bolca, 7, 67– 158. 31. 32 Tyler, J.C. & Bannikov, A.F. (1997). Relationships of the Fossil and Recent Genera of Rabbitfishes (Acanthuroidei: Siganidae). Smithsonian Contributions to Zoology, 84, 1-34. 32. Tyler, J.C., Johnson, G.D., Nakamura, I. & Collette, B.B. (1989). Morphology of Luvarus imperialis (Luvaridae), with a phylogenetic analysis of the Acanthuroidei. Smithsonian Contributions to Zoology, 485, 1-78. 33. Tyler, J.C. & Santini, F. (2005). A phylogeny of the fossil and extant zeiform-like fishes, Upper Cretaceous to Recent, with comments on the putative zeomorph clade (Acanthomorpha). Zoologica Scripta, 34, 157-175. 34. Wilson, M.V.H. & Murray, A.M. (2008). Osteoglossomorpha: phylogeny, biogeography, and fossil record and the significance of key African and Chinese fossil taxa. Geological Society, London, Special Publications, 295, 185219. 35. Xu, G.-H. & Gao, K.-Q. (2011). A new scanilepiform from the Lower Triassic of northern Gansu Province, China, and phylogenetic relationships of nonteleostean Actinopterygii. Zoological Journal of the Linnean Society, 161, 595612. 36. Xu, G.-H. & Wu, F.-X. (2012). A deep-bodied ginglymodian fish from the Middle Triassic of eastern Yunnan Province, China, and the phylogeny of lower neopterygians. Chinese Science Bulletin, 57, 111-118. 37. Zhang, J.-Y. (2006). Phylogeny of Osteoglossomorpha. Vert. Pal. Asiat., 44, 43-59. 33