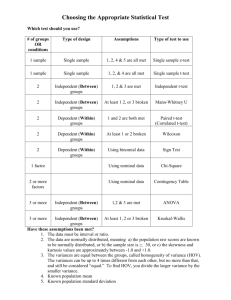
s1 <- .14
advertisement

Solutions to Exercises for Chapter 15 15.1. H 0 : s 12 = s 22 H1 : s 12 ¹ s 22 Given : n1 = 18, Test statistic : F= s12 s2 2 = .0196 .0484 s1 = .14, n2 = 13, s2 = .22 F, a = .10, F.05,17 ,12 = 0.4201, = 0.4050 Since 0.4050 < .4201, reject. With R: s1 <- .14 n1 <- 18 s2 <- .22 n2 <- 13 f.obs <- s1^2/s2^2 f.obs #Two-tailed critical values qf(c(.05,.95),n1-1,n2-1) > f.obs [1] 0.4049587 > #Two-tailed critical values > qf(c(.05,.95),n1-1,n2-1) [1] 0.4200526 2.5828389 F.95,17 ,12 = 2.583 15.2. H 0 : r1 = r 2 H1 : r1 ¹ r 2 Given : r1 = .93, n1 = 80, z1 = 1.658 r2 = .83, n2 = 30, z 2 = 1.188 Test statistic : z, a = .05, critical value = ±1.96 z= 1.658 -1.188 1 = 1 + 80 - 3 30 - 3 .47 .0129 + .0370 = .47 .2236 Since 2.10 > 1.96, reject. With R: r1 <- .93 n1 <- 80 r2 <- .83 n2 <- 30 r1.z <- .5*log(1+r1) - .5*log(1-r1) r2.z <- .5*log(1+r2) - .5*log(1-r2) z.obs <- (r1.z - r2.z)/sqrt((1/(n1-3))+(1/(n2-3))) z.obs > z.obs [1] 2.102533 > #Two-Tailed Critical values > round(qnorm(c(.025,.975)),3) [1] -1.96 1.96 = 2.10 15.3. This exercise calls for the comparison of two group having different sample sizes. We enter the data into R and create a data frame. All of our commands appear in Script 15.11. Next, we generate some descriptive statistics: > table(f.group) f.group white asian 12 25 > tapply(read,f.group,mean) white asian 24.00 30.32 > tapply(read,f.group,var) white asian 34.36364 249.31000 > tapply(read,f.group,sd) white asian 5.86205 15.78955 As you can see, there are more than twice as many Asian children as there are White children. The means appear to be different, but we also note that the variance of the reading scores of Asian children is approximately seven times that for the White children. Next, we will examine the distributions of scores for the two separate groups: > tapply(read,f.group,skewness) white asian 0.00974793 0.95786018 > tapply(read,f.group,SEsk) white asian 0.6373020 0.4636835 > tapply(read,f.group,kurtosis) white asian -0.72104644 0.01233870 > tapply(read,f.group,SEku) white asian 1.2322465 0.9017205 > boxplot(read ~ f.group) 60 50 40 30 20 10 white asian The indices for skewness and kurtosis suggest that the Asian scores are positively skewed. This is confirmed by the boxplot. At this point, we test the assumptions of normality and homogeneity of variance: >#Testing assumptions > tapply(read,f.group,shapiro.test) $white Shapiro-Wilk normality test W = 0.9604, p-value = 0.7894 $asian Shapiro-Wilk normality test W = 0.8958, p-value = 0.01492 > #Levene's test > leveneTest(read ~ f.group, data = data.var) Levene's Test for Homogeneity of Variance (center = median) Df F value group 1 Pr(>F) 5.1466 0.02957 35 > #Fligner-Killeen test > fligner.test(read ~ f.group) Fligner-Killeen test of homogeneity of variances Fligner-Killeen:med chi-squared = 4.9729, df = 1, p-value = 0.02575 The 12 reading scores for the White children are reasonably normally distributed; the 25 scores from the Asian children are not (p = .01492). Furthermore, both the Levene test and the Fligner–Killeen test indicate that the two sample variances suggest nonequivalent population variances with p-values of .02957 and .02575, respectively. Clearly, these data are problematic. As per the instructions, we now look at the pooled-sample t-test, the separate-sample t-test with the Welch correction, and the Mann–Whitney U/Wilcoxon test: > t.test(read ~ f.group,var.equal=T) Two Sample t-test data: read by f.group t = -1.3349, df = 35, p-value = 0.1905 alternative hypothesis: true difference in means is not equal to 0 95 percent confidence interval: -15.931763 3.291763 sample estimates: mean in group white mean in group asian 24.00 30.32 > t.test(read ~ f.group,var.equal=F) Welch Two Sample t-test data: read by f.group t = -1.764, df = 33.7, p-value = 0.0868 alternative hypothesis: true difference in means is not equal to 0 95 percent confidence interval: -13.603398 0.963398 sample estimates: mean in group white mean in group asian 24.00 30.32 > wilcox.test(read ~ f.group) Wilcoxon rank sum test with continuity correction data: read by f.group W = 124.5, p-value = 0.4165 alternative hypothesis: true location shift is not equal to 0 Warning message: In wilcox.test.default(x = c(29, 32, 23, 23, 20, 24, 17, 20, 33, : cannot compute exact p-value with ties As you can see from the output, the traditional pooled-sample t-test yields a p-value of .1905, indicating nonsignificance. The separate-sample t-test with the Welch correction to adjust for nonequivalent variances yields a p-value of .0868, also not significant, but “closer.” In looking back at the descriptive statistics, the Asian sample is more than twice the size of the White sample; it also hai the larger variance. Thus, when using the pooled-sample estimate of variance, the larger variance is weighted more heavily than the smaller variance, resulting in an overestimate of the population variance, which in turn results in an underestimate of the t-statistic. One might be tempted to turn to the nonparametric alternative, the Mann–Whitney U/Wilcoxon test in this context. It, too, indicates a nonsignificant difference between the two groups. However, recall that this procedure can only be interpreted in terms of location if the two group distributions are similar in shape and variability. Given that the charge was to compare the two groups with regard to central tendency, we would recommend reporting the results from the separate-sample estimate of the t-test.